

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.36 + phase: 0
(475 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
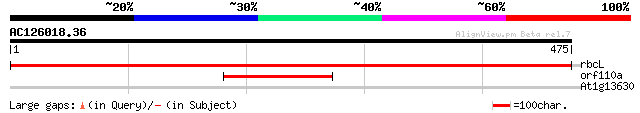
Score E
Sequences producing significant alignments: (bits) Value
rbcL -chloroplast genome- large subunit of riblose-1,5-bisphosph... 930 0.0
orf110a -mitochondrial genome- 181 8e-46
At1g13630 hypothetical protein 32 0.76
>rbcL -chloroplast genome- large subunit of riblose-1,5-bisphosphate
carboxylase/oxygenase
Length = 479
Score = 930 bits (2403), Expect = 0.0
Identities = 446/475 (93%), Positives = 463/475 (96%)
Query: 1 MSPQTETKATVGFKAGVKDYRLTYYTPDYETKDTDILAAFRVSPQPGVPAEEAGAAVAAE 60
MSPQTETKA+VGFKAGVK+Y+LTYYTP+YETKDTDILAAFRV+PQPGVP EEAGAAVAAE
Sbjct: 1 MSPQTETKASVGFKAGVKEYKLTYYTPEYETKDTDILAAFRVTPQPGVPPEEAGAAVAAE 60
Query: 61 SSTGTWTTVWTDGLTSLDRYKGRCYHIEPVAGEESQFIAYVAYPLDLFEEGSVTNMFTSI 120
SSTGTWTTVWTDGLTSLDRYKGRCYHIEPV GEE+QFIAYVAYPLDLFEEGSVTNMFTSI
Sbjct: 61 SSTGTWTTVWTDGLTSLDRYKGRCYHIEPVPGEETQFIAYVAYPLDLFEEGSVTNMFTSI 120
Query: 121 VGNVFGFKALRALRLEDLRIPVAYVKTFQGPPHGIQVERDKLNKYGRPLLGCTIKPKLGL 180
VGNVFGFKAL ALRLEDLRIP AY KTFQGPPHGIQVERDKLNKYGRPLLGCTIKPKLGL
Sbjct: 121 VGNVFGFKALAALRLEDLRIPPAYTKTFQGPPHGIQVERDKLNKYGRPLLGCTIKPKLGL 180
Query: 181 SAKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEAIYKAQAETGEIKGHYL 240
SAKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEAIYK+QAETGEIKGHYL
Sbjct: 181 SAKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEAIYKSQAETGEIKGHYL 240
Query: 241 NATAGTCEDMMKRAVFARELGVPIVMHDYLTGGFTANTTLAHYCRDNGLLLHIHRAMHAV 300
NATAGTCE+M+KRAVFARELGVPIVMHDYLTGGFTANT+L+HYCRDNGLLLHIHRAMHAV
Sbjct: 241 NATAGTCEEMIKRAVFARELGVPIVMHDYLTGGFTANTSLSHYCRDNGLLLHIHRAMHAV 300
Query: 301 IDRQKNHGMHFRVLAKALRMSGGDHIHAGTVVGKLEGERDITLGFVDLLRDDFVEKDRSR 360
IDRQKNHGMHFRVLAKALR+SGGDHIHAGTVVGKLEG+R+ TLGFVDLLRDD+VEKDRSR
Sbjct: 301 IDRQKNHGMHFRVLAKALRLSGGDHIHAGTVVGKLEGDRESTLGFVDLLRDDYVEKDRSR 360
Query: 361 GIFFTQDWVSLPGVLPVASGGIHVWHMPALTEIFGDDSVLQFGGGTLGHPWGNAPGAVAN 420
GIFFTQDWVSLPGVLPVASGGIHVWHMPALTEIFGDDSVLQFGGGTLGHPWGNAPGAVAN
Sbjct: 361 GIFFTQDWVSLPGVLPVASGGIHVWHMPALTEIFGDDSVLQFGGGTLGHPWGNAPGAVAN 420
Query: 421 RVALEACVQARNEGRDLAREGNEIIREATKWSPELAAACEVWKEIKFEFPAMDTI 475
RVALEACVQARNEGRDLA EGNEIIREA KWSPELAAACEVWKEI F FP +D +
Sbjct: 421 RVALEACVQARNEGRDLAVEGNEIIREACKWSPELAAACEVWKEITFNFPTIDKL 475
>orf110a -mitochondrial genome-
Length = 110
Score = 181 bits (459), Expect = 8e-46
Identities = 85/92 (92%), Positives = 88/92 (95%)
Query: 182 AKNYGRAVYECLRGGLDFTKDDENVNSQPFMRWRDRFLFCAEAIYKAQAETGEIKGHYLN 241
AKNYGRAVYECLRGGL FTKDDENVNSQPFMRWRDRFLFCAEA+YKAQAETG IKGHYLN
Sbjct: 14 AKNYGRAVYECLRGGLYFTKDDENVNSQPFMRWRDRFLFCAEAVYKAQAETGGIKGHYLN 73
Query: 242 ATAGTCEDMMKRAVFARELGVPIVMHDYLTGG 273
ATAGTCE+M+KRAVFARELGVPIVMHDYL G
Sbjct: 74 ATAGTCEEMIKRAVFARELGVPIVMHDYLNRG 105
>At1g13630 hypothetical protein
Length = 764
Score = 32.0 bits (71), Expect = 0.76
Identities = 26/86 (30%), Positives = 38/86 (43%), Gaps = 10/86 (11%)
Query: 275 TANTTLAHYCRDNGLLLHIHRAMHAVIDRQKNHGMHFRVLAKALRMSGGDHIHAGTVVGK 334
T N+T H G I M + R+ HG HFR L LR H+H ++ +
Sbjct: 8 TTNSTSDH----RGFYKEILFGMKKIGFREFLHGYHFRGLVSELR-----HVHVEEIMDE 58
Query: 335 LEGE-RDITLGFVDLLRDDFVEKDRS 359
L E D+++ F LRD + + S
Sbjct: 59 LMSESSDLSVWFFKELRDIYAFRHSS 84
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.138 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,963,290
Number of Sequences: 26719
Number of extensions: 471739
Number of successful extensions: 855
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 853
Number of HSP's gapped (non-prelim): 3
length of query: 475
length of database: 11,318,596
effective HSP length: 103
effective length of query: 372
effective length of database: 8,566,539
effective search space: 3186752508
effective search space used: 3186752508
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC126018.36


