

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
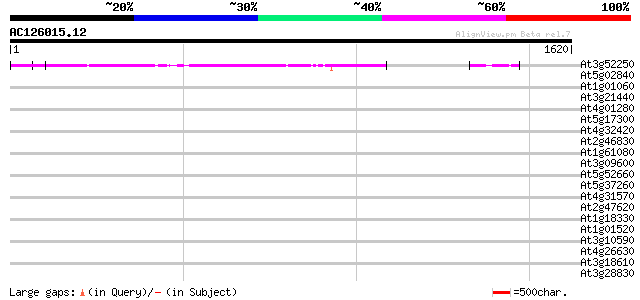
Query= AC126015.12 - phase: 0
(1620 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g52250 putative protein 593 e-169
At5g02840 unknown protein 43 0.001
At1g01060 DNA-binding protein, putative 43 0.001
At3g21440 hypothetical protein 40 0.008
At4g01280 myb-related DNA-binding protein like 39 0.019
At5g17300 unknown protein 39 0.025
At4g32420 cyclophilin (AtCYP95) 39 0.032
At2g46830 CCA1 39 0.032
At1g61080 hypothetical protein 39 0.032
At3g09600 putative MYB-related protein 38 0.042
At5g52660 unknown protein 38 0.055
At5g37260 putative protein 38 0.055
At4g31570 putative protein 37 0.071
At2g47620 SWI/SNF family like transcription activator 37 0.093
At1g18330 hypothetical protein 37 0.093
At1g01520 hypothetical protein 37 0.093
At3g10590 hypothetical protein 36 0.16
At4g26630 unknown protein 36 0.21
At3g18610 unknown protein 35 0.27
At3g28830 hypothetical protein 35 0.35
>At3g52250 putative protein
Length = 1677
Score = 593 bits (1529), Expect = e-169
Identities = 391/1051 (37%), Positives = 580/1051 (54%), Gaps = 79/1051 (7%)
Query: 67 SRRPQDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTG--GVNGLGTGPRCDR 124
+++ D+ +R V D+ + P+++ TWEQ LKD G+N + +C +
Sbjct: 249 AKQCNDLMYGRRLVSDNSLDAPIPNAELEGTWEQLRLKDPQDNNSLHGINDIDGDRKCAK 308
Query: 125 ENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEGKPNLKHKNVTAVESNS 184
E+SL + PL W SGS +S+ SGFSHSSS +S+ DS + K + K VT +S+S
Sbjct: 309 ESSLGATGKLPL-WNSSGSFASQSSGFSHSSSLKSLGAVDSSDRKIEVLPKIVTVTQSSS 367
Query: 185 GEATACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSAGNMEPC 244
G+ATAC T++ SE+ +SRKK RL WGEGLAKYEKKKVDV N+DG+ +E
Sbjct: 368 GDATACATTTHLSEEMSSRKKQRLGWGEGLAKYEKKKVDV---NPNEDGTTLMENGLEEL 424
Query: 245 SSISPNLVDKSPKVTGFSDCASPATPSSVACSSSPGVDDKLLGKVGNADNDVSNLTDSPA 304
S++ N+ DKSP D SP TPSSVACSSSPG DK K A +DVSN+ SP+
Sbjct: 425 HSLNKNIADKSPTAAIVPDYGSPTTPSSVACSSSPGFADKSSPKAAIAASDVSNMCRSPS 484
Query: 305 PGFQNHLQKFYLNLDKLDVDSLNSLGSSIVELVQSDDPSSDDSGLVRSNAINKLLIWKAD 364
P HL++F +N+++LD S+ G + EL+ +DD + DS V+ ++N LL WK +
Sbjct: 485 PVSSIHLERFPINIEELDNISMERFGCLLNELLGTDDSGTGDSSSVQLTSMNTLLAWKGE 544
Query: 365 ISKVLEMTESEIDLLENELKSLKSESVDRSECPVASGSQQADSSSKFYEERVEVSQKVIR 424
I K +EMTESEIDLLEN+ ++LK E S V S D + +E+ S
Sbjct: 545 ILKAVEMTESEIDLLENKHRTLKLEGRRHSRV-VGPSSYCCDGDANVPKEQASCSLD--- 600
Query: 425 PVPLKIISSDEPNTVKMP-QSTNLCSIHENDKEEDIDSPGSATSKFVEPLPVNAVSSSYT 483
P SS V+ P L + + E DSPG V+PL +
Sbjct: 601 --PKATASSVAKTLVRAPVHQAGLAKVPADVFE---DSPGE-----VKPL---------S 641
Query: 484 RGYDNLSRDMNAVQSTMMKCFVRCNRKNTSVSACNNVNTPTEVKDSLGDVTFGANLCSSY 543
+ + + R+ + + MK V + +NTP +V+ + +S
Sbjct: 642 QSFATVEREEDILPIPSMKAAV----------SSKEINTPAFANQETIEVSSADDSMASK 691
Query: 544 GDTY-KSIIASNKESANRAHKLFTKLVPKECKKHGNM---GVSNDSFSHTSILQKFAEKK 599
D + ++++NK+ A + +F +L+P++ N G+ F + + +K A++
Sbjct: 692 EDLFWAKLLSANKKYACESSGVFNQLLPRDFNSSDNSRFPGICQTQFD-SHVQEKIADRV 750
Query: 600 QFERFKERVIALKFKALHHLWKEDMRLLSIRKCRPKSHKKNELNVRTTCSSNMKNRSSIR 659
R +E+++ L+FKA WK+D+ L++ K + KS KK EL +K S+R
Sbjct: 751 GLLRAREKILLLQFKAFQLSWKKDLDQLALAKYQSKSSKKTELYPNAKNGGYLKLPQSVR 810
Query: 660 SRFTFPAGNHLSLVPTTEIINFTSKLLSESQAQLQRNTLKMPALILDEKEKMVTKFISSN 719
RF+ A S+VPTTE++++ KLL + + R+ LKMPA+ILDEKE+++++FISSN
Sbjct: 811 LRFSSSAPRRDSVVPTTELVSYMEKLLPGTHLKPFRDILKMPAMILDEKERVMSRFISSN 870
Query: 720 GLVEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKN 779
GL+EDP +EKER+MINPWTSEEKE+FL A GKDF+KIAS L KTTADCI++YYKN
Sbjct: 871 GLIEDPCDVEKERTMINPWTSEEKEIFLNLLAMHGKDFKKIASSLTQKTTADCIDYYYKN 930
Query: 780 HKSECFEKLKR-KDIGKLGKSYAAKTNLMASGKKWNHEVNVSSLDILSAASVM---ADVI 835
HKS+CF K+K+ + GK GK T ++A KKW E+ +SLDIL S++ A +
Sbjct: 931 HKSDCFGKIKKQRAYGKEGK----HTYMLAPRKKWKREMGAASLDILGDVSIIAANAGKV 986
Query: 836 AGNKRMRGRRYLL-GYGNVKASRGEDSIIER-SNSFDTLGDERETAAAADVLAGICGSFS 893
A + + ++ L G + + + + + E S SFD R+ A ADVLA G S
Sbjct: 987 ASTRPISSKKITLRGCSSANSLQHDGNNSEGCSYSFDF---PRKRTAGADVLA--VGPLS 1041
Query: 894 SEAMSSCITSSIDPVDGNKETKFLKANPLFKQP------------LTPDISQNADDETCS 941
E ++SC+ +S+ + LK N + K+P + + N +D++CS
Sbjct: 1042 PEQINSCLRTSVS--SRERCMDHLKFNHVVKKPRISHTLHNENSNTLHNENSNEEDDSCS 1099
Query: 942 DESCGEA--TEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFFSKTRKCLGLN 999
+ESCGE WTDDE +AF+Q S FGK+F ISR VGT++ + CK FFSK RKCLGL
Sbjct: 1100 EESCGETGPIHWTDDERSAFIQGFSLFGKNFASISRYVGTRSPDQCKVFFSKVRKCLGLE 1159
Query: 1000 LANPVPG-INGSPLNDDAN-GGESDTDDACVVEAGSVVDADKSGNKTDEDLPSDALNTFH 1057
G ++ S D+ N GG SD +D C +E+ S + + K + P+ N
Sbjct: 1160 SIKFGSGNVSTSVSVDNGNEGGGSDLEDPCPMESNSGIVNNGVCAKMGMNSPTSPFNMNQ 1219
Query: 1058 DESNPLEATSLSAKLNESREISGTE-VCLEN 1087
D N + ++ A L+ S E +G + +CL++
Sbjct: 1220 DGVNQSGSANVKADLSRSEEENGQKYLCLKD 1250
Score = 80.9 bits (198), Expect = 6e-15
Identities = 43/101 (42%), Positives = 61/101 (59%), Gaps = 2/101 (1%)
Query: 2 FSEEPGHGYGVSRSGDKMLEEDGRPLVSRGDGKYGRSSRDNRGGPFGQRDWRGHSWEASN 61
F EE HGY SRS +M + + RP SRGD +Y R+ RD+R Q++W+ ++WE SN
Sbjct: 91 FVEETSHGYTSSRSSARMFD-NYRPSASRGDWRYTRNCRDDRVS-VSQKEWKCNTWEMSN 148
Query: 62 GSPNLSRRPQDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHN 102
GS RP + N +RSVD+ P ++S HS VN+ + N
Sbjct: 149 GSSRSFERPFGIRNGRRSVDERPLHASDTHSTVVNSLDPAN 189
Score = 80.5 bits (197), Expect = 7e-15
Identities = 56/146 (38%), Positives = 81/146 (55%), Gaps = 24/146 (16%)
Query: 1329 CSSSAAEFPLLPQKVKQTDGHFKP-SFHSSNSEKTSRNGDVKLFGKILTNPSSTQNPNLT 1387
C+SSA + PL + GH + SF S++E+ +NGDVKLFG +LT
Sbjct: 1500 CTSSAPK-PLAVSHKEGRSGHSRSHSFSLSDTERLHKNGDVKLFGTVLT----------- 1547
Query: 1388 AKRSEENG---SHHPKLNNKSSNLNFTGHQNSDENLNFLKFGLENVPVMSYGYWEGNAIQ 1444
++ENG H+P +SS+ H +N + L+NVP+ SYG+W+GN I
Sbjct: 1548 ---TDENGIKQKHNPCGIVRSSSTLSRDHDTRHHYIN--QQHLQNVPITSYGFWDGNRI- 1601
Query: 1445 SRQSGLSSLPDSSFLLAKYPAAFSNY 1470
Q+GL+SLP+S+ LLA P AFS +
Sbjct: 1602 --QTGLTSLPESAKLLASCPEAFSTH 1625
>At5g02840 unknown protein
Length = 293
Score = 43.1 bits (100), Expect = 0.001
Identities = 26/94 (27%), Positives = 45/94 (47%), Gaps = 19/94 (20%)
Query: 920 NPLFKQPLTPDISQNADDETCSDESCGEATE---------------WTDDETAAFLQAVS 964
NP+ + + + S +A + T + GEA E WT+ E FL+A+
Sbjct: 5 NPVVAEVIPAETSTDATETTIATTEAGEAPEKKVRKAYTITKSRESWTEGEHDKFLEALQ 64
Query: 965 SFGKDFEKISRCVGTKA----QEHCKRFFSKTRK 994
F +D++KI VG+K + H +++F K +K
Sbjct: 65 LFDRDWKKIEDFVGSKTVIQIRSHAQKYFLKVQK 98
Score = 32.3 bits (72), Expect = 2.3
Identities = 16/53 (30%), Positives = 27/53 (50%), Gaps = 5/53 (9%)
Query: 738 WTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSECFEKLKR 790
WT E + FLE F +D++KI F+ KT ++H + F K+++
Sbjct: 51 WTEGEHDKFLEALQLFDRDWKKIEDFVGSKTV-----IQIRSHAQKYFLKVQK 98
>At1g01060 DNA-binding protein, putative
Length = 645
Score = 43.1 bits (100), Expect = 0.001
Identities = 38/148 (25%), Positives = 60/148 (39%), Gaps = 28/148 (18%)
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRK---------CLG 997
WT+DE FL+A+ +G+ +++I +GTK + H ++FF+K K C
Sbjct: 27 WTEDEHERFLEALRLYGRAWQRIEEHIGTKTAVQIRSHAQKFFTKLEKEAEVKGIPVCQA 86
Query: 998 LNLANPVPGINGSP----LNDDANGGESDTDDACVVEAGSVVDADKSG-NKTDEDLPSDA 1052
L++ P P P N G S + + +A V A S N+ DL
Sbjct: 87 LDIEIPPPRPKRKPNTPYPRKPGNNGTSSSQVSSAKDAKLVSSASSSQLNQAFLDL---- 142
Query: 1053 LNTFHDESNPLEATSLSAKLNESREISG 1080
E P + + K N+ SG
Sbjct: 143 ------EKMPFSEKTSTGKENQDENCSG 164
Score = 34.3 bits (77), Expect = 0.60
Identities = 19/67 (28%), Positives = 33/67 (48%), Gaps = 8/67 (11%)
Query: 725 PLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSEC 784
P I K+R WT +E E FLE +G+ +++I + KT ++H +
Sbjct: 17 PYTITKQRER---WTEDEHERFLEALRLYGRAWQRIEEHIGTKTAVQ-----IRSHAQKF 68
Query: 785 FEKLKRK 791
F KL+++
Sbjct: 69 FTKLEKE 75
>At3g21440 hypothetical protein
Length = 193
Score = 40.4 bits (93), Expect = 0.008
Identities = 20/54 (37%), Positives = 33/54 (61%), Gaps = 2/54 (3%)
Query: 730 KERSMINP-WTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKS 782
K M+ P W+ EE E F E + FGK+++K+A F+ H +A+ +E Y +K+
Sbjct: 38 KLSDMLGPQWSKEELERFYEGYRKFGKEWKKVAGFV-HSRSAEMVEALYTMNKA 90
Score = 32.0 bits (71), Expect = 3.0
Identities = 17/82 (20%), Positives = 39/82 (46%), Gaps = 7/82 (8%)
Query: 924 KQPLTPDISQNADDETCSDESCGE-------ATEWTDDETAAFLQAVSSFGKDFEKISRC 976
K+P +S + D+E+ S + +W+ +E F + FGK+++K++
Sbjct: 13 KKPRAKAVSPHKDEESMSKTKQRKRKLSDMLGPQWSKEELERFYEGYRKFGKEWKKVAGF 72
Query: 977 VGTKAQEHCKRFFSKTRKCLGL 998
V +++ E + ++ + L L
Sbjct: 73 VHSRSAEMVEALYTMNKAYLSL 94
>At4g01280 myb-related DNA-binding protein like
Length = 302
Score = 39.3 bits (90), Expect = 0.019
Identities = 22/59 (37%), Positives = 33/59 (55%), Gaps = 5/59 (8%)
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRKCLGLNLANPVP 1005
WTD E FL+A+ F +D++KI VG+K + H +++F K +K G N P P
Sbjct: 62 WTDQEHDKFLEALHLFDRDWKKIEAFVGSKTVVQIRSHAQKYFLKVQKS-GANEHLPPP 119
Score = 37.0 bits (84), Expect = 0.093
Identities = 20/69 (28%), Positives = 35/69 (49%), Gaps = 8/69 (11%)
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
+ P I+K R WT +E + FLE F +D++KI +F+ KT ++H
Sbjct: 49 IRKPYTIKKSREN---WTDQEHDKFLEALHLFDRDWKKIEAFVGSKTVVQ-----IRSHA 100
Query: 782 SECFEKLKR 790
+ F K+++
Sbjct: 101 QKYFLKVQK 109
>At5g17300 unknown protein
Length = 387
Score = 38.9 bits (89), Expect = 0.025
Identities = 24/78 (30%), Positives = 36/78 (45%), Gaps = 8/78 (10%)
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
V P I KER WT EE + F+E +G+ +R+I + KT ++H
Sbjct: 45 VRKPYTITKERER---WTDEEHKKFVEALKLYGRAWRRIEEHVGSKTAVQ-----IRSHA 96
Query: 782 SECFEKLKRKDIGKLGKS 799
+ F K+ R+ G G S
Sbjct: 97 QKFFSKVAREATGGDGSS 114
Score = 38.5 bits (88), Expect = 0.032
Identities = 17/52 (32%), Positives = 30/52 (57%), Gaps = 4/52 (7%)
Query: 947 EATEWTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRK 994
E WTD+E F++A+ +G+ + +I VG+K + H ++FFSK +
Sbjct: 54 ERERWTDEEHKKFVEALKLYGRAWRRIEEHVGSKTAVQIRSHAQKFFSKVAR 105
>At4g32420 cyclophilin (AtCYP95)
Length = 837
Score = 38.5 bits (88), Expect = 0.032
Identities = 102/496 (20%), Positives = 186/496 (36%), Gaps = 74/496 (14%)
Query: 15 SGDKMLEEDGRPLVSRGDGKYGRSSRDNRGGPFGQRDWRGHSWEASNGSPNLSRRPQDMN 74
SGDK + DG+ +GK+ +S R R + R HS S S + + +
Sbjct: 176 SGDKK-KSDGKK-----NGKHKKSLRVRR------KKRRRHSSSESESSSDSETDSSESD 223
Query: 75 NEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTGGVNGLGTGPRCDRENSLSSIDWK 134
+E S SP++ S + ++ + KD+H ++ R +++ S+ D +
Sbjct: 224 SESDSDLSSPSFLSSSSHERQKKRKRSSKKDKHRRS-----KQRDKRHEKKRSMR--DKR 276
Query: 135 PLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEGKPNLKHKNVTAVESNSGEATACVTSS 194
P + +R S S+S S S++ + G KH+ V +G + V
Sbjct: 277 PKRKSRRSPDSLED---SNSGSEASLSDVNVEIGAKKRKHR----VSRRTGNSAPAVEKE 329
Query: 195 MPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSAGNMEPCSSISPNLVDK 254
S RK P L GL + +S A + E S P++VD
Sbjct: 330 AESLHQGKRKGPDLLENRGL----------------RSNGISDAAS-EQISDRQPDIVDD 372
Query: 255 SP-----------KVTGFSDCASPATPSSVACSSSPGVDDKLLGKVGNADNDVSNLTDS- 302
P + S SP S + SSSP + G+ V NLT+S
Sbjct: 373 HPSKSRSRSLSPKRTVSKSTSVSPRRSQSKSPSSSPRWNGGRSPAKGS--RQVKNLTNSR 430
Query: 303 -PAPGFQ---NHLQKFYLNLDKLDVDSLNSLGSSIVELVQSDDPSSDDSGLVRSNAIN-- 356
+PG + H+++ S+ S V + + D S S + + +
Sbjct: 431 RESPGSEEKGRHVRR----------SPTKSVSRSPVRVKKERDISRSPSKSLSRSPLRSP 480
Query: 357 KLLIWKADISKVLEMTESEIDLLENELKSLKSESVDRSECPVASGSQQADSSSKFYEERV 416
K +I ++ + + + S + SL + RS S S S +
Sbjct: 481 KRVISRSPVRGRIARSPSRSPVRSASRGSLGRGPLRRSSRRSPSRSPVRSSRRSLSRSPI 540
Query: 417 EVSQKVIRPVPLKII-SSDEPNTVKMPQSTNLCSIHENDKEEDIDSPGSATSKFVEPLPV 475
++S++ + P ++ S + ++ P+ + S + ++ SP ++ + + PV
Sbjct: 541 QLSRRSLSRSPTRLSRRSLSRSPIRSPRKSVSRSPVRSSRKSVSRSPVRSSRRRISRSPV 600
Query: 476 NAVSSSYTRGYDNLSR 491
+ S +R LSR
Sbjct: 601 RSSRKSVSRSPIRLSR 616
>At2g46830 CCA1
Length = 608
Score = 38.5 bits (88), Expect = 0.032
Identities = 32/129 (24%), Positives = 57/129 (43%), Gaps = 17/129 (13%)
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRK-------CLG-- 997
WT++E F++A+ +G+ ++KI V TK + H ++FFSK K +G
Sbjct: 27 WTEEEHNRFIEALRLYGRAWQKIEEHVATKTAVQIRSHAQKFFSKVEKEAEAKGVAMGQA 86
Query: 998 LNLANPVPGINGSPLNDDANGGESDTDDACVVEAGSVVDADKSGNKTDEDLPSDALNTFH 1057
L++A P P P N T ++ + + V+ K +++ + N
Sbjct: 87 LDIAIPPP----RPKRKPNNPYPRKTGSGTILMSKTGVNDGKESLGSEKVSHPEMANEDR 142
Query: 1058 DESNPLEAT 1066
+S P E T
Sbjct: 143 QQSKPEEKT 151
Score = 32.3 bits (72), Expect = 2.3
Identities = 18/67 (26%), Positives = 32/67 (46%), Gaps = 8/67 (11%)
Query: 725 PLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSEC 784
P I K+R WT EE F+E +G+ ++KI + KT ++H +
Sbjct: 17 PYTITKQRER---WTEEEHNRFIEALRLYGRAWQKIEEHVATKTAVQ-----IRSHAQKF 68
Query: 785 FEKLKRK 791
F K++++
Sbjct: 69 FSKVEKE 75
>At1g61080 hypothetical protein
Length = 907
Score = 38.5 bits (88), Expect = 0.032
Identities = 55/236 (23%), Positives = 102/236 (42%), Gaps = 51/236 (21%)
Query: 212 EGLAKYEKKKVDV---------PDPGSNKDGSVSSAGNMEPCSSISPNLVDKSPKVTGFS 262
+GL K K++ D+ + S + G SS+ + P S S + KS + S
Sbjct: 215 DGLIKMSKERFDMMEIDEEEEKKESTSPQTGKTSSSRVLSPSESFSDS---KSSFGSRNS 271
Query: 263 DCASPATPSSVACSSSPGVDDKLLGKVGNADNDVSNLTDSPAPGFQNHLQKFYLNLDKLD 322
C SP TP SV S G G+VG+ N S+L + +Q L+KL
Sbjct: 272 FCGSPTTPRSVLPESMMGSP----GRVGDFANSASHLL------WNMRVQA----LEKLS 317
Query: 323 VDSLNSLGSSIVELVQSDDPS-SDDSGLVRSNAINKLLIWKADISKVLEMTESEIDLLEN 381
+ L I+ ++ +P+ S+D ++ + + ++ + EI+ ++
Sbjct: 318 PIDVKRLAIHILSQKEAQEPNESNDEDVI-------------SVVEEIKQKKDEIESIDV 364
Query: 382 ELKSLKSESVDRSECPVASGSQ---------QADSSSKF-YEERVEVSQKVIRPVP 427
++++ +S ++D E V +G Q ++ S SK + E+ E S ++ P P
Sbjct: 365 KMETEESVNLD-EESVVLNGEQDTIMKISSLESTSESKLNHSEKYENSSQLFPPPP 419
>At3g09600 putative MYB-related protein
Length = 125
Score = 38.1 bits (87), Expect = 0.042
Identities = 23/73 (31%), Positives = 35/73 (47%), Gaps = 8/73 (10%)
Query: 718 SNGLVEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYY 777
S+ V P I K R WT EE + FLE F +D++KI F+ KT
Sbjct: 29 SSKKVRKPYTITKSRES---WTEEEHDKFLEALQLFDRDWKKIEDFVGSKTV-----IQI 80
Query: 778 KNHKSECFEKLKR 790
++H + F K+++
Sbjct: 81 RSHAQKYFLKVQK 93
Score = 37.7 bits (86), Expect = 0.055
Identities = 17/48 (35%), Positives = 30/48 (62%), Gaps = 4/48 (8%)
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRK 994
WT++E FL+A+ F +D++KI VG+K + H +++F K +K
Sbjct: 46 WTEEEHDKFLEALQLFDRDWKKIEDFVGSKTVIQIRSHAQKYFLKVQK 93
>At5g52660 unknown protein
Length = 331
Score = 37.7 bits (86), Expect = 0.055
Identities = 21/74 (28%), Positives = 35/74 (46%), Gaps = 8/74 (10%)
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
+ P I K R WT E + FLE F +D++KI +F+ KT ++H
Sbjct: 62 IRKPYTITKSRES---WTEPEHDKFLEALQLFDRDWKKIEAFIGSKTV-----IQIRSHA 113
Query: 782 SECFEKLKRKDIGK 795
+ F K+++ G+
Sbjct: 114 QKYFLKVQKSGTGE 127
Score = 37.4 bits (85), Expect = 0.071
Identities = 20/60 (33%), Positives = 34/60 (56%), Gaps = 5/60 (8%)
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRKC-LGLNLANPVP 1005
WT+ E FL+A+ F +D++KI +G+K + H +++F K +K G +L P P
Sbjct: 75 WTEPEHDKFLEALQLFDRDWKKIEAFIGSKTVIQIRSHAQKYFLKVQKSGTGEHLPPPRP 134
>At5g37260 putative protein
Length = 287
Score = 37.7 bits (86), Expect = 0.055
Identities = 35/145 (24%), Positives = 63/145 (43%), Gaps = 26/145 (17%)
Query: 950 EWTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRKCLGL---NLAN 1002
+WT+ E F++A+ +G+ + +I VGTK + H ++FF+K + G+ ++
Sbjct: 38 KWTEAEHEKFVEALKLYGRAWRRIEEHVGTKTAVQIRSHAQKFFTKVARDFGVSSESIEI 97
Query: 1003 PVPGINGSPLNDDANGGESDTDDACVVEAGSVVDADKSGNK--TDED----------LPS 1050
P P P++ + +A +V A+ +G+K DED S
Sbjct: 98 PPPRPKRKPMHPYPR-------KLVIPDAKEMVYAELTGSKLIQDEDNRSPTSVLSAHGS 150
Query: 1051 DALNTFHDESNPLEATSLSAKLNES 1075
D L + S + LS+ ES
Sbjct: 151 DGLGSIGSNSPNSSSAELSSHTEES 175
Score = 32.3 bits (72), Expect = 2.3
Identities = 22/75 (29%), Positives = 34/75 (45%), Gaps = 9/75 (12%)
Query: 725 PLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSEC 784
P I K+R WT E E F+E +G+ +R+I + KT ++H +
Sbjct: 29 PYTITKQREK---WTEAEHEKFVEALKLYGRAWRRIEEHVGTKTAVQ-----IRSHAQKF 80
Query: 785 FEKLKRKDIGKLGKS 799
F K+ R D G +S
Sbjct: 81 FTKVAR-DFGVSSES 94
>At4g31570 putative protein
Length = 2712
Score = 37.4 bits (85), Expect = 0.071
Identities = 24/84 (28%), Positives = 43/84 (50%), Gaps = 2/84 (2%)
Query: 311 LQKFYLNLDKLDVDSLNS--LGSSIVELVQSDDPSSDDSGLVRSNAINKLLIWKADISKV 368
+QK Y +L KL +S S + S VE V DP D S A+ K+L + ++ V
Sbjct: 1163 MQKVYADLTKLITESCGSAEMTSLEVENVAVFDPFRDGSFENLLEAVRKILSERLELQSV 1222
Query: 369 LEMTESEIDLLENELKSLKSESVD 392
++ +S++ N+++ + S+D
Sbjct: 1223 IDKLQSDLSSKSNDMEEMTQRSLD 1246
>At2g47620 SWI/SNF family like transcription activator
Length = 512
Score = 37.0 bits (84), Expect = 0.093
Identities = 38/145 (26%), Positives = 64/145 (43%), Gaps = 13/145 (8%)
Query: 948 ATEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFFSKTRKC-LGLNLANPVPG 1006
A WT++E L++V G D+E IS+ V TK++ C SK + G L G
Sbjct: 225 AAVWTEEEILLLLESVLKHGDDWELISQSVSTKSRLDC---ISKLIELPFGEFLMGSASG 281
Query: 1007 -INGSPLNDDANGGESDTDDACVVEAGSVVDADKSGNKTDEDLPSD-----ALNTFHDES 1060
+N S L +D N + TD E ++ ++ +ED P AL + D S
Sbjct: 282 RLNPSILTEDENTEQVQTDGQ---EHEETETREEKEDRVNEDEPPAKRKRVALISEGDSS 338
Query: 1061 NPLEATSLSAKLNESREISGTEVCL 1085
+ ++++K+ S + + L
Sbjct: 339 LMKQVAAMASKVGPSVATAAAKAAL 363
>At1g18330 hypothetical protein
Length = 346
Score = 37.0 bits (84), Expect = 0.093
Identities = 15/46 (32%), Positives = 29/46 (62%), Gaps = 4/46 (8%)
Query: 950 EWTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSK 991
+W+++E FL+A+ +G+ + +I +GTK + H ++FFSK
Sbjct: 52 KWSEEEHDRFLEAIKLYGRGWRQIQEHIGTKTAVQIRSHAQKFFSK 97
Score = 33.5 bits (75), Expect = 1.0
Identities = 18/70 (25%), Positives = 33/70 (46%), Gaps = 8/70 (11%)
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
V P + K+R W+ EE + FLE +G+ +R+I + KT ++H
Sbjct: 40 VRKPYTVTKQREK---WSEEEHDRFLEAIKLYGRGWRQIQEHIGTKTAVQ-----IRSHA 91
Query: 782 SECFEKLKRK 791
+ F K+ ++
Sbjct: 92 QKFFSKMAQE 101
>At1g01520 hypothetical protein
Length = 287
Score = 37.0 bits (84), Expect = 0.093
Identities = 17/48 (35%), Positives = 29/48 (60%), Gaps = 4/48 (8%)
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRK 994
WT+ E FL+A+ F +D++KI VG+K + H +++F K +K
Sbjct: 64 WTEQEHDKFLEALHLFDRDWKKIKAFVGSKTVIQIRSHAQKYFLKVQK 111
Score = 36.2 bits (82), Expect = 0.16
Identities = 21/69 (30%), Positives = 34/69 (48%), Gaps = 8/69 (11%)
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
V P I K R WT +E + FLE F +D++KI +F+ KT ++H
Sbjct: 51 VRKPYTITKSREN---WTEQEHDKFLEALHLFDRDWKKIKAFVGSKTV-----IQIRSHA 102
Query: 782 SECFEKLKR 790
+ F K+++
Sbjct: 103 QKYFLKVQK 111
>At3g10590 hypothetical protein
Length = 206
Score = 36.2 bits (82), Expect = 0.16
Identities = 25/77 (32%), Positives = 37/77 (47%), Gaps = 5/77 (6%)
Query: 721 LVEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIAS--FLDHKTTADCIEF--Y 776
L E + +++ NPWT EE LFL+ +G+ + S F+ KT Y
Sbjct: 96 LAESSQSKRRKKDTPNPWTEEEHRLFLQGLKKYGEGASTLTSTNFVKTKTPRQVSSHAQY 155
Query: 777 YKNHKSECFEKLKRKDI 793
YK KS+ +K KR+ I
Sbjct: 156 YKRQKSD-NKKEKRRSI 171
>At4g26630 unknown protein
Length = 763
Score = 35.8 bits (81), Expect = 0.21
Identities = 52/236 (22%), Positives = 82/236 (34%), Gaps = 15/236 (6%)
Query: 938 ETCSDESCGEATEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFFSKTRKCLG 997
E DE EAT +D Q + + V +K + K +T++
Sbjct: 91 EVTKDEGQAEATNMDEDADGKKEQTDDGVSVEDTVMKENVESKDNNYAKDDEKETKETDI 150
Query: 998 LNLANPVPGINGSPLNDDANGGESDTDDACVVEAGSVVDADKSGNKTDEDLPSDALNTF- 1056
+ G D G D + + E G++VD DK G DE + + N
Sbjct: 151 TEADHKKAGKEDIQHEADKANGTKDGNTGDIKEEGTLVDEDK-GTDMDEKVENGDENKQV 209
Query: 1057 ----------HDESNPLEATSLSAKLNESR---EISGTEVCLENVDVASVACAINVESKL 1103
+E+ E + A+++ES+ E G+E +N V S + + +
Sbjct: 210 ENVEGKEKEDKEENKTKEVEAAKAEVDESKVEDEKEGSEDENDNEKVESKDAKEDEKEET 269
Query: 1104 GSDVSGVGLCTTDKSGSVNGVGLGGTVRESISASEIIKPRECGSVALDRTVSEGSS 1159
D + G GG VRE E+ K E + DR V E S
Sbjct: 270 NDDKEDEKEESKGSKKRGKGTSSGGKVREKNKTEEVKKDAEPRTPFSDRPVRERKS 325
>At3g18610 unknown protein
Length = 636
Score = 35.4 bits (80), Expect = 0.27
Identities = 81/340 (23%), Positives = 135/340 (38%), Gaps = 51/340 (15%)
Query: 157 SRSMAGTDSYEGKPNLKHKNVTAVESNSGEATACVTSSMPSEDATSRKKPRLNWGEGLAK 216
S S +DS E + + + ES+S E SS E A ++K+P
Sbjct: 64 SSSSDASDSDEEEKTKETPSKLKDESSSEEED---DSSSDEEIAPAKKRPE--------P 112
Query: 217 YEKKKVDVPDPGSNKDGSVSSAGNMEPCSSISPNLVDKSPKVTGFSDCASPATPSSVACS 276
+K KV+ S+ D +S P P +++K+ + SD S + +V
Sbjct: 113 IKKAKVE----SSSSDDDSTSDEETAPVKK-QPAVLEKAKVESSSSDDDSSSDEETVPVK 167
Query: 277 SSPGVDDKLLGKVGNADNDVSNLTDSPAPGFQNHLQKFYLNLDKLDVDSLNSLGSS---- 332
P V +K + ++D+D S+ + P ++K L+K +S +S S
Sbjct: 168 KQPAVLEKAKIESSSSDDD-SSSDEETVP-----MKKQTAVLEKAKAESSSSDDGSSSDE 221
Query: 333 ----------IVELVQSDDPSSD-DSGLVRSNAINKLLIWKADISKVLEMTESEIDLLEN 381
+V+ SD+ SSD ++ +V+ + KA+ S E + S ++
Sbjct: 222 EPTPAKKEPIVVKKDSSDESSSDEETPVVKKKPTTVVKDAKAESSSSEEESSS-----DD 276
Query: 382 ELKSLKSESVDRSECPVASGSQQADSSSKFYEERVEVSQKVIRPVPLKIISSDEPNTVKM 441
E K +V ++ P A DSSS + E S P +SS T K
Sbjct: 277 EPTPAKKPTVVKNAKPAAK-----DSSSSEEDSDEEESDDEKPPTKKAKVSS---KTSKQ 328
Query: 442 PQSTNLCSIHENDKEEDIDSPGSATSKFVEPLPVNAVSSS 481
S++ S E+DKEE D + K + V+A S
Sbjct: 329 ESSSDESS-DESDKEESKDEKVTPKKKDSDVEMVDAEQKS 367
>At3g28830 hypothetical protein
Length = 536
Score = 35.0 bits (79), Expect = 0.35
Identities = 65/339 (19%), Positives = 129/339 (37%), Gaps = 18/339 (5%)
Query: 145 SSRGSGFSHSSSSRSMAGTDSYEGKPNLKHKNVTAVESNSGEATACVTSSMPSEDATSRK 204
SS G + S SS + +G+ S G + K K ++ S S + + T S S ++
Sbjct: 190 SSIVGGAASSESSSTKSGSVSSSGSVSTKSKESSS--SGSSASGSVATKSKESSGGSAAT 247
Query: 205 KPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSAGNMEPCSSISPNLVDKSPKVTGFSDC 264
K + + G A K+ GS G S + + P +S S ++ KS S
Sbjct: 248 KSKESSGGSAATKSKES----SGGSATTGKTSGSPSGSPKASPSGSVSGKSSSKGSASAQ 303
Query: 265 ASPATPSSVACSSSPGVDDKLLGKVGNADNDVSNLTDSPAPGFQNHLQKFYLNLDKLDVD 324
S + S + S G + + S + + Q+ + +
Sbjct: 304 GSASAQGSASAQGSASAQ----GSASAQRRESGAMAMSKSRETKTSSQRQSKSSSE---S 356
Query: 325 SLNSLGSSIVELVQSDDPSSDDSGLVRSNAINKLLIWKADISKVLEMTESEIDLLENELK 384
S +S ++ V+ V+S+ S +++ + K KA++ E +S + +
Sbjct: 357 SSSSTTTTTVKQVESETSKEVMSFIMQ---LEKKYAAKAELKVFFESLKSSMQASASVGS 413
Query: 385 SLKSESVDRSECPVA--SGSQQADSSSKFYEERVEVSQKVIRPVPLKIISSDEPNTVKMP 442
+ V S+ S + + SS +++ + + LK + + KM
Sbjct: 414 KTAKDYVSASKAATGKLSEAMASVSSKNVKSAKMKSNLDTSKDEMLKCVKQIQDINGKMV 473
Query: 443 QSTNLCSIHENDKEEDIDSPGSATSKFVEPLPVNAVSSS 481
+ S +++ ++ I T++FVE ++ SSS
Sbjct: 474 SGKTVSSTQQSELKQTITKWEKVTTQFVETAASSSSSSS 512
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.310 0.128 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,342,919
Number of Sequences: 26719
Number of extensions: 1782664
Number of successful extensions: 4851
Number of sequences better than 10.0: 80
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 58
Number of HSP's that attempted gapping in prelim test: 4729
Number of HSP's gapped (non-prelim): 169
length of query: 1620
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1507
effective length of database: 8,299,349
effective search space: 12507118943
effective search space used: 12507118943
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC126015.12


