

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.19 - phase: 0
(686 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
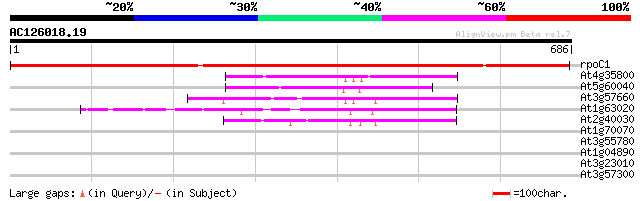
Sequences producing significant alignments: (bits) Value
rpoC1 -chloroplast genome- RNA polymerase beta' subunit-1 1243 0.0
At4g35800 DNA-directed RNA polymerase (EC 2.7.7.6) II largest chain 129 7e-30
At5g60040 DNA-directed RNA polymerase - like protein 107 2e-23
At3g57660 DNA-directed RNA polymerase I 190K chain - like protein 87 3e-17
At1g63020 RNA polymerase IIA largest subunit, putative 79 1e-14
At2g40030 putative DNA-directed RNA polymerase II subunit 75 9e-14
At1g70070 Similar to Synechocystis antiviral protein (At1g70070) 33 0.69
At3g55780 beta-1,3-glucanase - like protein 32 1.5
At1g04890 hypothetical protein 31 2.0
At3g23010 disease resistance protein, putative 31 2.6
At3g57300 helicase-like protein 29 7.7
>rpoC1 -chloroplast genome- RNA polymerase beta' subunit-1
Length = 680
Score = 1243 bits (3215), Expect = 0.0
Identities = 605/684 (88%), Positives = 642/684 (93%), Gaps = 6/684 (0%)
Query: 1 MIDQYKHQHLRIGSVSPEQISAWAKKILPNGEIVGEVTKPYTLHYKTNKPEKDGLFCERI 60
MID+YKHQ LRIG VSP+QISAWA KI+PNGEIVGEVTKPYT HYKTNKPEKDGLFCERI
Sbjct: 1 MIDRYKHQQLRIGLVSPQQISAWATKIIPNGEIVGEVTKPYTFHYKTNKPEKDGLFCERI 60
Query: 61 FGPIKSGICACGNYRVIGDKKDQPKFCEQCGVEFVDSRVRRYQMGYIKLACPVTHVWYLK 120
FGPIKSGICACGNYRVIGD+K+ PKFCEQCGVEFVDSR+RRYQMGYIKL CPVTHVWYLK
Sbjct: 61 FGPIKSGICACGNYRVIGDEKEDPKFCEQCGVEFVDSRIRRYQMGYIKLTCPVTHVWYLK 120
Query: 121 RLPSYIASLLDKPLKELEGLVYCDFSFARPVVKKPTFLRLRGSFEYEIQSWKYSIPLFFA 180
RLPSYIA+LLDKPLKELEGLVYCDFSFARP+ KKPTFLRLRGSFEYEIQSWKYSIPLFF
Sbjct: 121 RLPSYIANLLDKPLKELEGLVYCDFSFARPITKKPTFLRLRGSFEYEIQSWKYSIPLFFT 180
Query: 181 TQGFDTFRNREISSGAGAIREQLVDLDLRIIMDSSLVEWKELGEEGSADNENENEWEDRK 240
TQGFD FRNREIS+GAGAIREQL DLDLRII+++SLVEWK+LGEEG NE WEDRK
Sbjct: 181 TQGFDIFRNREISTGAGAIREQLADLDLRIIIENSLVEWKQLGEEGPTGNE----WEDRK 236
Query: 241 VGRRKNFLVRRMELVKHFIRTNIEPEWMVLSLLPVLPPELRPIIQIDGGKLMSSDINELY 300
+ RRK+FLVRRMEL KHFIRTNIEPEWMVL LLPVLPPELRPIIQI+GGKLMSSDINELY
Sbjct: 237 IVRRKDFLVRRMELAKHFIRTNIEPEWMVLCLLPVLPPELRPIIQIEGGKLMSSDINELY 296
Query: 301 RRVIYRNNTLIDLLTTSRSTPGELVMCQEKLVQEAVDTLLDNGIRGQPMRDGHNKVYKSF 360
RRVIYRNNTL DLLTTSRSTPGELVMCQEKLVQEAVDTLLDNGIRGQPMRDGHNKVYKSF
Sbjct: 297 RRVIYRNNTLTDLLTTSRSTPGELVMCQEKLVQEAVDTLLDNGIRGQPMRDGHNKVYKSF 356
Query: 361 SDIIEGKEGRFRETLLGKRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTFLIRGLIR 420
SD+IEGKEGRFRETLLGKRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTF+IRGLIR
Sbjct: 357 SDVIEGKEGRFRETLLGKRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTFVIRGLIR 416
Query: 421 KHFASNIGVAKSKIREKEPIVWEILQEVMRGHPVLLNRAPTLHRLGIQAFQPILVEGRAI 480
+H ASNIGVAKS+IREK+PIVWEILQEVM+GHPVLLNRAPTLHRLGIQ+FQPILVEGR I
Sbjct: 417 QHLASNIGVAKSQIREKKPIVWEILQEVMQGHPVLLNRAPTLHRLGIQSFQPILVEGRTI 476
Query: 481 CLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSHTNLLSPAIGDPISVPTQDML 540
CLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSH NLLSPAIGDPISVPTQDML
Sbjct: 477 CLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSHMNLLSPAIGDPISVPTQDML 536
Query: 541 IGLYVLTSGNRRGICANRYNPFNCRNSKNEKISNNNSKYMKKKEPFFCNSYDAIGAYRQK 600
IGLYVLTSG RRGICANRYNP N +N +NE+I N KY KEPFFCNSYDAIGAYRQK
Sbjct: 537 IGLYVLTSGTRRGICANRYNPCNRKNYQNERIYETNYKY--TKEPFFCNSYDAIGAYRQK 594
Query: 601 RINLDSPFWLRWRIDQCIMSSREVPIEVHYESFGTYYEIYGHYLVIRSIKKEIRCIYIRT 660
+INLDSP WLRW++DQ +++SREVPIEVHYESFG Y+EIY HYL++RS+KKE CIYIRT
Sbjct: 595 KINLDSPLWLRWQLDQRVIASREVPIEVHYESFGNYHEIYAHYLIVRSVKKENFCIYIRT 654
Query: 661 TVGHISFYREIEEAIQGFSRAYSY 684
TVGHISFYREIEEAIQGFS+A SY
Sbjct: 655 TVGHISFYREIEEAIQGFSQACSY 678
>At4g35800 DNA-directed RNA polymerase (EC 2.7.7.6) II largest chain
Length = 1840
Score = 129 bits (323), Expect = 7e-30
Identities = 89/311 (28%), Positives = 147/311 (46%), Gaps = 31/311 (9%)
Query: 265 PEWMVLSLLPVLPPELRPIIQIDGGKLMSSDINELYRRVIYRNNTLIDLLTTSRSTPGEL 324
P+WM+L +LP+ PP +RP + +D D+ +I N L P +
Sbjct: 243 PDWMILEVLPIPPPPVRPSVMMDATSRSEDDLTHQLAMIIRHNENL--KRQEKNGAPAHI 300
Query: 325 VMCQEKLVQEAVDTLLDNGIRGQPMRDGHN-KVYKSFSDIIEGKEGRFRETLLGKRVDYS 383
+ +L+Q + T DN + GQP + + KS ++ KEGR R L+GKRVD+S
Sbjct: 301 ISEFTQLLQFHIATYFDNELPGQPRATQKSGRPIKSICSRLKAKEGRIRGNLMGKRVDFS 360
Query: 384 GRSVIVVGPSLSLHRCGLPREIAIEL--------FQTFLIRGLI------------RKHF 423
R+VI P++++ G+P IA+ L + ++ L+ K+
Sbjct: 361 ARTVITPDPTINIDELGVPWSIALNLTYPETVTPYNIERLKELVDYGPHPPPGKTGAKYI 420
Query: 424 ASNIG-------VAKSKIREKEPIVWEILQEVMRGHPVLLNRAPTLHRLGIQAFQPILVE 476
+ G + KS + E + +++ + + G VL NR P+LH++ I + ++
Sbjct: 421 IRDDGQRLDLRYLKKSSDQHLE-LGYKVERHLQDGDFVLFNRQPSLHKMSIMGHRIRIMP 479
Query: 477 GRAICLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSHTNLLSPAIGDPISVPT 536
L+ V +NADFDGD+M +HVP S E +AE LM ++SP P+
Sbjct: 480 YSTFRLNLSVTSPYNADFDGDEMNMHVPQSFETRAEVLELMMVPKCIVSPQANRPVMGIV 539
Query: 537 QDMLIGLYVLT 547
QD L+G +T
Sbjct: 540 QDTLLGCRKIT 550
>At5g60040 DNA-directed RNA polymerase - like protein
Length = 1328
Score = 107 bits (267), Expect = 2e-23
Identities = 78/277 (28%), Positives = 137/277 (49%), Gaps = 27/277 (9%)
Query: 265 PEWMVLSLLPVLPPELRPIIQIDGGKLMSSDINELYRRVIYRNNTLIDLLTTSRSTPGEL 324
PE ++++ + V P +RP + I G + +D+ +++I N +L +L+ S+P +
Sbjct: 250 PENLIITCMLVPPLSIRPSVMIGGIQSNENDLTARLKQIILGNASLHKILSQPTSSPKNM 309
Query: 325 VMCQEKLVQEAVDTLLDNGIRGQPMRDGHNKVYKSFSDIIEGKEGRFRETLLGKRVDYSG 384
+ VQ V +++ +RG + + + ++GK GRFR L GKRV+++G
Sbjct: 310 QVWDT--VQIEVARYINSEVRGCQNQPEEHPL-SGILQRLKGKGGRFRANLSGKRVEFTG 366
Query: 385 RSVIVVGPSLSLHRCGLPREIA-------------IELFQTFLIRGLIRKHFASN----- 426
R+VI P+L + G+P +A IE + + G + A N
Sbjct: 367 RTVISPDPNLKITEVGIPILMAQILTFPECVSRHNIEKLRQCVRNGPNKYPGARNVRYPD 426
Query: 427 ------IGVAKSKIREKEPIVWEILQEVMRGHPVLLNRAPTLHRLGIQAFQPILVEGRAI 480
+G + +I ++ I + + + G VL NR P+LHR+ I + ++ R +
Sbjct: 427 GSSRTLVGDYRKRIADELAIGCIVDRHLQEGDVVLFNRQPSLHRMSIMCHRARIMPWRTL 486
Query: 481 CLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLM 517
+ VC +NADFDGD+M +HVP + EA+ EA LM
Sbjct: 487 RFNESVCNPYNADFDGDEMNMHVPQTEEARTEAITLM 523
>At3g57660 DNA-directed RNA polymerase I 190K chain - like protein
Length = 1670
Score = 87.0 bits (214), Expect = 3e-17
Identities = 95/384 (24%), Positives = 159/384 (40%), Gaps = 62/384 (16%)
Query: 218 EWKELGEEGSADNENENEWEDRKVGRRKNFLVRRMELVKHFIR----------TNIEPEW 267
E +E E +A+ E N D +N L + F I+
Sbjct: 281 ETREKSTEVAAEFEEHNSKRDLLPSEVRNILKHLWQNEHEFCSFIGDLWQSGSEKIDYSM 340
Query: 268 MVLSLLPVLPPELRPIIQIDGGKLMSSDINELYRRVIYRNNTLIDLLTTSRSTPGELVMC 327
L + V P + RP G +M +VI NN L + T V+
Sbjct: 341 FFLESVLVPPTKFRPPTT-GGDSVMEHPQTVGLNKVIESNNILGNACTNKLDQ--SKVIF 397
Query: 328 QEKLVQEAVDTLLDNGIRG-QPMRDGHNKVYKSFSDIIEGKEGRFRETLLGKRVDYSGRS 386
+ + +QE+V+ L D+ Q RD ++E KEG FR+ ++GKRV+++ RS
Sbjct: 398 RWRNLQESVNVLFDSKTATVQSQRDS-----SGICQLLEKKEGLFRQKMMGKRVNHACRS 452
Query: 387 VIVVGPSLSLHRCGLPREIAIEL-------------FQTFLIRGLI----RKHFASNIGV 429
VI P ++++ G+P A++L + +I G H++
Sbjct: 453 VISPDPYIAVNDIGIPPCFALKLTYPERVTPWNVEKLREAIINGPDIHPGATHYSDKSST 512
Query: 430 AKSKIREK--EPIVWEIL-----------------------QEVMRGHPVLLNRAPTLHR 464
K EK I ++L + + G VL+NR PTLH+
Sbjct: 513 MKLPSTEKARRAIARKLLSSRGATTELGKTCDINFEGKTVHRHMRDGDIVLVNRQPTLHK 572
Query: 465 LGIQAFQPILVEG-RAICLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSHTNL 523
+ A + +++G + + LH C +NADFDGD+M VH P ++AEA ++ ++
Sbjct: 573 PSLMAHKVRVLKGEKTLRLHYANCSTYNADFDGDEMNVHFPQDEISRAEAYNIVNANNQY 632
Query: 524 LSPAIGDPISVPTQDMLIGLYVLT 547
P+ G+P+ QD ++ +LT
Sbjct: 633 ARPSNGEPLRALIQDHIVSSVLLT 656
>At1g63020 RNA polymerase IIA largest subunit, putative
Length = 1453
Score = 78.6 bits (192), Expect = 1e-14
Identities = 108/485 (22%), Positives = 198/485 (40%), Gaps = 65/485 (13%)
Query: 87 CEQCGVEFVDSRVRRYQMGYIKLACPVTHVWYLKRLPSYIASLLDKPLKELEGLVYCDFS 146
C CG + D +V G I A + + ++LK +A+LL+K + + F
Sbjct: 56 CRTCGSK--DRKVCEGHFGVINFAYSIINPYFLKE----VAALLNKICPGCKYIRKKQFQ 109
Query: 147 FARPVVKKPTFLRLRGSFEYEIQSWKYSIPLFFATQGFDTFRNREISSGAGAIREQLVDL 206
++ + L Y + ++ + F G N E L+ L
Sbjct: 110 ITEDQPERCRYCTLNTG--YPLMKFRVTTKEVFRRSGIVVEVNEE----------SLMKL 157
Query: 207 DLRIIMDSSLVEWKELGEEGSADNENENEWEDRKVGRRKNFLVRRMELVKHFIRTNIEPE 266
R ++ W L ++ + D R++ + + + I+ +I P
Sbjct: 158 KKRGVLTLPPDYWSFLPQDSNIDESCLKP--TRRIITHAQVYALLLGIDQRLIKKDI-PM 214
Query: 267 WMVLSL--LPVLPPELRP---IIQIDGGKLMSSDINELYRRVI-YRNNTLIDLLTTSRST 320
+ L L PV P R + Q +G +L+ + +Y++++ + NTL
Sbjct: 215 FNSLGLTSFPVTPNGYRVTEIVHQFNGARLIFDERTRIYKKLVGFEGNTL--------EL 266
Query: 321 PGELVMCQE--KLVQEAVDTLLDNGIRGQPMRDGHNKVYKSFSDIIEGKEGRF-RETLLG 377
++ C + +L E V + D+ Y+ SD + RF ++ LLG
Sbjct: 267 SSRVMECMQYSRLFSETVSSSKDSA-----------NPYQKKSDTPKLCGLRFMKDVLLG 315
Query: 378 KRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTFLI-----RGLIRKHFASNIGVAKS 432
KR D++ R+V+V PSL L+ G+P IA L + + L+ + + +
Sbjct: 316 KRSDHTFRTVVVGDPSLKLNEIGIPESIAKRLQVSEHLNQCNKERLVTSFVPTLLDNKEM 375
Query: 433 KIREKEPIVW----------EILQEVMRGHPVLLNRAPTLHRLGIQAFQ-PILVEGRAIC 481
+R + +V +I + +M G VL+NR P++H+ + A IL +
Sbjct: 376 HVRRGDRLVAIQVNDLQTGDKIFRSLMDGDTVLMNRPPSIHQHSLIAMTVRILPTTSVVS 435
Query: 482 LHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSHTNLLSPAIGDPISVPTQDMLI 541
L+P+ C F DFDGD + +VP S++A+ E L+ L++ G + QD L
Sbjct: 436 LNPICCLPFRGDFDGDCLHGYVPQSIQAKVELDELVALDKQLINRQNGRNLLSLGQDSLT 495
Query: 542 GLYVL 546
Y++
Sbjct: 496 AAYLV 500
>At2g40030 putative DNA-directed RNA polymerase II subunit
Length = 888
Score = 75.5 bits (184), Expect = 9e-14
Identities = 84/308 (27%), Positives = 129/308 (41%), Gaps = 28/308 (9%)
Query: 262 NIEPEWMVLSLLPVLPPELRPIIQIDGGKLMSSDINELYRRVIYRNNTLIDLLTTSRS-- 319
+I E +L LPV P L DG MS D + + + + + + + +SRS
Sbjct: 151 HIPQEGYILEYLPVPPNCLSVPEASDGFSTMSVDPSRIELKDVLKK---VIAIKSSRSGE 207
Query: 320 TPGELVMCQEKLVQEAVDTLLD-----NGIRGQPMRDGHNKVYKSFSDIIEGKEGRFRET 374
T E + + VDT L R MR G +K+ S S + + R
Sbjct: 208 TNFESHKAEASEMFRVVDTYLQVRGTAKAARNIDMRYGVSKISDSSSS--KAWTEKMRTL 265
Query: 375 LLGKRVDYSGRSVIVVGPSLSLHRCGLPREIAIELFQTFLI----RGLIRKHFASNI--- 427
+ K +S RSVI ++ G+P EIA + + RG ++K +
Sbjct: 266 FIRKGSGFSSRSVITGDAYRHVNEVGIPIEIAQRITFEERVSVHNRGYLQKLVDDKLCLS 325
Query: 428 ---GVAKSKIREKEPIVWEIL------QEVMRGHPVLLNRAPTLHRLGIQAFQPILVEGR 478
G +R+ E+ + VM G V +NR PT H+ +QA + + E
Sbjct: 326 YTQGSTTYSLRDGSKGHTELKPGQVVHRRVMDGDVVFINRPPTTHKHSLQALRVYVHEDN 385
Query: 479 AICLHPLVCKGFNADFDGDQMAVHVPLSLEAQAEARLLMFSHTNLLSPAIGDPISVPTQD 538
+ ++PL+C +ADFDGD + + P SL A+AE L LLS G I D
Sbjct: 386 TVKINPLMCSPLSADFDGDCVHLFYPQSLSAKAEVMELFSVEKQLLSSHTGQLILQMGSD 445
Query: 539 MLIGLYVL 546
L+ L V+
Sbjct: 446 SLLSLRVM 453
>At1g70070 Similar to Synechocystis antiviral protein (At1g70070)
Length = 1171
Score = 32.7 bits (73), Expect = 0.69
Identities = 52/218 (23%), Positives = 90/218 (40%), Gaps = 39/218 (17%)
Query: 159 RLRGSFEYEIQSWKYSIPLFFATQGFDTF--RNREISSGAGAIREQ---LVDLDLRIIMD 213
RL GS E SW ++P+ + D + E + A +EQ + L ++
Sbjct: 879 RLEGSLE--TWSWSLNVPVLSSLSDEDEVLHMSEEYDNAAQKYKEQRSKISRLKKKMSRS 936
Query: 214 SSLVEWKELGEEGSADNENENEWEDRKVGRRKNFLVRRMELV-----KHFIR-TNIEPEW 267
E+K++ E N N + +++ R L+ R+E + K F+R +N+ E
Sbjct: 937 EGFREYKKILE-----NANLTVEKMKRLKARSRRLINRLEQIEPSGWKDFMRISNVIHES 991
Query: 268 MVLSLLPVLPPELRPIIQIDGGKLMSSDINELYRRVIYRNNTLIDLLTTSRSTPGELVMC 327
L + L L G+ NEL+ ++ RN L+DL P +L
Sbjct: 992 RALDINTHLIFPLGETAAAIRGE------NELWLAMVLRNKALVDL------KPPQLAGV 1039
Query: 328 QEKLVQEAVDTLLDNGIRGQPMRDGHNKVYKSFSDIIE 365
LV E GI+ +P RD +N +Y+ +++
Sbjct: 1040 CASLVSE--------GIKVRPWRD-NNYIYEPSDTVVD 1068
>At3g55780 beta-1,3-glucanase - like protein
Length = 429
Score = 31.6 bits (70), Expect = 1.5
Identities = 36/114 (31%), Positives = 51/114 (44%), Gaps = 6/114 (5%)
Query: 269 VLSLLPVLPPELRPIIQIDGGKLMSS-DINELYRRVIYRNNTLIDLLTTSRS---TPGEL 324
V+S + V+ E P+I + G S D +E+ ++Y L LLT RS TP
Sbjct: 256 VISSMAVMGHENLPVIVAETGWPSSGIDASEVDATLLYSEMFLKALLTHLRSGCGTPLRK 315
Query: 325 VMCQEKLVQEAVDTLLDNGIRGQPMRDGHNKVYKSFSDIIEG-KEGRFRETLLG 377
E + E V+ GIR + HN K D +G K RF+E L+G
Sbjct: 316 EGVSEVYIFELVEKDAKQGIRNWGLLH-HNMTSKYSFDFSDGGKVRRFKEILVG 368
>At1g04890 hypothetical protein
Length = 494
Score = 31.2 bits (69), Expect = 2.0
Identities = 44/186 (23%), Positives = 74/186 (39%), Gaps = 31/186 (16%)
Query: 298 ELYRRVIYRNNTLIDLLTTSRSTPG----ELVMCQEK---LVQEAVDTLLDNGIRGQPMR 350
E YR+++ L L ++ P + CQE+ LVQE T+LD R + R
Sbjct: 290 EAYRQLLEETEELECSLIKEKNVPEPEHKQNKDCQERRALLVQELDGTVLDMPYREEGNR 349
Query: 351 DGHNKVYKSFSDIIE----------------GKEGRFRETLLGK-RVDYSGRSVIVVGPS 393
D + +YKS S++ K+ E+ +GK + S+IV G +
Sbjct: 350 DKNRDLYKSDSEVAYSRVRDVYMVKDETENISKKKNLEESSVGKPKESLDENSIIVSGIA 409
Query: 394 LSLHRCGLPREIAIELFQTFLIRGLIRKHFASNIGVAKSKIREKEPIVWEILQEVMRGHP 453
L PR+ ++ + R+ S + + KI + ++ E L+ V
Sbjct: 410 RKLPPLCRPRKKSLSSSGS-------RRKSMSAVDYERLKIENEVELLRERLKAVQEERE 462
Query: 454 VLLNRA 459
L RA
Sbjct: 463 ELTRRA 468
>At3g23010 disease resistance protein, putative
Length = 595
Score = 30.8 bits (68), Expect = 2.6
Identities = 18/58 (31%), Positives = 34/58 (58%), Gaps = 4/58 (6%)
Query: 486 VCKGFNA-DFDGDQMAVHVPLSLEAQAEARLLMFS---HTNLLSPAIGDPISVPTQDM 539
+ +GFNA DF G++ + H+P S+ +E RLL S T + P++ + ++ + D+
Sbjct: 426 IFEGFNAIDFSGNRFSGHIPGSIGLLSELRLLNLSGNAFTGNIPPSLANITNLESLDL 483
>At3g57300 helicase-like protein
Length = 1507
Score = 29.3 bits (64), Expect = 7.7
Identities = 16/71 (22%), Positives = 37/71 (51%), Gaps = 5/71 (7%)
Query: 199 IREQLVDLDLRIIM---DSSLVEWKELGEEGSAD--NENENEWEDRKVGRRKNFLVRRME 253
IR + + D+ + D + E ++ E+ +A+ + + E ++ +R NFL+++ E
Sbjct: 404 IRTRKISRDMLLFWKRYDKQMAEERKKQEKEAAEAFKREQEQRESKRQQQRLNFLIKQTE 463
Query: 254 LVKHFIRTNIE 264
L HF++ +
Sbjct: 464 LYSHFMQNKTD 474
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.141 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,332,118
Number of Sequences: 26719
Number of extensions: 745078
Number of successful extensions: 1647
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1625
Number of HSP's gapped (non-prelim): 17
length of query: 686
length of database: 11,318,596
effective HSP length: 106
effective length of query: 580
effective length of database: 8,486,382
effective search space: 4922101560
effective search space used: 4922101560
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC126018.19


