

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126015.14 + phase: 0 /pseudo
(241 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
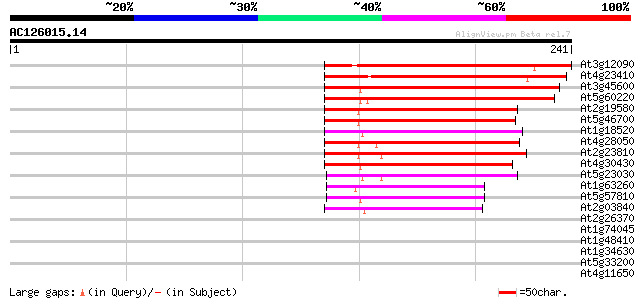
Score E
Sequences producing significant alignments: (bits) Value
At3g12090 putative protein 143 9e-35
At4g23410 unknown protein 142 2e-34
At3g45600 unknown protein 108 3e-24
At5g60220 senescence-associated protein - like 105 2e-23
At2g19580 putative senescence-associated protein 5 102 1e-22
At5g46700 senescence-associated protein 5-like protein 97 8e-21
At1g18520 unknown protein 95 4e-20
At4g28050 senescence-associated protein -like 94 7e-20
At2g23810 similar to senescence-associated protein 87 6e-18
At4g30430 senescence-associated protein homolog 85 4e-17
At5g23030 senescence-associated protein 5-like protein 80 1e-15
At1g63260 unknown protein 63 2e-10
At5g57810 unknown protein 62 2e-10
At2g03840 putative senescence-associated protein 57 9e-09
At2g26370 hypothetical protein 29 2.6
At1g74045 putative protein 28 4.4
At1g48410 Argonaute protein (AGO1) 28 4.4
At1g34630 unknown protein 28 5.7
At5g33200 putative protein 27 7.5
At4g11650 osmotin precursor like protein 27 7.5
>At3g12090 putative protein
Length = 282
Score = 143 bits (360), Expect = 9e-35
Identities = 62/107 (57%), Positives = 81/107 (74%), Gaps = 3/107 (2%)
Query: 136 TGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVL 195
+GCCKPPTAC Y EA ++ DC++W+N +LCYECD+CKAGVLE+IR +W KLSV+
Sbjct: 165 SGCCKPPTACTY--EAGVVDGGGDCFRWNNGVEMLCYECDACKAGVLEEIRLDWRKLSVV 222
Query: 196 TVTMLILLIGIYSIGCCAFRNARRAETDY-PYGENRMTKVRPRWDYH 241
+ +L+LLI +Y+ GCCAF N R A Y P +NRMT+VRPRWDY+
Sbjct: 223 NILVLVLLIAVYAAGCCAFHNTRHAAHPYHPSDDNRMTRVRPRWDYY 269
>At4g23410 unknown protein
Length = 281
Score = 142 bits (357), Expect = 2e-34
Identities = 58/105 (55%), Positives = 82/105 (77%), Gaps = 2/105 (1%)
Query: 136 TGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVL 195
+GCCKPPT+C YN + V+ QD DCY+W+N T+LCY+CD+C+AGVLE +RR+WHKLS++
Sbjct: 165 SGCCKPPTSCVYNTDTVIQ-QDPDCYRWNNAATVLCYDCDTCRAGVLETVRRDWHKLSLV 223
Query: 196 TVTMLILLIGIYSIGCCAFRNARRAE-TDYPYGENRMTKVRPRWD 239
V ++I LI +Y +GCCAF+NA+R + +PYG M+K RP W+
Sbjct: 224 NVIVVIFLIAVYCVGCCAFKNAKRPQHYGFPYGRYGMSKSRPGWE 268
>At3g45600 unknown protein
Length = 285
Score = 108 bits (270), Expect = 3e-24
Identities = 47/109 (43%), Positives = 69/109 (63%), Gaps = 8/109 (7%)
Query: 136 TGCCKPPTACNYNM--------EAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKPPT C ++ M+ + DC WSN+ ++LCY+C SCKAGVL +++
Sbjct: 172 SGCCKPPTDCGFSYVNETGWDTRGGMIGPNQDCMVWSNDQSMLCYQCSSCKAGVLGSLKK 231
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRP 236
+W K+SV+ + +LI+L+ Y I A+RN +R + D P GE RMTK P
Sbjct: 232 SWRKVSVINIVVLIILVIFYVIAYAAYRNVKRIDNDEPAGEARMTKSHP 280
>At5g60220 senescence-associated protein - like
Length = 327
Score = 105 bits (262), Expect = 2e-23
Identities = 47/107 (43%), Positives = 68/107 (62%), Gaps = 8/107 (7%)
Query: 136 TGCCKPPTACNYNM--EAV------MMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKPPT C Y E V M+ + DC W+N+ LLCY+C SCKAGVL +++
Sbjct: 172 SGCCKPPTDCGYTYVNETVWIPGGEMVGPNPDCMLWNNDQRLLCYQCSSCKAGVLGSLKK 231
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKV 234
+W K+SV+ + ++I+L+ Y I C A++N +R D P GE RMT +
Sbjct: 232 SWRKVSVINIVVVIILVIFYVIACAAYQNVKRMYNDEPVGEARMTNL 278
>At2g19580 putative senescence-associated protein 5
Length = 270
Score = 102 bits (255), Expect = 1e-22
Identities = 42/90 (46%), Positives = 61/90 (67%), Gaps = 7/90 (7%)
Query: 136 TGCCKPPTACNYN-------MEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKPPTAC YN + M D+DCY WSN+ + LCY C+SCKAG+L ++R+
Sbjct: 168 SGCCKPPTACGYNFVNPTLWLNPTNMAADADCYLWSNDQSQLCYNCNSCKAGLLGNLRKE 227
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNAR 218
W K +++ + +++LI +Y I C AFRNA+
Sbjct: 228 WRKANLILIITVVVLIWVYVIACSAFRNAQ 257
>At5g46700 senescence-associated protein 5-like protein
Length = 269
Score = 97.1 bits (240), Expect = 8e-21
Identities = 43/89 (48%), Positives = 56/89 (62%), Gaps = 7/89 (7%)
Query: 136 TGCCKPPTACNYN-------MEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKPPT C + + + M+ D DC WSN+ LCY CDSCKAG+L +I+ +
Sbjct: 167 SGCCKPPTKCGFTFVNPTYWISPIDMSADMDCLNWSNDQNTLCYTCDSCKAGLLANIKVD 226
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNA 217
W K + + LI LI +Y IGCCAFRNA
Sbjct: 227 WLKADIFLLLALIGLIIVYIIGCCAFRNA 255
>At1g18520 unknown protein
Length = 271
Score = 94.7 bits (234), Expect = 4e-20
Identities = 43/96 (44%), Positives = 56/96 (57%), Gaps = 11/96 (11%)
Query: 136 TGCCKPPTACNYNME-----------AVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLED 184
+GCCKPP+ CN+ + ++ DC WSN T LC+ C++CKAGVL +
Sbjct: 170 SGCCKPPSDCNFEFRNATFWIPPSKNETAVAENGDCGTWSNVQTELCFNCNACKAGVLAN 229
Query: 185 IRRNWHKLSVLTVTMLILLIGIYSIGCCAFRNARRA 220
IR W L V + +LILLI +YS GCCA RN R A
Sbjct: 230 IREKWRNLLVFNICLLILLITVYSCGCCARRNNRTA 265
>At4g28050 senescence-associated protein -like
Length = 263
Score = 94.0 bits (232), Expect = 7e-20
Identities = 39/91 (42%), Positives = 60/91 (65%), Gaps = 7/91 (7%)
Query: 136 TGCCKPPTACNYN-MEAVMMTQ------DSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKP CN+ + T+ + DC W N+P LCY+C++CKAG+L++I+ +
Sbjct: 170 SGCCKPSNDCNFTYVNPTTWTKTPGPYKNEDCNVWDNKPGTLCYDCEACKAGLLDNIKNS 229
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNARR 219
W K++ + + LI LI +YS+GCCAFRN R+
Sbjct: 230 WKKVAKVNIVFLIFLIIVYSVGCCAFRNNRK 260
>At2g23810 similar to senescence-associated protein
Length = 195
Score = 87.4 bits (215), Expect = 6e-18
Identities = 35/95 (36%), Positives = 59/95 (61%), Gaps = 8/95 (8%)
Query: 136 TGCCKPPTACNYN-MEAVMMTQDS-------DCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKP C + + T+++ DC W N LC++C SCKAG+L++++
Sbjct: 92 SGCCKPSDECGFEYVNPTTWTKNTTGTHTNPDCQTWDNAKEKLCFDCQSCKAGLLDNVKS 151
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAET 222
W K++++ + L+ LI +YS+GCCAFRN +R ++
Sbjct: 152 AWKKVAIVNIVFLVFLIIVYSVGCCAFRNNKRDDS 186
>At4g30430 senescence-associated protein homolog
Length = 272
Score = 84.7 bits (208), Expect = 4e-17
Identities = 33/88 (37%), Positives = 55/88 (62%), Gaps = 7/88 (7%)
Query: 136 TGCCKPPTACNYNM-------EAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKP C++ + ++SDC W NE LCY C +CKAG L++++
Sbjct: 170 SGCCKPSNDCDFTYITSTTWNKTSGTHKNSDCQLWDNEKHKLCYNCKACKAGFLDNLKAA 229
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRN 216
W +++++ + L+LL+ +Y++GCCAFRN
Sbjct: 230 WKRVAIVNIIFLVLLVVVYAMGCCAFRN 257
>At5g23030 senescence-associated protein 5-like protein
Length = 264
Score = 79.7 bits (195), Expect = 1e-15
Identities = 36/91 (39%), Positives = 52/91 (56%), Gaps = 9/91 (9%)
Query: 137 GCCKPPTACNYNME-----AVMMTQDS----DCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
GCC+PP C + + V T + DC WSN LCY C+SCK GVL+ IR+
Sbjct: 166 GCCRPPVECGFESKNATWWTVPATATTAIIGDCKAWSNTQRQLCYACESCKIGVLKGIRK 225
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNAR 218
W L V+ + +++L++ +YS GCC +N R
Sbjct: 226 RWRILIVVNLLLILLVVFLYSCGCCVRKNNR 256
>At1g63260 unknown protein
Length = 284
Score = 62.8 bits (151), Expect = 2e-10
Identities = 25/76 (32%), Positives = 43/76 (55%), Gaps = 8/76 (10%)
Query: 137 GCCKPPTACNY--------NMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
GCC+PP+ C Y ++ ++ + DC + N T+ CY CDSCKAGV + ++
Sbjct: 169 GCCRPPSECGYPAVNASYYDLSFHSISSNKDCKLYKNLRTIKCYNCDSCKAGVAQYMKTE 228
Query: 189 WHKLSVLTVTMLILLI 204
W +++ V + ++LI
Sbjct: 229 WRLVAIFNVVLFVVLI 244
>At5g57810 unknown protein
Length = 317
Score = 62.4 bits (150), Expect = 2e-10
Identities = 28/91 (30%), Positives = 44/91 (47%), Gaps = 23/91 (25%)
Query: 137 GCCKPPTACNYNM-----------------------EAVMMTQDSDCYKWSNEPTLLCYE 173
GCC PP CN + + +M+ + SDC W N+ ++LCY+
Sbjct: 210 GCCMPPETCNMDAINATFWYRRKDEGPPSSMNLMYGDEMMVGRISDCQLWRNDWSILCYD 269
Query: 174 CDSCKAGVLEDIRRNWHKLSVLTVTMLILLI 204
C SCK G + +RR W +L + + + ILL+
Sbjct: 270 CRSCKFGFIRSVRRKWWQLGIFLIVISILLL 300
>At2g03840 putative senescence-associated protein
Length = 278
Score = 57.0 bits (136), Expect = 9e-09
Identities = 24/76 (31%), Positives = 39/76 (50%), Gaps = 8/76 (10%)
Query: 136 TGCCKPPTACNYNMEA--------VMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKPP +C N E + DC +W+N LC++CDSCKA ++ D+
Sbjct: 184 SGCCKPPLSCGLNYEKPNNWTVSRYYNNLEVDCKRWNNSADTLCFDCDSCKAVIIADVHN 243
Query: 188 NWHKLSVLTVTMLILL 203
++V + ++ L
Sbjct: 244 TSFSITVNIIHIIFSL 259
>At2g26370 hypothetical protein
Length = 173
Score = 28.9 bits (63), Expect = 2.6
Identities = 13/45 (28%), Positives = 20/45 (43%)
Query: 190 HKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKV 234
H VL + + + + G F N + A DY YG +T+V
Sbjct: 5 HAQPVLLLLSSLYFLSAFGAGAYNFENCKNAPIDYNYGITNVTRV 49
>At1g74045 putative protein
Length = 215
Score = 28.1 bits (61), Expect = 4.4
Identities = 16/75 (21%), Positives = 26/75 (34%), Gaps = 14/75 (18%)
Query: 138 CCKPPTACNYNMEAVMMTQDSDCYKWSNEPTL----------LCYECDSCKAGVLEDIRR 187
CC P C + M + W ++ C +C C+ +L+ I
Sbjct: 119 CCAQPVGCG----TITMFDKPGEWSWKHQYERNQVPEECSYEYCLDCRGCQLSILKAIVH 174
Query: 188 NWHKLSVLTVTMLIL 202
W LS+ L+L
Sbjct: 175 QWKYLSMFAYPALVL 189
>At1g48410 Argonaute protein (AGO1)
Length = 870
Score = 28.1 bits (61), Expect = 4.4
Identities = 15/39 (38%), Positives = 25/39 (63%), Gaps = 1/39 (2%)
Query: 84 VSSYRLFTLVEEKNTRSSLLEYYKELY-FRV*NL*QTCL 121
V++ L V+E+NT+ S++EY+ E Y FR+ + CL
Sbjct: 265 VATRELTFPVDERNTQKSVVEYFHETYGFRIQHTQLPCL 303
>At1g34630 unknown protein
Length = 481
Score = 27.7 bits (60), Expect = 5.7
Identities = 11/36 (30%), Positives = 18/36 (49%)
Query: 21 RFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSS 56
RF C ++W H LF C+ + +L Y++ S
Sbjct: 193 RFGTICKPLTWKHGDLFLMCLSSSQILSAYILKQES 228
>At5g33200 putative protein
Length = 426
Score = 27.3 bits (59), Expect = 7.5
Identities = 11/35 (31%), Positives = 17/35 (48%)
Query: 34 WCLFSCCMCTMVVLGDYVIANSSTIRFDYIWFWCD 68
W FSC C V + N+++++ FWCD
Sbjct: 212 WYYFSCRNCNKKVTHIHAGVNTTSLKSSKPRFWCD 246
>At4g11650 osmotin precursor like protein
Length = 244
Score = 27.3 bits (59), Expect = 7.5
Identities = 9/18 (50%), Positives = 14/18 (77%)
Query: 11 KFSPNTTSCNRFYCTCDI 28
+FSP +++C+R CT DI
Sbjct: 130 EFSPTSSNCHRILCTADI 147
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.343 0.149 0.575
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,476,643
Number of Sequences: 26719
Number of extensions: 208244
Number of successful extensions: 865
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 828
Number of HSP's gapped (non-prelim): 22
length of query: 241
length of database: 11,318,596
effective HSP length: 96
effective length of query: 145
effective length of database: 8,753,572
effective search space: 1269267940
effective search space used: 1269267940
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC126015.14


