

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126016.2 - phase: 0
(327 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
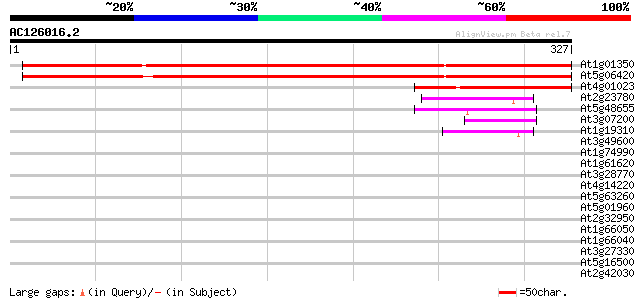
Score E
Sequences producing significant alignments: (bits) Value
At1g01350 unknown protein (At1g01350/F6F3_27) 477 e-135
At5g06420 unknown protein 473 e-134
At4g01023 unknown protein 110 8e-25
At2g23780 putative RING zinc finger protein 45 7e-05
At5g48655 RING zinc finger like protein 43 3e-04
At3g07200 putative RING zinc finger protein 42 3e-04
At1g19310 unknown protein 41 8e-04
At3g49600 putative protein 40 0.001
At1g74990 RING zinc finger like protein 40 0.001
At1g61620 unknown protein 40 0.001
At3g28770 hypothetical protein 40 0.002
At4g14220 RING-H2 finger protein RHF1a 39 0.004
At5g63260 unknown protein 39 0.005
At5g01960 unknown protein 38 0.007
At2g32950 photomorphogenesis repressor (COP1) 38 0.007
At1g66050 hypothetical protein 38 0.007
At1g66040 hypothetical protein 38 0.007
At3g27330 hypothetical protein 38 0.009
At5g16500 protein kinase-like protein 37 0.015
At2g42030 putative RING zinc finger protein 37 0.015
>At1g01350 unknown protein (At1g01350/F6F3_27)
Length = 343
Score = 477 bits (1228), Expect = e-135
Identities = 223/320 (69%), Positives = 266/320 (82%), Gaps = 3/320 (0%)
Query: 8 STENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFS 67
+ + Q++QVC+FF+KP KN+RKRTI+ ++ + DS +E S++ KK K D+KL+FS
Sbjct: 20 AAQETQSQQVCTFFKKPTKSKNIRKRTIDADEEDGDSKSESSILQNLKKVAKPDSKLYFS 79
Query: 68 TGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALK 127
+G SKSS + + E+ FH++SSKEIQVQ+DS ATATLETETDF++DARAIRER LK
Sbjct: 80 SGPSKSSTTT--SGAPERSVFHYDSSKEIQVQNDSGATATLETETDFNQDARAIRERVLK 137
Query: 128 QATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSA 187
+A E+LKG + D KLYKGI+ YTDHKAGFRREQTI+SEKAGGSHGPLRASAHIRVSA
Sbjct: 138 KADEALKGNKKKASDEKLYKGIHGYTDHKAGFRREQTISSEKAGGSHGPLRASAHIRVSA 197
Query: 188 RFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDA 247
RFDYQPDICKDYKETGYCGYGDSCKF+HDRGDYK GWQ+EKEW+EAEK RK A G +
Sbjct: 198 RFDYQPDICKDYKETGYCGYGDSCKFLHDRGDYKPGWQIEKEWEEAEKVRKRNKAMGVED 257
Query: 248 EEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
E++ A D D+DE+ALPFACFICR PFVDPV TKCKHYFCEHCALKHH KNKKCFVCNQ
Sbjct: 258 EDDEAD-KDSDEDENALPFACFICREPFVDPVVTKCKHYFCEHCALKHHTKNKKCFVCNQ 316
Query: 308 PTLGIFNVAHEIRKKMAEDK 327
PT+GIFN AHEI+K+MAE++
Sbjct: 317 PTMGIFNAAHEIKKRMAEER 336
>At5g06420 unknown protein
Length = 378
Score = 473 bits (1217), Expect = e-134
Identities = 219/320 (68%), Positives = 265/320 (82%), Gaps = 6/320 (1%)
Query: 8 STENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFS 67
++ NQ+++QVC+FF+KP KN+RKRTI+ ++ + DS +E S++ KK K D+ L+FS
Sbjct: 58 NSNNQESQQVCTFFKKPTKSKNIRKRTIDADEEDGDSKSESSILQNLKKVAKPDSNLYFS 117
Query: 68 TGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALK 127
+G S ++ A E+P FH++SSKEIQVQ+DS ATATLETETDF++DARAIRER LK
Sbjct: 118 SGPSTRTSGAP-----ERPVFHYDSSKEIQVQNDSGATATLETETDFNQDARAIRERVLK 172
Query: 128 QATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSA 187
+A +LKG + D KLYKGI+ YTDHKAGFRREQTI+SEKAGGSHGPLRASAHIRVSA
Sbjct: 173 KADHALKGNKKKASDEKLYKGIHGYTDHKAGFRREQTISSEKAGGSHGPLRASAHIRVSA 232
Query: 188 RFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDA 247
RFDYQPDICKDYKETGYCGYGDSCKF+HDRGDYK GWQ+EKEW+EAEK RK A G +
Sbjct: 233 RFDYQPDICKDYKETGYCGYGDSCKFLHDRGDYKPGWQIEKEWEEAEKVRKRNKAMGVED 292
Query: 248 EEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
+++ A D D+DE+ALPFACFICR PF+DPV TKCKHYFCEHCALKHH KNKKCFVCNQ
Sbjct: 293 DDDEAD-KDSDEDENALPFACFICREPFLDPVVTKCKHYFCEHCALKHHTKNKKCFVCNQ 351
Query: 308 PTLGIFNVAHEIRKKMAEDK 327
PT+GIFN AHEI+K+MAE++
Sbjct: 352 PTMGIFNAAHEIKKRMAEER 371
>At4g01023 unknown protein
Length = 255
Score = 110 bits (276), Expect = 8e-25
Identities = 47/91 (51%), Positives = 65/91 (70%), Gaps = 2/91 (2%)
Query: 237 RKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHH 296
+K++ T E + +EDDD ALP AC IC+NPF+DPV T C HYFC+ CALKHH
Sbjct: 76 KKIKKETARKLETDFVLTGEEDDD--ALPLACSICQNPFLDPVVTNCNHYFCDKCALKHH 133
Query: 297 AKNKKCFVCNQPTLGIFNVAHEIRKKMAEDK 327
+N CFVCN+PTLG+F+ A EI++++ E++
Sbjct: 134 TENDTCFVCNEPTLGLFDTAVEIKERIEEER 164
Score = 30.0 bits (66), Expect = 1.8
Identities = 43/171 (25%), Positives = 69/171 (40%), Gaps = 28/171 (16%)
Query: 1 MEDS-EAKSTENQ-QTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNT 58
M DS EAK++E + Q + + ++ KN++KR ++ ++E DSNNE S + +K
Sbjct: 20 MSDSDEAKTSEEESQLAEQETETQEGEMTKNLKKRDLDAVEDE-DSNNESSRLEKLEKKI 78
Query: 59 KAD--NKL---FFSTGSSKSSASAKPNEESEKPSF--------HFESSKEIQVQHDSKAT 105
K + KL F TG A + P H+ K H T
Sbjct: 79 KKETARKLETDFVLTGEEDDDALPLACSICQNPFLDPVVTNCNHYFCDKCALKHHTENDT 138
Query: 106 ---------ATLETETDFSRDARAIRERA---LKQATESLKGKSTSSEDVK 144
+T + RE+A +K+ T L+ ST ++D K
Sbjct: 139 CFVCNEPTLGLFDTAVEIKERIEEEREKARAMVKEVTAMLEKASTMADDAK 189
>At2g23780 putative RING zinc finger protein
Length = 227
Score = 44.7 bits (104), Expect = 7e-05
Identities = 22/68 (32%), Positives = 33/68 (48%), Gaps = 3/68 (4%)
Query: 241 LATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCA---LKHHA 297
+ GE + S ++ D ++ F C IC DP+ T C H FC C L HH+
Sbjct: 1 MVNGESSTSTSYSDSNNDTNDQGGDFECNICFELAQDPIVTLCGHLFCWPCLYRWLHHHS 60
Query: 298 KNKKCFVC 305
+++C VC
Sbjct: 61 HSQECPVC 68
>At5g48655 RING zinc finger like protein
Length = 203
Score = 42.7 bits (99), Expect = 3e-04
Identities = 25/85 (29%), Positives = 39/85 (45%), Gaps = 14/85 (16%)
Query: 237 RKMRLATGEDAEE-EGASLNDEDDDEDALP-------------FACFICRNPFVDPVSTK 282
++ R+ + E + E AS+NDE + + F C IC PF + +STK
Sbjct: 103 KRRRIPSSESVIDCEHASVNDEVNMSSRVSRSKAPAPPPEEPKFTCPICMCPFTEEMSTK 162
Query: 283 CKHYFCEHCALKHHAKNKKCFVCNQ 307
C H FC+ C ++ KC C +
Sbjct: 163 CGHIFCKGCIKMAISRQGKCPTCRK 187
>At3g07200 putative RING zinc finger protein
Length = 182
Score = 42.4 bits (98), Expect = 3e-04
Identities = 18/42 (42%), Positives = 22/42 (51%)
Query: 266 FACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
F+C IC PF VSTKC H FC+ C + KC C +
Sbjct: 125 FSCPICLCPFTQEVSTKCGHIFCKKCIKNALSLQAKCPTCRK 166
>At1g19310 unknown protein
Length = 226
Score = 41.2 bits (95), Expect = 8e-04
Identities = 20/56 (35%), Positives = 30/56 (52%), Gaps = 3/56 (5%)
Query: 253 SLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKH---HAKNKKCFVC 305
S ++ ++ D+ F C IC + DP+ T C H FC C K H+++K C VC
Sbjct: 8 SSGNDTNNNDSSNFECNICLDLAQDPIVTLCGHLFCWPCLYKWLHLHSQSKDCPVC 63
>At3g49600 putative protein
Length = 1672
Score = 40.4 bits (93), Expect = 0.001
Identities = 52/258 (20%), Positives = 109/258 (42%), Gaps = 34/258 (13%)
Query: 22 RKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFSTGSSKSSASAKPNE 81
+K RK + + D+E+DS++++ +KK TK K S S + ++
Sbjct: 242 KKAKGRKQESESDSSSSDSESDSDSDDGKKRGRKKPTKTTKKRSRRKRSVSSESEEVESD 301
Query: 82 ESEK---------PSFHFESSKEIQVQHDSKATATLET-ETDFSRDARAIRERALKQATE 131
+S+K PS + SKE++ +HD ++ A + ++D S ++ L++ E
Sbjct: 302 DSKKLRKSHKKSLPS-NRSGSKELRDKHDEQSRAGRKRHDSDVSEPESEDNKQPLRKKEE 360
Query: 132 SLKG---KSTSSEDVKLYKGINNYTDH-KAGFRREQTIASEKAGGSHGPLRASAHIRVSA 187
+ +G + EDV+ DH K + R+ A+ + S + +R
Sbjct: 361 AYRGGQKQKRDDEDVE--------ADHLKDRYTRDDKKAARDSDDSEIEYQNKKQLRSKV 412
Query: 188 RFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKS---GWQMEKEWDEAEKARKMRLATG 244
Y + + KE + H + Y+S G ++ ++ D++E + R
Sbjct: 413 EV-YSAGMSQKRKE-------EEDVTKHGKDKYRSDSRGKEVARDSDDSEAEYENRKKLK 464
Query: 245 EDAEEEGASLNDEDDDED 262
++ + G E+D+++
Sbjct: 465 NESYQRGRKHKREEDEDN 482
Score = 33.1 bits (74), Expect = 0.21
Identities = 42/261 (16%), Positives = 109/261 (41%), Gaps = 16/261 (6%)
Query: 3 DSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKK---NTK 59
DSE+ S + ++ K +++ RKR++ +E E +S++ + L KK + +
Sbjct: 259 DSESDSDSDDGKKRGRKKPTKTTKKRSRRKRSVSSESEEVESDDSKKLRKSHKKSLPSNR 318
Query: 60 ADNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDAR 119
+ +K +S A K ++ ++ + ++ + ++ + D +A
Sbjct: 319 SGSKELRDKHDEQSRAGRKRHDSDVSEPESEDNKQPLRKKEEAYRGGQKQKRDDEDVEAD 378
Query: 120 AIRERALKQATESLKGKSTSSEDVKLYKGINNYTD-HKAGFRREQTIASEKAGGSHGPLR 178
+++R + ++ + S + + K + + + + AG +++ E+ HG +
Sbjct: 379 HLKDRYTRDDKKAARDSDDSEIEYQNKKQLRSKVEVYSAGMSQKR--KEEEDVTKHGKDK 436
Query: 179 ASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARK 238
+ R ++ +D + Y + K ++ Y+ G + ++E DE
Sbjct: 437 YRSDSR-------GKEVARD-SDDSEAEYENRKKLKNE--SYQRGRKHKREEDEDNDNHG 486
Query: 239 MRLATGEDAEEEGASLNDEDD 259
G+DA + ++ ++DD
Sbjct: 487 RDRYRGDDAVKRYGTIKEDDD 507
>At1g74990 RING zinc finger like protein
Length = 137
Score = 40.4 bits (93), Expect = 0.001
Identities = 21/54 (38%), Positives = 28/54 (50%), Gaps = 5/54 (9%)
Query: 255 NDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALK---HHAKNKKCFVC 305
N+EDD + F C IC +P+ T C H FC C K +H+K+ C VC
Sbjct: 8 NEEDDASNN--FGCNICLELAREPIVTLCGHLFCWPCLYKWLHYHSKSNHCPVC 59
>At1g61620 unknown protein
Length = 310
Score = 40.4 bits (93), Expect = 0.001
Identities = 21/64 (32%), Positives = 31/64 (47%), Gaps = 8/64 (12%)
Query: 265 PF-ACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQPTLGIFNVAHEIRKKM 323
PF AC +C PF+DP+ H FC C L +CF+ + + AH +KK
Sbjct: 39 PFDACSLCLKPFIDPMCCHKGHVFCRECIL-------ECFLAQKKDIQRRLAAHSSQKKQ 91
Query: 324 AEDK 327
+D+
Sbjct: 92 DKDE 95
Score = 37.7 bits (86), Expect = 0.009
Identities = 19/70 (27%), Positives = 31/70 (44%), Gaps = 4/70 (5%)
Query: 243 TGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVS----TKCKHYFCEHCALKHHAK 298
T +++EEE + C C+ + +S + C H FC+ CA K
Sbjct: 196 TEDNSEEEETKTKSASSSSYDKSYICPSCKVTLTNTMSLVALSSCGHVFCKKCAEKFMPV 255
Query: 299 NKKCFVCNQP 308
+K C VC++P
Sbjct: 256 DKVCLVCDKP 265
>At3g28770 hypothetical protein
Length = 2081
Score = 40.0 bits (92), Expect = 0.002
Identities = 53/269 (19%), Positives = 104/269 (37%), Gaps = 21/269 (7%)
Query: 2 EDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTI------ENEDNENDSNNEESLMHVQK 55
++S K E + E V + +K + K ++ EN+DN+ +E+S ++
Sbjct: 952 KNSNMKKKEEDKKEYVNNELKKQEDNKKETTKSENSKLKEENKDNKEKKESEDSASKNRE 1011
Query: 56 KNTKADNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFS 115
K + K + K ++ + EK S +S KE + D KA E ET
Sbjct: 1012 KKEYEEKKSKTKEEAKKEKKKSQDKKREEKDSEERKSKKEKEESRDLKAKKK-EEETKEK 1070
Query: 116 RDARAIRERALKQATESLKGKSTSSEDVKLYKGINNYT------DHKAGFRREQTIASEK 169
+++ + + + E KS E+ K K + + + K + + S K
Sbjct: 1071 KESENHKSKKKEDKKEHEDNKSMKKEEDKKEKKKHEESKSRKKEEDKKDMEKLEDQNSNK 1130
Query: 170 AGGSHGPLRASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKE 229
+ S H+++ + + + K+ +E +S K + D K + +
Sbjct: 1131 KKEDKNEKKKSQHVKLVKKESDKKE-KKENEEKSETKEIESSKSQKNEVDKKEKKSSKDQ 1189
Query: 230 WDEAEKARKMRLATGEDAEEEGASLNDED 258
+ EK K ++EE+ N+ED
Sbjct: 1190 QKKKEKEMK-------ESEEKKLKKNEED 1211
Score = 36.6 bits (83), Expect = 0.019
Identities = 43/193 (22%), Positives = 75/193 (38%), Gaps = 23/193 (11%)
Query: 1 MEDSEAKSTENQQTEQVCSFFRKPV-----NRKNMRKRT-IENEDNENDSNNEESLMHVQ 54
++ E KS+++QQ ++ N ++ +K+T +E + ++ E++
Sbjct: 1178 VDKKEKKSSKDQQKKKEKEMKESEEKKLKKNEEDRKKQTSVEENKKQKETKKEKNKPKDD 1237
Query: 55 KKNT-KADNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETD 113
KKNT K S S A + ++ + ES EI +Q DS+A + +++ D
Sbjct: 1238 KKNTTKQSGGKKESMESESKEAENQQKSQATTQADSDESKNEILMQADSQADSHSDSQAD 1297
Query: 114 FSRDARAIRERALKQAT----------------ESLKGKSTSSEDVKLYKGINNYTDHKA 157
I +A QAT E+ K K T E K N T
Sbjct: 1298 SDESKNEILMQADSQATTQRNNEEDRKKQTSVAENKKQKETKEEKNKPKDDKKNTTKQSG 1357
Query: 158 GFRREQTIASEKA 170
G + S++A
Sbjct: 1358 GKKESMESESKEA 1370
Score = 36.2 bits (82), Expect = 0.025
Identities = 46/197 (23%), Positives = 71/197 (35%), Gaps = 37/197 (18%)
Query: 3 DSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADN 62
DS+A S E++ + + + R N R + EN E ++KN D+
Sbjct: 1293 DSQADSDESKNEILMQADSQATTQRNNEEDRKKQTSVAENKKQKETK----EEKNKPKDD 1348
Query: 63 KLFFSTGSSKSSASAKPNEESEKPSFHF-------------ESSKEIQVQHDSKATATLE 109
K ++K S K + ESE ES EI +Q DS+A + +
Sbjct: 1349 K----KNTTKQSGGKKESMESESKEAENQQKSQATTQADSDESKNEILMQADSQADSHSD 1404
Query: 110 TETDFSRDARAIRERALKQAT----------------ESLKGKSTSSEDVKLYKGINNYT 153
++ D I +A QAT E+ K K T E K N T
Sbjct: 1405 SQADSDESKNEILMQADSQATTQRNNEEDRKKQTSVAENKKQKETKEEKNKPKDDKKNTT 1464
Query: 154 DHKAGFRREQTIASEKA 170
+ G + S++A
Sbjct: 1465 EQSGGKKESMESESKEA 1481
Score = 35.4 bits (80), Expect = 0.043
Identities = 51/271 (18%), Positives = 98/271 (35%), Gaps = 53/271 (19%)
Query: 26 NRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFSTGSSKSSASAKPNEESEK 85
N + +++ + D+ ND+N E K++TK++ ++ + GSS+ K N +
Sbjct: 653 NADSNKEKEVHVGDSTNDNNMES------KEDTKSEVEVKKNDGSSEKGEEGKENNKDSM 706
Query: 86 PSFHFESSKEIQVQHDSKATATLETETDF----SRDARAI-----------------RER 124
E+ + D K+ + E S+D +++ E
Sbjct: 707 EDKKLENKESQTDSKDDKSVDDKQEEAQIYGGESKDDKSVEAKGKKKESKENKKTKTNEN 766
Query: 125 ALKQATESLKGKSTSSEDV--------KLYKGINNYTDHKAGFRREQTIASEKAGGSHGP 176
++ E+++G SE V K K + + K + A E++G +
Sbjct: 767 RVRNKEENVQGNKKESEKVEKGEKKESKDAKSVETKDNKKLSSTENRDEAKERSGEDNKE 826
Query: 177 LRASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKA 236
+ + DYQ K+ E G G + D D K +E + ++ E
Sbjct: 827 DKEESK-------DYQSVEAKEKNENG--GVDTNVGNKEDSKDLKDDRSVEVKANKEESM 877
Query: 237 RKMRLATGEDAEEEGASLNDEDDDEDALPFA 267
+K R E ND+ ++ FA
Sbjct: 878 KKKR---------EEVQRNDKSSTKEVRDFA 899
Score = 35.4 bits (80), Expect = 0.043
Identities = 35/153 (22%), Positives = 70/153 (44%), Gaps = 8/153 (5%)
Query: 2 EDSEAKSTENQQTEQVCSFFRKPVNRKNMRK----RTIENEDNENDSNNEESLMHVQKKN 57
E+ ++K E+++ + +K ++K +K ++ + E+++ D E +KK
Sbjct: 1074 ENHKSKKKEDKKEHEDNKSMKKEEDKKEKKKHEESKSRKKEEDKKDMEKLEDQNSNKKKE 1133
Query: 58 TKADNKLFFSTGSSKSSASAKPNEESEKPSF--HFESSKEIQVQHDSKAT-ATLETETDF 114
K + K K + K +E+E+ S ESSK + + D K ++ + +
Sbjct: 1134 DKNEKKKSQHVKLVKKESDKKEKKENEEKSETKEIESSKSQKNEVDKKEKKSSKDQQKKK 1193
Query: 115 SRDARAIRERALKQATESLKGKSTSSEDVKLYK 147
++ + E+ LK+ E K K TS E+ K K
Sbjct: 1194 EKEMKESEEKKLKKNEEDRK-KQTSVEENKKQK 1225
Score = 34.7 bits (78), Expect = 0.073
Identities = 53/266 (19%), Positives = 95/266 (34%), Gaps = 19/266 (7%)
Query: 1 MEDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDN------ENDSNNEESLMHVQ 54
+ED + + + E+ S K V +++ +K ENE+ E+ + + + +
Sbjct: 1123 LEDQNSNKKKEDKNEKKKSQHVKLVKKESDKKEKKENEEKSETKEIESSKSQKNEVDKKE 1182
Query: 55 KKNTKADNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQ--VQHDSKATATLETET 112
KK++K K K NEE K E +K+ + + +K + T
Sbjct: 1183 KKSSKDQQKKKEKEMKESEEKKLKKNEEDRKKQTSVEENKKQKETKKEKNKPKDDKKNTT 1242
Query: 113 DFSRDARAIRERALKQATESLKGKSTSSEDVKLYKG---------INNYTDHKAGFRREQ 163
S + E K+A K ++T+ D K ++++D +A +
Sbjct: 1243 KQSGGKKESMESESKEAENQQKSQATTQADSDESKNEILMQADSQADSHSDSQADSDESK 1302
Query: 164 TIASEKAGGSHGPLRASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSG 223
+A R + R + K+ KE D G K
Sbjct: 1303 NEILMQADSQATTQRNNEEDRKKQTSVAENKKQKETKEEKNKPKDDKKNTTKQSGGKKE- 1361
Query: 224 WQMEKEWDEAEKARKMRLATGEDAEE 249
ME E EAE +K + T D++E
Sbjct: 1362 -SMESESKEAENQQKSQATTQADSDE 1386
Score = 34.3 bits (77), Expect = 0.095
Identities = 36/164 (21%), Positives = 67/164 (39%), Gaps = 18/164 (10%)
Query: 1 MEDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIEN-------EDNENDSNNEESLMHV 53
M S E+ + V + ++K ++ T EN E+ EN + NEES+
Sbjct: 390 MNSENKGSGESTNDKMVNATTNDEDHKKENKEETHENNGESVKGENLENKAGNEESM--- 446
Query: 54 QKKNTKADNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKAT----ATLE 109
K +NK+ +S AK N ES K ES + +V + + T ++
Sbjct: 447 --KGENLENKVGNEELKGNASVEAKTNNESSKEEKREESQRSNEVYMNKETTKGENVNIQ 504
Query: 110 TET--DFSRDARAIRERALKQATESLKGKSTSSEDVKLYKGINN 151
E+ D ++D + +K ++ + S+++ +NN
Sbjct: 505 GESIGDSTKDNSLENKEDVKPKVDANESDGNSTKERHQEAQVNN 548
Score = 30.0 bits (66), Expect = 1.8
Identities = 33/148 (22%), Positives = 55/148 (36%), Gaps = 13/148 (8%)
Query: 3 DSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADN 62
DS+A S E++ + + + R N R + EN E ++KN D+
Sbjct: 1404 DSQADSDESKNEILMQADSQATTQRNNEEDRKKQTSVAENKKQKETK----EEKNKPKDD 1459
Query: 63 KLFFS--TGSSKSSASAKPNEESEKPSFHF-------ESSKEIQVQHDSKATATLETETD 113
K + +G K S ++ E + ES EI +Q DS+A ++ D
Sbjct: 1460 KKNTTEQSGGKKESMESESKEAENQQKSQATTQGESDESKNEILMQADSQADTHANSQGD 1519
Query: 114 FSRDARAIRERALKQATESLKGKSTSSE 141
I +A QA + +E
Sbjct: 1520 SDESKNEILMQADSQADSQTDSDESKNE 1547
>At4g14220 RING-H2 finger protein RHF1a
Length = 371
Score = 38.9 bits (89), Expect = 0.004
Identities = 23/69 (33%), Positives = 34/69 (48%), Gaps = 4/69 (5%)
Query: 242 ATGEDAEEEGASLNDEDDDEDALPFACFICRNPFV--DPVS-TKCKHYFCEHCALKHHAK 298
A G + A + DDD + AC IC PF DP + T CKH + C ++ +
Sbjct: 21 AIGSSSSSSSALVVASDDDNNT-DDACSICLEPFTLQDPSTVTSCKHEYHLQCIIEWSQR 79
Query: 299 NKKCFVCNQ 307
+K+C +C Q
Sbjct: 80 SKECPICWQ 88
>At5g63260 unknown protein
Length = 435
Score = 38.5 bits (88), Expect = 0.005
Identities = 47/165 (28%), Positives = 69/165 (41%), Gaps = 25/165 (15%)
Query: 71 SKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALKQAT 130
SK ++ PN PS SS E+ V +A + L ++ D +R L +
Sbjct: 2 SKPEETSDPNPTGPDPSR--SSSDEVTVTVADRAPSDLNHVSEELSDQ--LRNVGLDDSA 57
Query: 131 ESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSARFD 190
+ L S + + +G N TD +A F +Q E+ GS R+ +
Sbjct: 58 KEL------SVPISVPQG-NVETDSRALFGSDQ---KEEEEGSEK--------RMMMVYP 99
Query: 191 YQPDI--CKDYKETGYCGYGDSCKFMHD-RGDYKSGWQMEKEWDE 232
+PD C Y TG C YG SCKF H R + G + +E DE
Sbjct: 100 VRPDSEDCSFYMRTGSCKYGSSCKFNHPVRRKLQIGRERVRERDE 144
Score = 32.7 bits (73), Expect = 0.28
Identities = 12/20 (60%), Positives = 14/20 (70%)
Query: 196 CKDYKETGYCGYGDSCKFMH 215
CK Y TG C YG+SC+F H
Sbjct: 154 CKYYFRTGGCKYGESCRFSH 173
>At5g01960 unknown protein
Length = 426
Score = 38.1 bits (87), Expect = 0.007
Identities = 13/38 (34%), Positives = 20/38 (52%)
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVC 305
C IC+ P+ +CKH FCE C + + + C +C
Sbjct: 365 CAICQEKMHTPILLRCKHMFCEDCVSEWFERERTCPLC 402
>At2g32950 photomorphogenesis repressor (COP1)
Length = 675
Score = 38.1 bits (87), Expect = 0.007
Identities = 20/63 (31%), Positives = 26/63 (40%), Gaps = 3/63 (4%)
Query: 245 EDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFV 304
+D G+ + D D+D L C IC D T C H FC C + H C
Sbjct: 32 DDGGSGGSEIGAPDLDKDLL---CPICMQIIKDAFLTACGHSFCYMCIITHLRNKSDCPC 88
Query: 305 CNQ 307
C+Q
Sbjct: 89 CSQ 91
>At1g66050 hypothetical protein
Length = 598
Score = 38.1 bits (87), Expect = 0.007
Identities = 27/77 (35%), Positives = 35/77 (45%), Gaps = 9/77 (11%)
Query: 232 EAEKARK-MRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEH 290
EAEKA+K RL +G G +D+++ L C IC PV+T C H FC
Sbjct: 98 EAEKAKKRQRLMSG------GGDDGVDDEEKKKLEIFCSICIQLPERPVTTPCGHNFCLK 151
Query: 291 CALKHHAKNKK--CFVC 305
C K K C +C
Sbjct: 152 CFEKWAVGQGKLTCMIC 168
Score = 35.8 bits (81), Expect = 0.033
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 229 EWDEAEKARKMRLATGEDAEEEGASLNDEDDDEDALP-FACFICRNPFVDPVSTKCKHYF 287
+W ++ +M L T E + + A + L F+C ICR PV+T C H F
Sbjct: 433 KWMKSPPVSRMALDTEERKKNKRAKKGNNAMKARLLKEFSCQICRKVLSLPVTTPCAHNF 492
Query: 288 CEHC 291
C+ C
Sbjct: 493 CKAC 496
>At1g66040 hypothetical protein
Length = 622
Score = 38.1 bits (87), Expect = 0.007
Identities = 27/77 (35%), Positives = 35/77 (45%), Gaps = 9/77 (11%)
Query: 232 EAEKARK-MRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEH 290
EAEKA+K RL +G G +D+++ L C IC PV+T C H FC
Sbjct: 98 EAEKAKKRQRLMSG------GGDDGVDDEEKKKLEIFCSICIQLPERPVTTPCGHNFCLK 151
Query: 291 CALKHHAKNKK--CFVC 305
C K K C +C
Sbjct: 152 CFEKWAVGQGKLTCMIC 168
Score = 35.8 bits (81), Expect = 0.033
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 229 EWDEAEKARKMRLATGEDAEEEGASLNDEDDDEDALP-FACFICRNPFVDPVSTKCKHYF 287
+W ++ +M L T E + + A + L F+C ICR PV+T C H F
Sbjct: 458 KWMKSPPVSRMALDTEERKKNKRAKKGNNAMKARLLKEFSCQICRKVLSLPVTTPCAHNF 517
Query: 288 CEHC 291
C+ C
Sbjct: 518 CKAC 521
>At3g27330 hypothetical protein
Length = 913
Score = 37.7 bits (86), Expect = 0.009
Identities = 28/90 (31%), Positives = 42/90 (46%), Gaps = 11/90 (12%)
Query: 227 EKEWDEAEKARKMRLATGEDA--EEEGASLNDE------DDDEDALPFACFICRNPFVDP 278
EK ++ +K R+M T E+ +++ SL+D D D L +C IC +P
Sbjct: 677 EKSAEKKKKKREMETETKEEKKIDKDEKSLSDHVVLPCIDKLRDEL--SCAICLEICFEP 734
Query: 279 VSTKCKHYFCEHCALKHHAK-NKKCFVCNQ 307
+T C H FC+ C K +KC C Q
Sbjct: 735 STTTCGHSFCKKCLRSAADKCGRKCPKCRQ 764
>At5g16500 protein kinase-like protein
Length = 636
Score = 37.0 bits (84), Expect = 0.015
Identities = 45/244 (18%), Positives = 105/244 (42%), Gaps = 18/244 (7%)
Query: 18 CSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFSTGSSKSSASA 77
CS ++RK++ ++ + D+E++ +E ++++T + + +
Sbjct: 384 CSVTPFCISRKDVGNKSSSSSDSEDEEEEKEQKAEKEEESTSKKRQ-------EQEETAT 436
Query: 78 KPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALKQATESLKGKS 137
++ES+ S E +E + KA + + +D + R+I E + + +
Sbjct: 437 DSDDESDSNS---EKDQEEEQSQLEKARESSSSSSDSGSERRSIDETNATAQSLKISYSN 493
Query: 138 TSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVS--ARFDYQPDI 195
SSE+ K +++ + K+ E T + +G H + +R++ A D + D
Sbjct: 494 YSSEEEDNEK-LSSKSSCKS--NEESTFSRYDSGRDHDDSSRNTSMRINSLAHDDKEEDE 550
Query: 196 CKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDAEEEGASLN 255
++++ Y + DS + R + S + + +E E+ + RL ++ E S+
Sbjct: 551 EENHETRSYSDHDDSPRNTSMRINSLS---HDDDEEEEEENHQTRLEHIHSSKSEDQSVY 607
Query: 256 DEDD 259
+DD
Sbjct: 608 SDDD 611
Score = 33.1 bits (74), Expect = 0.21
Identities = 29/118 (24%), Positives = 53/118 (44%), Gaps = 5/118 (4%)
Query: 4 SEAKSTENQQTEQVCSFFRKPVNRKNM-RKRTIENEDNENDSNNEESLMHVQKKNTKADN 62
S E ++ EQ + ++K ++ T + D+E+DSN+E+ Q + KA
Sbjct: 403 SSDSEDEEEEKEQKAEKEEESTSKKRQEQEETATDSDDESDSNSEKDQEEEQSQLEKARE 462
Query: 63 KLFFSTGSS---KSSASAKPNEESEKPSF-HFESSKEIQVQHDSKATATLETETDFSR 116
S+ S +S +S K S+ ++ S +E + SK++ E+ FSR
Sbjct: 463 SSSSSSDSGSERRSIDETNATAQSLKISYSNYSSEEEDNEKLSSKSSCKSNEESTFSR 520
>At2g42030 putative RING zinc finger protein
Length = 425
Score = 37.0 bits (84), Expect = 0.015
Identities = 22/81 (27%), Positives = 36/81 (44%), Gaps = 2/81 (2%)
Query: 227 EKEWDEAEKARKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHY 286
E+ ++++K ED E ++ D F C+IC + DPV T C H
Sbjct: 100 ERVIEDSKKCENGSKVMAEDNVTEEKRDVEKSVGSDGSFFDCYICLDLSKDPVVTNCGHL 159
Query: 287 FCEHCALK--HHAKNKKCFVC 305
+C C + ++ K+C VC
Sbjct: 160 YCWSCLYQWLQVSEAKECPVC 180
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.127 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,683,836
Number of Sequences: 26719
Number of extensions: 343997
Number of successful extensions: 2438
Number of sequences better than 10.0: 254
Number of HSP's better than 10.0 without gapping: 90
Number of HSP's successfully gapped in prelim test: 164
Number of HSP's that attempted gapping in prelim test: 2134
Number of HSP's gapped (non-prelim): 432
length of query: 327
length of database: 11,318,596
effective HSP length: 100
effective length of query: 227
effective length of database: 8,646,696
effective search space: 1962799992
effective search space used: 1962799992
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC126016.2


