

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126780.2 - phase: 0 /pseudo
(297 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
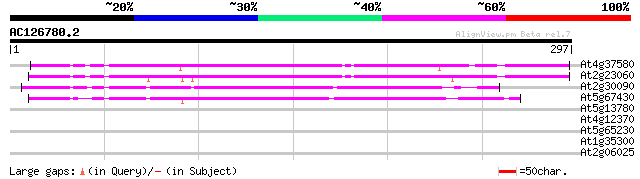
At4g37580 probable N-acetyltransferase hookless 1 167 5e-42
At2g23060 similar to hookless1 (HLS1) 162 3e-40
At2g30090 hookless1-like protein 148 3e-36
At5g67430 N-acetyltransferase hookless1-like protein 142 2e-34
At5g13780 Unknown protein (MXE10.5) 33 0.18
At4g12370 putative protein 30 2.0
At5g65230 transcription factor-like protein 28 5.9
At1g35300 hypothetical protein 28 5.9
At2g06025 unknown protein 28 7.8
>At4g37580 probable N-acetyltransferase hookless 1
Length = 403
Score = 167 bits (424), Expect = 5e-42
Identities = 110/306 (35%), Positives = 164/306 (52%), Gaps = 35/306 (11%)
Query: 12 VIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHVMLVAE 71
V+RE+D RD+ V +ER C E+ + K S+FT+++ GDP+ RIR P ++MLVAE
Sbjct: 3 VVREYDPTRDLVGVEDVERRC-EVGPSGK--LSLFTDLL--GDPICRIRHSPSYLMLVAE 57
Query: 72 M-VESKELVGVVKGCIKSV--------------QTPSGSLFKMGCILGLRVSPIHRRKGV 116
M E KE+VG+++GCIK+V K+ +LGLRVSP HRR+G+
Sbjct: 58 MGTEKKEIVGMIRGCIKTVTCGQKLDLNHKSQNDVVKPLYTKLAYVLGLRVSPFHRRQGI 117
Query: 117 GLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTN 176
G KLV +EEW NGA+Y+++ATE +N AS NLFT KC Y F + I ++P + N
Sbjct: 118 GFKLVKMMEEWFRQNGAEYSYIATENDNQASVNLFTGKCGYSEFRTPSILVNPVYAHRVN 177
Query: 177 HISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSLGTWVS------YYKD 230
+S++ V + K+ A + Y T E +P D+D +L KLSLGT+V+ Y
Sbjct: 178 -VSRR-VTVIKLEPVDAETLYRIRFSTTEFFPRDIDSVLNNKLSLGTFVAVPRGSCYGSG 235
Query: 231 EGFKLNIEDIITHKSTTSSSWIIFSLWNTCEACDNNLQVKTKLFQPLRFLHATLNHAKDK 290
G + + SW + S+WN C ++ ++ + LR + A DK
Sbjct: 236 SGSWPGSAKFLEY---PPESWAVLSVWN----CKDSFLLEVRGASRLRRVVAKTTRVVDK 288
Query: 291 ICPLFE 296
P +
Sbjct: 289 TLPFLK 294
>At2g23060 similar to hookless1 (HLS1)
Length = 413
Score = 162 bits (409), Expect = 3e-40
Identities = 108/311 (34%), Positives = 166/311 (52%), Gaps = 36/311 (11%)
Query: 11 VVIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHVMLVA 70
V +RE+D +D+ V +ER C E+ K S+FT+++ GDP+ R+R P ++MLVA
Sbjct: 5 VEVREYDPSKDLATVEDVERRC-EVGPAGK--LSLFTDLL--GDPICRVRHSPSYLMLVA 59
Query: 71 EM--VESKELVGVVKGCIKSVQ----------TPSGS----------LFKMGCILGLRVS 108
E+ E KELVG+++GCIK+V T + S K+ ILGLRVS
Sbjct: 60 EIGPKEKKELVGMIRGCIKTVTCGITTKRLDLTHNKSQNDVVITKPLYTKLAYILGLRVS 119
Query: 109 PIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLH 168
P HRR+G+G KLV ++E+W NGA+Y++ ATE +N+AS NLFT KC Y F + I ++
Sbjct: 120 PTHRRQGIGFKLVKAMEDWFSQNGAEYSYFATENDNHASVNLFTGKCGYAEFRTPSILVN 179
Query: 169 PPTSFPTNHISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSLGTWVSYY 228
P + N IS++ V + K+ A Y T E +P D+D +L KLSLGT+V+
Sbjct: 180 PVYAHRVN-ISRR-VTVIKLEPSDAELLYRLRFSTTEFFPRDIDSVLNNKLSLGTFVAVP 237
Query: 229 KDEGF---KLNIEDIITHKSTTSSSWIIFSLWNTCEACDNNLQVKTKLFQPLRFLHATLN 285
+ + + SW + S+WN C ++ +++ + LR + +
Sbjct: 238 RGSCYGSGSRSWPGSAKFLEYPPDSWAVLSVWN----CKDSFRLEVRGASRLRRVVSKAT 293
Query: 286 HAKDKICPLFE 296
DK P +
Sbjct: 294 RMVDKTLPFLK 304
>At2g30090 hookless1-like protein
Length = 386
Score = 148 bits (374), Expect = 3e-36
Identities = 93/255 (36%), Positives = 152/255 (59%), Gaps = 26/255 (10%)
Query: 7 IESKVVIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHV 66
++ +VVIR +D+ RD +G++E++C EI + +FT+ + GDP+ RIR P +
Sbjct: 9 VDEEVVIRCYDDRRDRIQMGRMEKSC-EIGHDHQT--LLFTDTL--GDPICRIRNSPFFI 63
Query: 67 MLVAEMVESKELVGVVKGCIKSVQTPSGSLFKMGCILGLRVSPIHRRKGVGLKLVTSIEE 126
MLVA + +LVG ++G +K V+ S+ ++G +LGLRV P +RR+G+G LV +EE
Sbjct: 64 MLVAGV--GNKLVGSIQGSVKPVEFHDKSV-RVGYVLGLRVVPSYRRRGIGSILVRKLEE 120
Query: 127 WMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTNHIS-KKDVKI 185
W ++ ADYA++ATEK+N AS LF + Y F + I ++P P + D+ I
Sbjct: 121 WFESHNADYAYMATEKDNEASHGLFIGRLGYVVFRNPAILVNPVN--PGRGLKLPSDIGI 178
Query: 186 DKISIDQAISFYTR-ILKTKELYPLDMDIILKEKLSLGTWVSYYKDEGFKLNIEDIITHK 244
K+ + +A S Y R + T E +P D++ IL+ KLS+GTWV+YY N+++
Sbjct: 179 RKLKVKEAESLYRRNVAATTEFFPDDINKILRNKLSIGTWVAYYN------NVDN----- 227
Query: 245 STTSSSWIIFSLWNT 259
+ SW + S+W++
Sbjct: 228 ---TRSWAMLSVWDS 239
>At5g67430 N-acetyltransferase hookless1-like protein
Length = 386
Score = 142 bits (359), Expect = 2e-34
Identities = 98/272 (36%), Positives = 145/272 (53%), Gaps = 34/272 (12%)
Query: 11 VVIREFDEDRDVKVVGKLERNCTEINGTTKKGFSIFTNMMSNGDPLSRIRFYPLHVMLVA 70
VV+RE+D RD+ V +LE +C E+ S+ ++M GDPL+RIR P MLVA
Sbjct: 8 VVVREYDPKRDLTSVEELEESC-EVG-------SLLVDLM--GDPLARIRQSPSFHMLVA 57
Query: 71 EMVESKELVGVVKGCIKSVQ------------TPSGSLFKMGCILGLRVSPIHRRKGVGL 118
E+ E+VG+++G IK V +P + K+ + GLRVSP +RR G+GL
Sbjct: 58 EI--GNEIVGMIRGTIKMVTRGVNALRQADDVSPEINTTKLAFVSGLRVSPFYRRMGIGL 115
Query: 119 KLVTSIEEWMLTNGADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTNHI 178
KLV +EEW L N A Y+++ TE +N AS LFT K Y F + ++P F
Sbjct: 116 KLVQRLEEWFLRNDAVYSYVQTENDNIASVKLFTEKSGYSKFRTPTFLVNP--VFNHRVT 173
Query: 179 SKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSLGTWVSYYKDEGFKLNIE 238
+ VKI K++ A S Y T E +P D++ IL KLSLGT+++ + +
Sbjct: 174 VSRRVKIIKLAPSDAESLYRNRFSTTEFFPSDINSILTNKLSLGTYLAVPRGG------D 227
Query: 239 DIITHKSTTSSSWIIFSLWNTCEACDNNLQVK 270
++ + SW + S+WN+ + LQVK
Sbjct: 228 NVSGSLPDQTGSWAVISIWNSKDV--YRLQVK 257
>At5g13780 Unknown protein (MXE10.5)
Length = 192
Score = 33.1 bits (74), Expect = 0.18
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 1/59 (1%)
Query: 100 GCILGLRVSPIHRRKGVGLKLVTSIEEWM-LTNGADYAFLATEKNNNASKNLFTNKCNY 157
G I L V HR+ G+ KL+T+ + M A+Y L ++N A+ NL+T Y
Sbjct: 70 GHITSLAVLRTHRKLGLATKLMTAAQAAMEQVYEAEYVSLHVRRSNRAAFNLYTETLGY 128
>At4g12370 putative protein
Length = 300
Score = 29.6 bits (65), Expect = 2.0
Identities = 18/48 (37%), Positives = 26/48 (53%), Gaps = 1/48 (2%)
Query: 6 SIESKVVIREFDEDRDVKVVGKLER-NCTEINGTTKKGFSIFTNMMSN 52
++ V + F D + +V KLE N EIN TT G +IFT + S+
Sbjct: 188 AVRKGVFFKIFTCDGEKPIVHKLEDINWEEINSTTIDGLTIFTGLYSS 235
>At5g65230 transcription factor-like protein
Length = 310
Score = 28.1 bits (61), Expect = 5.9
Identities = 23/78 (29%), Positives = 37/78 (46%), Gaps = 6/78 (7%)
Query: 124 IEEWMLTNGADYAFLAT---EKNNNASKNLFTNKCNYFNFTSLIIFLHPPTSFPTNHISK 180
I+E +T+ D FL++ + NNN +L T N+F+ + I+LH P S +
Sbjct: 201 IKENSVTSNIDLGFLSSHLQDFNNNNLPSLKTLDDNHFSQNTSPIWLHEPPSLNQTMLPT 260
Query: 181 KD---VKIDKISIDQAIS 195
D +D +QA S
Sbjct: 261 HDPCAQSVDGFGSNQASS 278
>At1g35300 hypothetical protein
Length = 477
Score = 28.1 bits (61), Expect = 5.9
Identities = 17/45 (37%), Positives = 25/45 (54%), Gaps = 3/45 (6%)
Query: 217 EKLSLGTWVSYYKDEGFKLNIEDIITHKSTTSSS--WIIFSLWNT 259
+ L +GT V YY +E K ++DI++ TT WI +LW T
Sbjct: 274 DPLIIGT-VQYYFNEIVKRRLKDIVSTARTTREQPPWIGETLWGT 317
>At2g06025 unknown protein
Length = 288
Score = 27.7 bits (60), Expect = 7.8
Identities = 13/54 (24%), Positives = 27/54 (49%)
Query: 98 KMGCILGLRVSPIHRRKGVGLKLVTSIEEWMLTNGADYAFLATEKNNNASKNLF 151
+ G I L V+ RR+G+ ++ E +G + ++ KNN+ ++ L+
Sbjct: 208 RYGYIANLCVAKSARRQGIACNMLRFAVESARLSGVEQVYVHVHKNNSVAQELY 261
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,825,332
Number of Sequences: 26719
Number of extensions: 285723
Number of successful extensions: 684
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 662
Number of HSP's gapped (non-prelim): 11
length of query: 297
length of database: 11,318,596
effective HSP length: 99
effective length of query: 198
effective length of database: 8,673,415
effective search space: 1717336170
effective search space used: 1717336170
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC126780.2


