

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.13 - phase: 0
(320 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
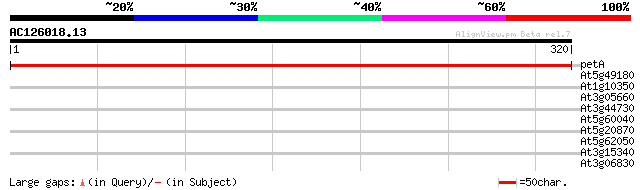
Score E
Sequences producing significant alignments: (bits) Value
petA -chloroplast genome- cytochrome f 583 e-167
At5g49180 pectin methylesterase 34 0.093
At1g10350 putative heat-shock protein 30 1.7
At3g05660 putative disease resistance protein 29 3.0
At3g44730 kinesin-like protein heavy chain (KP1) 29 3.9
At5g60040 DNA-directed RNA polymerase - like protein 28 5.1
At5g20870 beta-1,3-glucanase - like protein 28 6.6
At5g62050 AtOXA1 28 8.7
At3g15340 hypothetical protein 28 8.7
At3g06830 pectin methylesterase like protein 28 8.7
>petA -chloroplast genome- cytochrome f
Length = 320
Score = 583 bits (1503), Expect = e-167
Identities = 289/320 (90%), Positives = 307/320 (95%)
Query: 1 MQTRNAFSWIKEEITRSISVLLMIYIITRAPISNAYPIFAQQGYENPREATGRIVCANCH 60
MQTRN FSWI+EEITRSISV L+IYIIT A IS+AYPIFAQQ YENPREATGRIVCANCH
Sbjct: 1 MQTRNTFSWIREEITRSISVSLIIYIITWASISSAYPIFAQQNYENPREATGRIVCANCH 60
Query: 61 LANKPVDIEVPQAVLPDTVFEAVVRIPYDMQVKQVLANGKKGALNVGAVLILPEGFELAP 120
LANKPVDIEVPQ VLPDTVFEAVV+IPYDMQ+KQVLANGKKGALNVGAVLILPEGFELAP
Sbjct: 61 LANKPVDIEVPQTVLPDTVFEAVVKIPYDMQLKQVLANGKKGALNVGAVLILPEGFELAP 120
Query: 121 PDRISPEIKEKIGNLSFQSYRPTKKNILVVGPVPGKKYSEITFPILSPDPATKRDVHFLK 180
PDRISPE+KEKIGNLSFQ+YRP KKNILV+GPVPG+KYSEITFPIL+PDPAT +DVHFLK
Sbjct: 121 PDRISPEMKEKIGNLSFQNYRPNKKNILVIGPVPGQKYSEITFPILAPDPATNKDVHFLK 180
Query: 181 YPIYVGGNRGRGQIYPDGSKSNNNVYNATATGIVNKIIRKEKGGYEITIVDASDGREVID 240
YPIYVGGNRGRGQIYPDGSKSNN VYNATA GI++KI+RKEKGGYEITIVDAS+GREVID
Sbjct: 181 YPIYVGGNRGRGQIYPDGSKSNNTVYNATAGGIISKILRKEKGGYEITIVDASNGREVID 240
Query: 241 IIPPGPELLVSEGESMKLDQPLTSNPNVGGFGQGDAEIVLQDPLRVQGLLFFLASIILAQ 300
IIP G ELLVSEGES+KLDQPLTSNPNVGGFGQGDAEIVLQDPLRVQGLLFFL S++LAQ
Sbjct: 241 IIPRGLELLVSEGESIKLDQPLTSNPNVGGFGQGDAEIVLQDPLRVQGLLFFLGSVVLAQ 300
Query: 301 IFLVLKKKQFEKVQLSEMNF 320
IFLVLKKKQFEKVQLSEMNF
Sbjct: 301 IFLVLKKKQFEKVQLSEMNF 320
>At5g49180 pectin methylesterase
Length = 571
Score = 34.3 bits (77), Expect = 0.093
Identities = 19/53 (35%), Positives = 28/53 (51%), Gaps = 1/53 (1%)
Query: 18 ISVLLMIYIITRAPISNAYPIFAQQGYENPREATGRIVCANCHLANKPVDIEV 70
I VL I+ R P+ + + QG + RE+TG +V NCH+ +P I V
Sbjct: 416 IVVLQNCNIVVRKPMKSQSCMITAQGRSDKRESTG-LVLQNCHITGEPAYIPV 467
>At1g10350 putative heat-shock protein
Length = 349
Score = 30.0 bits (66), Expect = 1.7
Identities = 40/168 (23%), Positives = 68/168 (39%), Gaps = 41/168 (24%)
Query: 120 PPDRISPEIKEKIGNLSFQSYRPTKKNILVVGPVP---GKKYSEITFPILSPDPATKRDV 176
P +R +P I+ K+ + Y+ KK + + VP GK P T +++
Sbjct: 166 PANRKAPAIESKLACTLEELYKGAKKKMRISRVVPDDFGK-------------PKTVQEI 212
Query: 177 HFLKYPIYVGGNRGRGQIYPDGSKSNNNVYNATATGIVNK---IIRKEKGG-----YEIT 228
LK I G +G +P+ V A +V++ + K G +++
Sbjct: 213 --LKIDIKPGWKKGTKITFPEKGNQEPGVTPADLIFVVDEKPHSVFKRDGNDLILEKKVS 270
Query: 229 IVDAS----------DGRE----VIDIIPPGPELLVSEGESMKLDQPL 262
++DA DGR V+DI+ PG E+++ E M PL
Sbjct: 271 LIDALTGLTISVTTLDGRSLTIPVLDIVKPGQEIVI-PNEGMPTKDPL 317
>At3g05660 putative disease resistance protein
Length = 883
Score = 29.3 bits (64), Expect = 3.0
Identities = 15/34 (44%), Positives = 23/34 (67%), Gaps = 1/34 (2%)
Query: 122 DRISPEIKEKIGNLSFQSYRPTKKNILVVGPVPG 155
+++S EI +++GNLS+ +Y N L VG VPG
Sbjct: 745 NKLSGEIPQELGNLSYLAYMNFSHNQL-VGQVPG 777
>At3g44730 kinesin-like protein heavy chain (KP1)
Length = 1087
Score = 28.9 bits (63), Expect = 3.9
Identities = 36/174 (20%), Positives = 67/174 (37%), Gaps = 22/174 (12%)
Query: 123 RISPEIKEKIGNLSFQSYRPTKKNILVVGPVPGKKYSEITFPI--------------LSP 168
R+ P +E+ S Y NI++ P +K + F +
Sbjct: 383 RVRPFFQEQKDMQSTVDYIGENGNIIINNPFKQEKDARKIFSFNKVFGQTVSQEQIYIDT 442
Query: 169 DPATKRDVHFLKYPIYVGGNRGRGQIYPDGSKSNNNVYNATATGIVNKIIRK--EKGGYE 226
P + + I+ G G G+ Y + S ++ T G+ + +R +
Sbjct: 443 QPVIRSVLDGFNVCIFAYGQTGSGKTY---TMSGPDLMTETTWGVNYRALRDLFQLSNAR 499
Query: 227 ITIVDASDGREVIDIIPPGP-ELLVSEGESMKLDQPLTSNPNVGGFGQGDAEIV 279
+V G ++I+I +LLVS+G S +LD + +N + G DA ++
Sbjct: 500 THVVTYEIGVQMIEIYNEQVRDLLVSDGSSRRLD--IRNNSQLNGLNVPDANLI 551
>At5g60040 DNA-directed RNA polymerase - like protein
Length = 1328
Score = 28.5 bits (62), Expect = 5.1
Identities = 18/60 (30%), Positives = 29/60 (48%), Gaps = 9/60 (15%)
Query: 149 VVGPVPGKKYSEITFPILS------PDPATKRDVHFLKYPIYVGGNR---GRGQIYPDGS 199
V+ P P K +E+ PIL P+ ++ ++ L+ + G N+ R YPDGS
Sbjct: 369 VISPDPNLKITEVGIPILMAQILTFPECVSRHNIEKLRQCVRNGPNKYPGARNVRYPDGS 428
>At5g20870 beta-1,3-glucanase - like protein
Length = 501
Score = 28.1 bits (61), Expect = 6.6
Identities = 24/78 (30%), Positives = 36/78 (45%), Gaps = 12/78 (15%)
Query: 159 SEITFPI-----LSPDPATKRDVHFLKYPIYVGGNRGRGQIYPDGSKSNNNVYNATATGI 213
S ITF I L+ DP R+ F GG G + DGS S NV++A +
Sbjct: 202 SPITFNIYPFLSLNADPNFPREYAFFPN----GGGGGGAKPVVDGSISYTNVFDANFDTL 257
Query: 214 VNKIIRKEKGGYEITIVD 231
V+ + EK G++ ++
Sbjct: 258 VSAL---EKNGFDANKIE 272
>At5g62050 AtOXA1
Length = 429
Score = 27.7 bits (60), Expect = 8.7
Identities = 16/56 (28%), Positives = 27/56 (47%), Gaps = 1/56 (1%)
Query: 40 AQQGYE-NPREATGRIVCANCHLANKPVDIEVPQAVLPDTVFEAVVRIPYDMQVKQ 94
AQ+G E NP T + VC L P+ + PQA+ + + + Y + +K+
Sbjct: 290 AQEGMEGNPMAGTVKTVCRVFALLTVPMTMSFPQAIFCYWITSNLFSLMYGLVIKR 345
>At3g15340 hypothetical protein
Length = 478
Score = 27.7 bits (60), Expect = 8.7
Identities = 27/87 (31%), Positives = 41/87 (47%), Gaps = 18/87 (20%)
Query: 106 VGAVLILPEGFELAPPDRISPEIKEKIGNLSFQSYRPTKKNILVVGPVPGKKYSEITFPI 165
+GA +IL EGFE+ PP PE+ + + S QS + ++ V G +S +
Sbjct: 1 MGAQIILSEGFEVVPP----PEMNDLVLFGSNQSV--SSCDVSTVTTEDGTVFSGDS--- 51
Query: 166 LSPDPATKRD--------VHFLKYPIY 184
SP AT+ D +F+K P Y
Sbjct: 52 -SPGGATEEDFPEEKPFSFYFVKQPAY 77
>At3g06830 pectin methylesterase like protein
Length = 568
Score = 27.7 bits (60), Expect = 8.7
Identities = 15/43 (34%), Positives = 20/43 (45%), Gaps = 1/43 (2%)
Query: 26 IITRAPISNAYPIFAQQGYENPREATGRIVCANCHLANKPVDI 68
I+ R P + QG N RE+TG +V CH+ P I
Sbjct: 422 IVVRKPNKGQTCMVTAQGRSNVRESTG-LVLHGCHITGDPAYI 463
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.140 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,364,401
Number of Sequences: 26719
Number of extensions: 335857
Number of successful extensions: 707
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 706
Number of HSP's gapped (non-prelim): 11
length of query: 320
length of database: 11,318,596
effective HSP length: 99
effective length of query: 221
effective length of database: 8,673,415
effective search space: 1916824715
effective search space used: 1916824715
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC126018.13


