

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126015.11 + phase: 0
(371 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
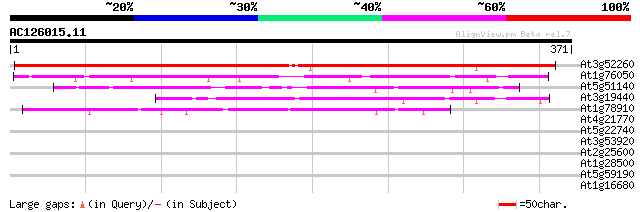
Score E
Sequences producing significant alignments: (bits) Value
At3g52260 unknown protein 487 e-138
At1g76050 putative RNA pseudouridylate synthase 92 5e-19
At5g51140 unknown protein 91 8e-19
At3g19440 unknown protein 73 3e-13
At1g78910 unknown protein 44 2e-04
At4g21770 unknown protein (At4g21770) 37 0.017
At5g22740 glucosyltransferase-like protein 31 1.2
At3g53920 sigma factor SigC 28 6.2
At2g25600 potassium transporter/channel like protein 28 6.2
At1g28500 hypothetical protein 28 6.2
At5g59190 cucumisin precursor - like 28 8.1
At1g16680 hypothetical protein 28 8.1
>At3g52260 unknown protein
Length = 369
Score = 487 bits (1254), Expect = e-138
Identities = 240/366 (65%), Positives = 300/366 (81%), Gaps = 10/366 (2%)
Query: 4 PKKIGIPWPEFNDGLSYDDVVRPSDAGLTLI-EFYSTKYKSSAPLQGWLQRIKSDQITLD 62
P++IG+PWPE NDGL+Y D + S++ LT + EFY TKYKSSAPL GW+QRI++ QI +D
Sbjct: 5 PQRIGLPWPELNDGLAYKDAISSSESELTTVSEFYFTKYKSSAPLLGWIQRIQNGQIQID 64
Query: 63 ARVVTDPNTILRVGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQ 122
VV DPNT+LR GSKLVY R+PWKEPD P+++EVLYEDDD+IALNKPSGLQVLPGGLFQ
Sbjct: 65 GEVVKDPNTLLRSGSKLVYSRLPWKEPDTPYSLEVLYEDDDLIALNKPSGLQVLPGGLFQ 124
Query: 123 QRTVLTQLQW-ETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLASYFADG 181
QRTVLTQLQW N S I + + PHPVPVHRLGRGTSGILLCAKTKLA+ +LA+YFA+G
Sbjct: 125 QRTVLTQLQWCFGKNDSYIGSRESPHPVPVHRLGRGTSGILLCAKTKLAKTKLAAYFAEG 184
Query: 182 TSRIEGKGRHTNQELG---KIAKIYRALVSGIINNDQVTINQPIGMVKYPGVAKGLYVAS 238
TS + G G + +QE G K++KIYRAL GI+ D+V I QPIG+V+YPGVA+GLYVAS
Sbjct: 185 TSLV-GSG-NLDQECGTGRKLSKIYRALADGIVEEDEVVIKQPIGVVRYPGVAQGLYVAS 242
Query: 239 QSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPK 298
GKPA S V VLER+ ++N +LV+V+IQSGRPHQIRIHL+++GHPL+GDPLY GGQPK
Sbjct: 243 PEGKPAFSKVFVLERDREKNCSLVKVEIQSGRPHQIRIHLAYMGHPLVGDPLYVAGGQPK 302
Query: 299 CLDCDFLDE--SFAEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPI-TNEIIEMIAPLPSI 355
C D D +D+ +FAEDGGY RP++ VPGDCGYHLHAH++ L + + T+++++++APLP I
Sbjct: 303 CFDPDLVDDAAAFAEDGGYRRPNQAVPGDCGYHLHAHQVELPNLLNTHKVVKIVAPLPPI 362
Query: 356 LQTHEE 361
LQT E
Sbjct: 363 LQTRCE 368
>At1g76050 putative RNA pseudouridylate synthase
Length = 430
Score = 92.0 bits (227), Expect = 5e-19
Identities = 100/399 (25%), Positives = 171/399 (42%), Gaps = 80/399 (20%)
Query: 3 SPKKIGIPWPEFNDGLSYDDVVRPSDAGLTLIEFYSTKYK--SSAPLQGWLQRIKSDQIT 60
SP +I IP N GL +++V + + L + S++ S A +Q I+ +T
Sbjct: 55 SPTEIIIPRVN-NAGLRIEEIVDAAKGKIRLDSWISSRINGVSRARVQS---SIRLGLVT 110
Query: 61 LDARVVTDPNTILRVGSKL---VYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLP 117
++ RVV + ++ G ++ + P K ++++YED ++ +NKP+ + V P
Sbjct: 111 VNGRVVDKVSHNVKSGDEVNCTISELQPLKAEAEDIPLDIVYEDKHVLVVNKPAHMVVHP 170
Query: 118 GGLFQQRTVLTQL---------------QWETNNKSTIEAHQKPHPVP-----------V 151
T++ + + + +++ T ++ P V
Sbjct: 171 APGNPTGTLVNGILHHCSLPCVDYSNSEEDDDSDEETFSDDEEMTTSPSSYAASVRPGIV 230
Query: 152 HRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGII 211
HRL +GT+G+L+ AK + + A LA F +L I ++Y +L +G+
Sbjct: 231 HRLDKGTTGLLVVAKDEHSHAHLAEQF----------------KLHTIERVYVSLTTGVP 274
Query: 212 NNDQVTINQPIG--------MVKYPGVAKGLYVASQSGKPALSVVHVLERNVKENSTLVQ 263
+ Q I PIG M PG +G + A S V+E S LV+
Sbjct: 275 SPPQGRIEIPIGRDSSNRIRMAAIPGGVRG-----GRARHAASRYKVIETFAGGGSALVE 329
Query: 264 VKIQSGRPHQIRIHLSFIGHPLLGDPLYAVGGQPKCLDCDFLDESFAEDGG------YTR 317
++++GR HQIR H ++G PLLGD +Y G K + L + + +R
Sbjct: 330 WRLETGRTHQIRAHAKYMGVPLLGDEVY---GGTKSMALSLLQKRVSRSDQEEIIELISR 386
Query: 318 PSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLPSIL 356
+P LHA L +HP T EI++ P PS L
Sbjct: 387 MDRPC-------LHAIVLGFTHPCTGEIVKFSCPPPSDL 418
>At5g51140 unknown protein
Length = 395
Score = 91.3 bits (225), Expect = 8e-19
Identities = 87/333 (26%), Positives = 148/333 (44%), Gaps = 57/333 (17%)
Query: 30 GLTLIEFYSTKYKSSAPLQGWLQRIKSDQITLDARVVTDPNTILRVGSKLVYHRIPWKEP 89
G T+++ ++ ++K P ++ +KS +I +D +V P + + S+ + H + EP
Sbjct: 73 GKTIVDLFADEFKGR-PRDYYVGAVKSGRIKVDGEIV--PVSYIVKSSQKITHFLHRHEP 129
Query: 90 DAP-HTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHP 148
+ +L+++ D++ + KP+ + V P G +++ T++ L E H
Sbjct: 130 PVMIDDVVILHQEPDVVTVCKPASVPVHPCGQYRKNTIVGILDAE---------HDLGTL 180
Query: 149 VPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVS 208
P+HRL R SG+L+ A+T A A +F +IEG G + K Y A V
Sbjct: 181 FPIHRLDRLVSGLLIIART----AAKADFFRQ---QIEG---------GMVKKRYIAKVI 224
Query: 209 GIINNDQVTINQPIGMVKYPGVAKGLYVASQS------GKPALSVVHVLERNVKENSTLV 262
G+ D++ ++ I G + S GKPA + ++ N +LV
Sbjct: 225 GVFPEDEMIVDANINYNGSEGRSTAEDANSSGDDKKVKGKPACTKFTRIDTN--GTHSLV 282
Query: 263 QVKIQSGRPHQIRIHLSFIGHPLLGDPLY---------------AVGGQPKCLDCD---F 304
+ +GR HQIR+HL + GHP+ DPLY G+ K + D +
Sbjct: 283 SCEPVTGRTHQIRVHLQYTGHPIANDPLYLNQHIDNLETYIAKRIDAGERKIVSPDDYVY 342
Query: 305 LDESFAEDGGYTRPSKPVPGDCGYHLHAHRLFL 337
E F+ D T K +P GY H L+L
Sbjct: 343 SSEDFSIDPMCTNCPKLIPQ--GYEEHDEALWL 373
>At3g19440 unknown protein
Length = 477
Score = 72.8 bits (177), Expect = 3e-13
Identities = 78/295 (26%), Positives = 124/295 (41%), Gaps = 52/295 (17%)
Query: 97 VLYEDDDIIALNKPSGLQVLPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHPVPVHRLGR 156
VLY+D II LNKP GL V G +T + +L S ++ + P VHRL R
Sbjct: 185 VLYKDPAIIVLNKPHGLAVQGGSGI--KTSIDELA-----ASCLKFDKSESPRLVHRLDR 237
Query: 157 GTSGILLCAKTKLARARLASYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGIINNDQV 216
SG+L+ A+T+ A L S F + T+ G + + + + Y ALV G +
Sbjct: 238 DCSGLLVLARTQTAATVLHSIFREKTTGASAYG--VKKNVKSLKRKYMALVIGCPPRQRG 295
Query: 217 TINQPIGMVKYPGVAKGLYVASQSGKPALSVVHVLERNVKENS----TLVQVKIQSGRPH 272
I+ P+ V + +G+ + + E V E+S T ++++ +GR H
Sbjct: 296 QISAPLRKVVVDDGKSERITVNDNGELVSTQHAITEYRVIESSPHGYTWLELRPLTGRKH 355
Query: 273 QIRIHLS-FIGHPLLGDPLYAVGGQPKCLDCDFLDE------------SFAEDGGYTRPS 319
Q+R+H + +G P++GD Y G Q F+ DGG
Sbjct: 356 QLRVHCAEVLGTPIVGD--YKYGWQAHKAREPFVSSENNPTKQSSSPFGLDLDGGDVSSK 413
Query: 320 KPVPGDCGYHLHAHRLFLSHPITNEIIEMI-----------------APLPSILQ 357
+P HLH H + P ++++E + APLPS +Q
Sbjct: 414 QP-------HLHLHSKQIDLPNISQLLEKMQVSSDSDISDLDSLKFDAPLPSHMQ 461
>At1g78910 unknown protein
Length = 491
Score = 43.5 bits (101), Expect = 2e-04
Identities = 75/308 (24%), Positives = 127/308 (40%), Gaps = 31/308 (10%)
Query: 9 IPWPEFNDGLSYDDVVRPS-DAGLTLIEFYSTKYKSSAPLQGWL--QRIKSDQITLDARV 65
I W ++ YD ++ GL +EF + S G + ++IK +++
Sbjct: 84 ISWVKYYFEEIYDKAIQTHFTKGLVQMEFRGRRDASREKEDGAIPMRKIKHNEVMQIGDK 143
Query: 66 VTDPNTILRVGSKLVYHRIPWKEPDAPHTIEVLY-------EDDDIIALNKPSGLQV--- 115
+ P +I + Y IP P+ E+ Y +D II LNKP L V
Sbjct: 144 IWLPVSIAEMRISKRYDTIP-SGTLYPNADEIAYLQRLVRFKDSAIIVLNKPPKLPVKGN 202
Query: 116 LPGGLFQQRTVLTQLQWETNNKSTIEAHQKPHPVPVHRLGRGTSGILLCAKTKLARARLA 175
+P L + + + K + VHRL R TSG+L+ +TK + L
Sbjct: 203 VPIHNSMDALAAAALSFGNDEGPRLV---KLTFLGVHRLDRETSGLLVMGRTKESIDYLH 259
Query: 176 SYFADGTSRIEGKGRHTNQELGKIAKIYRALVSGIINNDQVTINQPIGMVKY-PGVAKGL 234
S F+D R + N+ + + Y ALV G + I+ P+ V G +
Sbjct: 260 SVFSDYKGR-NSSCKAWNKACEAMYQQYWALVIGSPKEKEGLISAPLSKVLLDDGKTDRV 318
Query: 235 YVASQSG----KPALSVVHVLERNVKENSTLVQVKIQSGRPH------QIRIHLS-FIGH 283
+A SG + A++ VL + + V+++ + R H Q+R+H + +G
Sbjct: 319 VLAQGSGFEASQDAITEYKVLGPKI-NGCSWVELRPITSRKHQPPSKKQLRVHCAEALGT 377
Query: 284 PLLGDPLY 291
P++GD Y
Sbjct: 378 PIVGDYKY 385
>At4g21770 unknown protein (At4g21770)
Length = 472
Score = 37.0 bits (84), Expect = 0.017
Identities = 59/247 (23%), Positives = 94/247 (37%), Gaps = 47/247 (19%)
Query: 66 VTDPNTILRVGSKLVYHRIPWKEP---DAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQ 122
VT N G+ L H P + P + ++ D + L+KP+G V GG
Sbjct: 177 VTHTNQYAEAGTYLRVHVHPKRSPRCYEIDWKSRIVAVTDSYVILDKPAGTTV--GGT-- 232
Query: 123 QRTVLTQLQWETNNKSTIEAHQKPHPVPV-HRLGRGTSGILLCAKTKLARARLASYFADG 181
T E+ A P P+ H++ T G ++ A+TK S F
Sbjct: 233 -----TDNIEESCATFASRALDLPEPLKTTHQIDNCTEGCVVFARTK----EYCSVF--- 280
Query: 182 TSRIEGKGRHTNQELGKIAKIYRALVS-----GIINNDQVTINQPIGMVKYPGVAKGLYV 236
HT ++ K+YRAL + GII++ N +V + +G ++
Sbjct: 281 ---------HTKIRNKEVKKLYRALAAAPLPIGIISHYMRPKNMAPRLVAEDSI-QGWHL 330
Query: 237 ASQS---------GKPALSVVHVLER---NVKENSTLVQVKIQSGRPHQIRIHLSFIGHP 284
A H +E KE + + + +G+ HQIR L+ G P
Sbjct: 331 CQLEVLECKKIPWPDAATEKKHDIEDCGWTSKEFAYECTINLLTGKTHQIRAQLAACGAP 390
Query: 285 LLGDPLY 291
L+GD +Y
Sbjct: 391 LVGDSMY 397
>At5g22740 glucosyltransferase-like protein
Length = 534
Score = 30.8 bits (68), Expect = 1.2
Identities = 15/52 (28%), Positives = 31/52 (58%), Gaps = 2/52 (3%)
Query: 75 VGSKLVYHRIPWKEPDAPHTIEVLYEDDDIIALNKPSGLQVLPGGLFQQRTV 126
+G +V ++ WK+PD + E +++D+++ + N P L +P +F +R V
Sbjct: 62 MGIVIVLVKLFWKKPDKRYKFEPIHDDEELGSSNFPVVLVQIP--MFNEREV 111
>At3g53920 sigma factor SigC
Length = 571
Score = 28.5 bits (62), Expect = 6.2
Identities = 17/61 (27%), Positives = 35/61 (56%), Gaps = 3/61 (4%)
Query: 17 GLSYDDVVRPSDAGLT--LIEFYSTK-YKSSAPLQGWLQRIKSDQITLDARVVTDPNTIL 73
G++++D+++ G+ F T+ YK S +Q W+++ S ++ AR V P++I+
Sbjct: 355 GIAHEDLIQAGYVGVLQGAERFDHTRGYKFSTYVQYWIRKSMSTMVSRHARGVHIPSSII 414
Query: 74 R 74
R
Sbjct: 415 R 415
>At2g25600 potassium transporter/channel like protein
Length = 888
Score = 28.5 bits (62), Expect = 6.2
Identities = 15/48 (31%), Positives = 24/48 (49%)
Query: 310 AEDGGYTRPSKPVPGDCGYHLHAHRLFLSHPITNEIIEMIAPLPSILQ 357
A+DGG S VPG G ++ R+ +S P E + LP+ ++
Sbjct: 798 ADDGGEVPRSPAVPGGGGSMIYPERVTISSPENGETGGKVVLLPNSME 845
>At1g28500 hypothetical protein
Length = 326
Score = 28.5 bits (62), Expect = 6.2
Identities = 12/53 (22%), Positives = 30/53 (55%), Gaps = 3/53 (5%)
Query: 232 KGLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGHP 284
+G++ +Q+G P + ++ V+ +EN + ++ + R H ++++F G P
Sbjct: 214 RGMFGVAQTGLPQVQILKVVIETEEENEKPLDKRLNARRAH---VYITFTGLP 263
>At5g59190 cucumisin precursor - like
Length = 693
Score = 28.1 bits (61), Expect = 8.1
Identities = 16/45 (35%), Positives = 18/45 (39%)
Query: 148 PVPVHRLGRGTSGILLCAKTKLARARLASYFADGTSRIEGKGRHT 192
P P G G+ KL AR + FAD EG G HT
Sbjct: 120 PPPKKWKGSCKGGLKFACNNKLIGARFYNKFADSARDEEGHGTHT 164
>At1g16680 hypothetical protein
Length = 496
Score = 28.1 bits (61), Expect = 8.1
Identities = 26/90 (28%), Positives = 41/90 (44%), Gaps = 9/90 (10%)
Query: 232 KGLYVASQSGKPALSVVHVLERNVKENSTLVQVKIQSGRPHQIRIHLSFIGH-PLLGDPL 290
K ++ S KPA S V N+KE ST+ VKI++ +++ L + H LG PL
Sbjct: 185 KKVHEDKSSTKPASSSTVV--SNMKEISTVKVVKIETDSADEMKRILDSLNHYEALGLPL 242
Query: 291 YAVGGQPKCLDCDFLDESFAEDGGYTRPSK 320
+ K +D L + + + P K
Sbjct: 243 F------KKIDAALLKKDYRKKAMLVHPDK 266
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.137 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,673,428
Number of Sequences: 26719
Number of extensions: 391634
Number of successful extensions: 827
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 804
Number of HSP's gapped (non-prelim): 14
length of query: 371
length of database: 11,318,596
effective HSP length: 101
effective length of query: 270
effective length of database: 8,619,977
effective search space: 2327393790
effective search space used: 2327393790
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126015.11


