

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
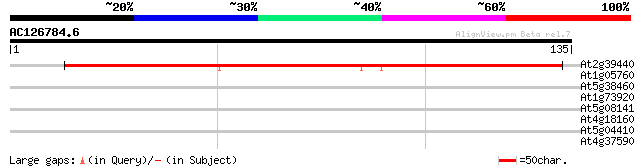
Query= AC126784.6 - phase: 0
(135 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g39440 hypothetical protein 133 3e-32
At1g05760 disease resistance protein RTM1 28 1.5
At5g38460 glucosyltransferase-like protein 27 4.2
At1g73920 putative lipase 27 4.2
At5g08141 bZip transcription factor AtbZip75 26 5.5
At4g18160 potassium channel - like protein 26 5.5
At5g04410 unknown protein 25 9.5
At4g37590 unknown protein 25 9.5
>At2g39440 hypothetical protein
Length = 773
Score = 133 bits (334), Expect = 3e-32
Identities = 67/142 (47%), Positives = 87/142 (61%), Gaps = 22/142 (15%)
Query: 14 DGIEDLSGRVHLLPCCIKHDGPTEVSQYFKPKPTGV----------VGEDGLPLQQSHFR 63
+ I DLS +VH LPCCI+ DGP EVS YFKPK + V DG+ +++HFR
Sbjct: 83 NSIVDLSSQVHQLPCCIRFDGPVEVSHYFKPKSSVRLMWKAVEFVDVEIDGVKTEEAHFR 142
Query: 64 GRLLEGTTLPLPHNYSGFVL---------GKKNS---VESSNSWETSVTFNDITYWNHDS 111
GR L+G T+ LP YSGFVL GK+ + E + WE FN++TYWNHDS
Sbjct: 143 GRKLQGATISLPSGYSGFVLRQASNLNANGKRKASMPTEDNQCWEVKAKFNNLTYWNHDS 202
Query: 112 VPSNNDDFSRAFHWIPVAEAVS 133
+PS +D F R+FHW +AEAV+
Sbjct: 203 LPSKDDTFFRSFHWFSIAEAVT 224
>At1g05760 disease resistance protein RTM1
Length = 174
Score = 28.1 bits (61), Expect = 1.5
Identities = 18/61 (29%), Positives = 28/61 (45%), Gaps = 2/61 (3%)
Query: 62 FRGRLLEGTTLPLPHNYSGFVLGKKNSVESSNSWETSVTFNDIT--YWNHDSVPSNNDDF 119
F G + + L PH Y + G+ E++N S+ FN T Y + S+ND F
Sbjct: 62 FSGNMFDVIELNYPHEYITGISGEYYKYEANNPHMRSLKFNTNTSEYGPFGTSGSSNDKF 121
Query: 120 S 120
+
Sbjct: 122 A 122
>At5g38460 glucosyltransferase-like protein
Length = 533
Score = 26.6 bits (57), Expect = 4.2
Identities = 9/19 (47%), Positives = 12/19 (62%)
Query: 95 WETSVTFNDITYWNHDSVP 113
W + T+ND+TYW D P
Sbjct: 91 WYRNGTYNDLTYWGLDYPP 109
>At1g73920 putative lipase
Length = 704
Score = 26.6 bits (57), Expect = 4.2
Identities = 13/36 (36%), Positives = 18/36 (49%), Gaps = 2/36 (5%)
Query: 84 GKKNSVESSNSWETSVTFND--ITYWNHDSVPSNND 117
GK N +E + WE TF + W H+ V S N+
Sbjct: 44 GKANILEGLHGWELRPTFRGPRLPRWMHNGVSSFNE 79
>At5g08141 bZip transcription factor AtbZip75
Length = 137
Score = 26.2 bits (56), Expect = 5.5
Identities = 13/33 (39%), Positives = 20/33 (60%), Gaps = 3/33 (9%)
Query: 90 ESSNSWETSVTFND---ITYWNHDSVPSNNDDF 119
E+S ET +F+D I+Y NH+ + N +DF
Sbjct: 94 ENSQLKETVSSFHDQYTISYGNHEGILGNTNDF 126
>At4g18160 potassium channel - like protein
Length = 436
Score = 26.2 bits (56), Expect = 5.5
Identities = 15/49 (30%), Positives = 24/49 (48%)
Query: 55 LPLQQSHFRGRLLEGTTLPLPHNYSGFVLGKKNSVESSNSWETSVTFND 103
+P+ S F+ RL+ G P + S F+ K SS+S + F+D
Sbjct: 38 MPMTPSEFKERLIFGPFSCSPRDSSHFIDSMKQPSPSSSSTAVNNPFSD 86
>At5g04410 unknown protein
Length = 567
Score = 25.4 bits (54), Expect = 9.5
Identities = 14/41 (34%), Positives = 18/41 (43%)
Query: 73 PLPHNYSGFVLGKKNSVESSNSWETSVTFNDITYWNHDSVP 113
P P V G K +VE S E S + D + + DS P
Sbjct: 401 PAPKELEKEVAGGKEAVEEKESGEGSSSKQDTDFKDFDSAP 441
>At4g37590 unknown protein
Length = 580
Score = 25.4 bits (54), Expect = 9.5
Identities = 14/41 (34%), Positives = 24/41 (58%)
Query: 58 QQSHFRGRLLEGTTLPLPHNYSGFVLGKKNSVESSNSWETS 98
Q+SH +G+L++G L + +S GK+ S + +S TS
Sbjct: 521 QESHNQGKLMKGGGLGVSRVFSKLWSGKERSGDMISSSGTS 561
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.136 0.429
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,585,426
Number of Sequences: 26719
Number of extensions: 156463
Number of successful extensions: 323
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 316
Number of HSP's gapped (non-prelim): 9
length of query: 135
length of database: 11,318,596
effective HSP length: 88
effective length of query: 47
effective length of database: 8,967,324
effective search space: 421464228
effective search space used: 421464228
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 54 (25.4 bits)
Medicago: description of AC126784.6


