

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
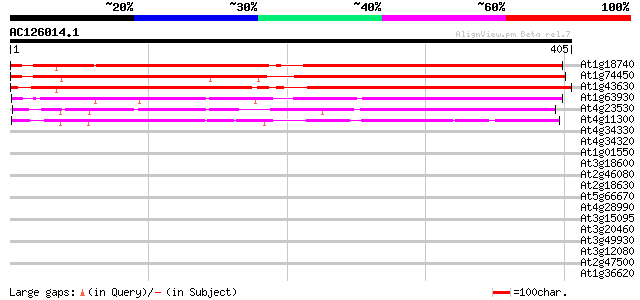
Query= AC126014.1 + phase: 0
(405 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g18740 unknown protein 479 e-136
At1g74450 unknown protein 474 e-134
At1g43630 unknown protein 461 e-130
At1g63930 hypothetical protein 268 4e-72
At4g23530 unknown protein (At4g23530) 227 1e-59
At4g11300 unknown protein 220 9e-58
At4g34330 putative protein 40 0.002
At4g34320 unknown protein 38 0.009
At1g01550 Unknown protein (F22L4.9) 37 0.019
At3g18600 DEAD box helicase protein, putative 36 0.043
At2g46080 unknown protein 35 0.056
At2g18630 unknown protein 33 0.28
At5g66670 At14a protein-like 32 0.48
At4g28990 unknown protein 32 0.48
At3g15095 unknown protein, 3' partial 32 0.48
At3g20460 sugar transporter, putative 32 0.62
At3g49930 zinc-finger-like protein 31 1.4
At3g12080 unknown protein 31 1.4
At2g47500 kinesin heavy chain like protein 31 1.4
At1g36620 hypothetical protein 30 1.8
>At1g18740 unknown protein
Length = 382
Score = 479 bits (1234), Expect = e-136
Identities = 243/410 (59%), Positives = 315/410 (76%), Gaps = 39/410 (9%)
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQIHSM----------EGSTLEIELDSFQQHVTD 50
MPATD+QGS FGRS+LS RR+Q+ S E ST+E+ELDSFQ+ V +
Sbjct: 1 MPATDFQGS-------FGRSLLSLRRDQVDSSTVVSGSSSHHEPSTMEVELDSFQRQVAE 53
Query: 51 RFVDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERS 110
+F+DL++ +D LLSL+W+GK+LD FL CQEEF+AI+ H+ Q+ + P+DR++S+YFERS
Sbjct: 54 KFIDLNASSND-LLSLEWIGKLLDSFLCCQEEFRAIVFNHRSQISKSPMDRLISDYFERS 112
Query: 111 VKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDG 170
+KALDVCNAIRDG+EQIR W+KL +IV+ ALD R IGEGQ RRAKKALIDL+I MLD+
Sbjct: 113 IKALDVCNAIRDGIEQIRQWEKLADIVISALDSHRPIGEGQLRRAKKALIDLAIGMLDEK 172
Query: 171 KE-SNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTW 229
S ++AHRNRSFGR +D H HRS+G FRSLSWSVSR+W
Sbjct: 173 DHPSGTNLAHRNRSFGRV-----KDSH---------------HRSIGHFRSLSWSVSRSW 212
Query: 230 SAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHF 289
SA++QLQA+ +N+ PR ND++A+NGLA+ VYTM SVLLFVMW LVAAIPCQDRGL V+F
Sbjct: 213 SASKQLQALASNLATPRPNDVVASNGLAVPVYTMTSVLLFVMWVLVAAIPCQDRGLQVNF 272
Query: 290 SIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFPLT 349
+PR + WA P++ LH++I+EESK+RDRKN CGLL+EI +IEK R+M++L+DS HFPL
Sbjct: 273 FVPRHFQWAAPVMSLHDKIVEESKRRDRKNCCGLLKEIDRIEKSSRLMNELIDSIHFPLN 332
Query: 350 EEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSL 399
++KE EV+Q+V E+ +V +AL++GLDP ER+VREVFHRIVRS+TE LDSL
Sbjct: 333 DDKEVEVKQRVDELVQVREALRNGLDPFERKVREVFHRIVRSRTESLDSL 382
>At1g74450 unknown protein
Length = 397
Score = 474 bits (1220), Expect = e-134
Identities = 245/418 (58%), Positives = 317/418 (75%), Gaps = 43/418 (10%)
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQ-IHSMEGST-------LEIELDSFQQHVTDRF 52
MPAT+YQ S FGRS L+ RR+ ++S+E +T +E EL SFQ+ V +RF
Sbjct: 1 MPATEYQRS-------FGRSFLNLRRDTAVNSVESTTVTPELTQMEAELVSFQRKVAERF 53
Query: 53 VDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVK 112
+DL++ ++LLSL+WVGK+LD FL CQEEF++I+ H+ + +PP+DR+VS+YFERSVK
Sbjct: 54 IDLNASSCEDLLSLEWVGKLLDSFLSCQEEFRSIVINHRSMITKPPMDRLVSDYFERSVK 113
Query: 113 ALDVCNAIRDGVEQIRVWQKLLEIVLYALDH-------QRSIGEGQFRRAKKALIDLSIS 165
ALDVCNAIRDGVEQIR WQKL+EIV+ A ++ +R +GEGQFRRA+K LI+L+I
Sbjct: 114 ALDVCNAIRDGVEQIRQWQKLIEIVICAFNNNGGGSSGKRPLGEGQFRRARKTLIELAIG 173
Query: 166 MLDDGKESNASVA--HRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSW 223
MLD+ S++SV+ HRNRSFGR N HR++G FRSLSW
Sbjct: 174 MLDEKDSSSSSVSSQHRNRSFGR-------------------NKEQLHHRTIGHFRSLSW 214
Query: 224 SVSRTWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDR 283
SVSR+WSA++QLQAIGNN+ PRA+D+ ATNGL + VYTM +VLLFVMWALVAAIPCQDR
Sbjct: 215 SVSRSWSASKQLQAIGNNLATPRASDITATNGLIVPVYTMTTVLLFVMWALVAAIPCQDR 274
Query: 284 GLNVHFSIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDS 343
GL VHF++PR+Y W L+ LH+RI+EESKKR+RKN CGLL+EI Q EK R+M++LVDS
Sbjct: 275 GLQVHFNVPRNYQWGGSLMSLHDRIIEESKKRERKNTCGLLKEIHQFEKTSRLMNELVDS 334
Query: 344 AHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGR 401
FPL+EEKE EVR++V E+ K+ +ALK+GLDP ER+VREVFHRIVRS+TEGLD++G+
Sbjct: 335 VQFPLSEEKEMEVRERVEELGKLQEALKNGLDPFERKVREVFHRIVRSRTEGLDTVGK 392
>At1g43630 unknown protein
Length = 383
Score = 461 bits (1187), Expect = e-130
Identities = 238/415 (57%), Positives = 311/415 (74%), Gaps = 43/415 (10%)
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQIHSM------EGSTLEIELDSFQQHVTDRFVD 54
MP T+Y +FGRS LS RR+Q H M E T+E+ELDSFQ+ V ++F+D
Sbjct: 1 MPETEY---------SFGRSFLSLRRDQAHLMDPTSFSEPMTMEVELDSFQRQVAEKFID 51
Query: 55 LS-SVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKA 113
L+ S E+LSL+W+GK+LD FL CQE+F+ I+ HK Q+++ P+DR++ EYFERSVKA
Sbjct: 52 LNASADEAEILSLEWIGKLLDSFLCCQEDFRVIIFNHKPQLLKQPMDRLIEEYFERSVKA 111
Query: 114 LDVCNAIRDGVEQIRVWQKLLEIVLYALD-HQRSIGEGQFRRAKKALIDLSISMLDDGKE 172
LDVCNAIRDG+EQIR WQKL+EIV+ ALD +QR +GEG+ RAKKALIDL+I MLD+
Sbjct: 112 LDVCNAIRDGIEQIRQWQKLIEIVISALDTNQRQLGEGEIHRAKKALIDLAIGMLDEKDS 171
Query: 173 SNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWSAA 232
SN HRNRSF R+ +DH+QH +G RSLSWSVSR+WSA+
Sbjct: 172 SNT---HRNRSFTRN-----KDHNQH----------------IGYIRSLSWSVSRSWSAS 207
Query: 233 RQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHFSIP 292
RQLQ IGNN+ PRA+D+MATNGLA++VYTM S+LLFV W LVAAIPCQDRGL+VHF P
Sbjct: 208 RQLQGIGNNLATPRASDVMATNGLALTVYTMTSILLFVTWVLVAAIPCQDRGLHVHFYFP 267
Query: 293 RSYTWAIPLLLLHERIMEESKKRD-RKNACGLLREIQQIEKCVRVMSDLVDSAHFPLTEE 351
R + WA+P++ LH++IM+ESKKRD +K CGLLREI QIE+ R++SDL+DS +F LT+E
Sbjct: 268 RHFQWAVPVMSLHDKIMDESKKRDKKKKGCGLLREINQIERNSRMLSDLIDSDNFSLTDE 327
Query: 352 K-EGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGRPNNE 405
K EV+++V E+ VC+A+K+GLDP +R+VR+VFH+IVR++TE LDSLG+ N+
Sbjct: 328 KCTLEVKERVQELMNVCEAIKEGLDPFDRKVRDVFHQIVRTRTEALDSLGKLQNQ 382
>At1g63930 hypothetical protein
Length = 415
Score = 268 bits (685), Expect = 4e-72
Identities = 153/428 (35%), Positives = 250/428 (57%), Gaps = 56/428 (13%)
Query: 2 PATDYQGSSPSSLTNFGRSILSFRREQIHSMEGSTLEIELDSFQQHVTDRFVDLSSVPH- 60
PA D QGS GR +S RR Q + + +L+ FQ+H+ DRF +L S P
Sbjct: 3 PAQDNQGSF------LGR--ISIRRNQFVDVNNEQEQEDLELFQKHIADRFTELLSPPQP 54
Query: 61 ---------------DELLSLKWVGKMLDCFLICQEEFKAILHTHKG--QVVRPPLDRMV 103
++++S+ W+ K++D FL C+ EFKAIL + Q+ +PP DR+V
Sbjct: 55 PPSDEINTVASVAATEQIMSVTWLRKLMDVFLCCEAEFKAILLMGRDPTQISKPPFDRLV 114
Query: 104 SEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLS 163
E +RS+KALD+C A+ +G++ +R +Q+L EI + AL+ QR +G+G RRAK+AL +L
Sbjct: 115 PEMLDRSIKALDICTAVVNGIDSVRHYQRLAEIAVTALE-QRPLGDGNVRRAKRALANLV 173
Query: 164 ISMLDDGKESNAS----------VAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHR 213
+++ + KE+ + R+ SFGR +GG ++ +
Sbjct: 174 VALSLEDKENVSGGGGGGGGGNKTTERSWSFGRRSGG--------------SSAASKGGA 219
Query: 214 SMGQFRSLSWSVSRTWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWA 273
++GQ +S SW+V R WSAA+Q+ A+ N+ PPR N+ GL ++ M++V++FVMW
Sbjct: 220 TIGQLKSSSWAVGRNWSAAKQIHAMTANLTPPRGNEAA---GLPQPMFIMSTVMVFVMWV 276
Query: 274 LVAAIPCQDR-GLNVHFSIPRSY-TWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIE 331
L AA+PCQ+R GL H +P + WA L+ +HE+I +E KK+++K + GL+ E+ ++E
Sbjct: 277 LTAAVPCQERSGLANHLPVPPKHLNWAQSLIGIHEKIGDEWKKKEKKGSAGLMEEMTRME 336
Query: 332 KCVRVMSDLVDSAHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRS 391
K + + D H+P ++ +V E++++C +++ L PL++Q+REVFHRIVRS
Sbjct: 337 KLGHSLMEFADGFHYPAEKDAAESAAVQVAEMAEICRRMEEELVPLQQQIREVFHRIVRS 396
Query: 392 KTEGLDSL 399
+ E L+ L
Sbjct: 397 RAEILEVL 404
>At4g23530 unknown protein (At4g23530)
Length = 396
Score = 227 bits (578), Expect = 1e-59
Identities = 151/418 (36%), Positives = 233/418 (55%), Gaps = 62/418 (14%)
Query: 3 ATDYQGSSPSSLTNFGRSILSFRREQIHSMEGS---TLEIELDSFQQHVTDRFVDLS--- 56
AT++QGS S + S RR QI SM+ + LE EL+ FQ+HV +RF DL
Sbjct: 4 ATEFQGSFLSRI--------SIRRNQIVSMDVNHEQELE-ELEYFQKHVAERFSDLITSP 54
Query: 57 -----------SVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPP-LDRMVS 104
S P D +LS+ W+ +LD F+ C+ EFKA+L T Q+ + P L+R++
Sbjct: 55 SPPPSSSSSAVSQPSDPILSIPWLQNLLDVFMSCEAEFKAVLSTT--QISKSPSLERVLP 112
Query: 105 EYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSI 164
E +R +KALD+CNA+ +G++ +R ++ EI + AL QR + +G RRAK+AL L I
Sbjct: 113 EMLDRILKALDLCNAVVNGIDSVRQSRRFAEIAVTALK-QRPLCDGSVRRAKRALTSLLI 171
Query: 165 SMLDDGKESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWS 224
+ N RD + G+ SN T + S G +++
Sbjct: 172 GL---------------------NADERRDRNSGGSGCSNQRRTTSRSWSFGTRSNVTGG 210
Query: 225 ------VSRTWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAI 278
VS+ WSA++Q+QA+ N+ PR + +G AM VY M+SV++ VMW LVAA+
Sbjct: 211 GLYGQVVSKNWSASKQIQAMVANLVLPRGAE---ASGPAMPVYIMSSVMVLVMWVLVAAV 267
Query: 279 PCQDRGLNVH-FSIPRSYTWAIPLLLLHERIMEESKKRDRK-NACGLLREIQQIEKCVRV 336
PCQ + V +P+ WA + + ERI EE K+++++ GL+ E+Q++EK
Sbjct: 268 PCQTSSVLVAPLPLPKHQNWASAAMSIQERIGEEIKRKEKRCGGGGLMEEMQRMEKIGLS 327
Query: 337 MSDLVDSAHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTE 394
+ + + FP EE+E EV +KV E+ ++C ++ GL+ L+RQVR+VFHR+VRS+ E
Sbjct: 328 LMEFAERFRFPADEEEEVEVAEKVDEMEEICRRMEVGLEDLQRQVRQVFHRLVRSRIE 385
>At4g11300 unknown protein
Length = 371
Score = 220 bits (561), Expect = 9e-58
Identities = 145/411 (35%), Positives = 230/411 (55%), Gaps = 61/411 (14%)
Query: 2 PATDYQGSSPSSLTNFGRSILSFRREQIHSMEGS--TLEIELDSFQQHVTDRFVDL---S 56
PAT++Q S S L+ RR Q+ SME + + EL+ FQ+HV +RF +L S
Sbjct: 3 PATEHQSSFLSRLS---------RRNQVVSMEVNHEQEQEELEDFQKHVAERFAELLPPS 53
Query: 57 SVPHD-ELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALD 115
P +LS++W+ K+LD F+ + EF ++L ++ Q+ +PPLD++V E +R VKALD
Sbjct: 54 DSPESYPILSIQWLRKLLDVFMSIESEFHSVLTSNPSQISKPPLDKLVPEMLDRIVKALD 113
Query: 116 VCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDGKESNA 175
+C A+ +GV+ +R Q+ EI + AL Q + +G RRAK+AL L ++ L+ K S +
Sbjct: 114 ICTAVVNGVDSVRQIQRCAEIAVTAL-KQTPLSDGSVRRAKRALTSL-LAALNADKNSGS 171
Query: 176 SVAHRNR--------SFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSR 227
S R SFGR +GG S G + VS+
Sbjct: 172 SGGGSGRRSSTDQWSSFGRRSGG-----------------------SSGGGGGGAGCVSK 208
Query: 228 TWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQ-DRGLN 286
WSAA+Q+QA+ N+ PR G A +Y M+SV++ VMW LV A+PCQ GL
Sbjct: 209 NWSAAKQIQAMTANLVAPR-------GGEASPMYIMSSVMVMVMWTLVVAVPCQTSNGLM 261
Query: 287 VHFSIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHF 346
VH +P++ WA + + ER+ EE K+++ + GL+ E+Q++E+ + + + F
Sbjct: 262 VHVPLPKNQVWANAAVSISERVGEEMKRKETRGG-GLMEEMQRMERIGLKLMEFSEGFRF 320
Query: 347 PLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLD 397
E +V +V E+ ++C ++DGL+ L+R+VREVFHR+V+S++E L+
Sbjct: 321 ----NGEEDVVAEVAEMEEICRKMEDGLEGLQRRVREVFHRLVKSRSEILE 367
>At4g34330 putative protein
Length = 354
Score = 40.4 bits (93), Expect = 0.002
Identities = 32/124 (25%), Positives = 56/124 (44%), Gaps = 8/124 (6%)
Query: 9 SSPSSLTNFGRSILSFRREQIHSMEGSTLEIELDSFQQHVTDRF---VDLSSVPHDELLS 65
S S N+ + S+ ME + + + + HV V++ S+ D L +
Sbjct: 9 SKEKSGRNYTTELRSYEAACKEDMEIQSFDTRMQARTSHVISTLATGVEVRSLSFDSLKA 68
Query: 66 LKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAIRDGVE 125
+ +G +LD + QE K IL K + V YFE S+K LD NA++ G++
Sbjct: 69 V--IGSLLD---MNQEVAKVILDCKKDIWKNQEMFEFVEAYFETSLKTLDFFNALKRGLQ 123
Query: 126 QIRV 129
+++
Sbjct: 124 GVQI 127
>At4g34320 unknown protein
Length = 374
Score = 38.1 bits (87), Expect = 0.009
Identities = 25/88 (28%), Positives = 41/88 (46%), Gaps = 1/88 (1%)
Query: 64 LSLKWVGKMLDCFL-ICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAIRD 122
LS + ++ C L + QE K IL K + +V +YFE S+K LD C A+
Sbjct: 63 LSFDSLKEVTQCLLEMNQEVVKVILDCKKDIWKNQEMFELVEDYFENSLKTLDFCAALEK 122
Query: 123 GVEQIRVWQKLLEIVLYALDHQRSIGEG 150
G+ + R L+ + L + + + G
Sbjct: 123 GLRRARDSHLLILVALQQFEDESLVQGG 150
>At1g01550 Unknown protein (F22L4.9)
Length = 349
Score = 37.0 bits (84), Expect = 0.019
Identities = 26/113 (23%), Positives = 52/113 (46%), Gaps = 1/113 (0%)
Query: 30 HSMEGSTLEIELDSFQQHVTDRFVDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHT 89
+S+ S L L++F+ ++ L ++L++ W+ + ++ K ++ T
Sbjct: 25 NSVLSSKLLPLLNNFETNLASSISKLVPKEKSDILTVSWMKQAMESLCETHNGIKTLI-T 83
Query: 90 HKGQVVRPPLDRMVSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALD 142
V D+ V Y + SVK LD+CNA + ++ LL+ L+ L+
Sbjct: 84 DLELPVSDWEDKWVDVYLDISVKLLDLCNAFSSELTRLNQGHLLLQFALHNLE 136
>At3g18600 DEAD box helicase protein, putative
Length = 568
Score = 35.8 bits (81), Expect = 0.043
Identities = 22/67 (32%), Positives = 36/67 (52%), Gaps = 1/67 (1%)
Query: 309 MEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFPLTEEKEGEVRQKVHEVSKVCD 368
+EE KKR RK + G E Q+ E+ + + D +E+K +VR K+ E + +
Sbjct: 9 VEELKKRVRKRSRGKKNEQQKAEEKTHTVEENADETQ-KKSEKKVKKVRGKIEEEEEKVE 67
Query: 369 ALKDGLD 375
A++DG D
Sbjct: 68 AMEDGED 74
>At2g46080 unknown protein
Length = 347
Score = 35.4 bits (80), Expect = 0.056
Identities = 28/119 (23%), Positives = 55/119 (45%), Gaps = 4/119 (3%)
Query: 41 LDSFQQHVTDRFVDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLD 100
L+ F+ + +R L D++L+L W+ ++ + ++ T V +
Sbjct: 36 LNGFELRLEERLKKLMPKNKDDILTLSWMKLAMESLCETHKNINTLI-TDLQLPVSDWEE 94
Query: 101 RMVSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKAL 159
+ V Y SV+ LD+CNA + ++ L+ VL+ L Q GE ++ +A+ +L
Sbjct: 95 KWVDVYLNISVRLLDLCNAFSSELTRLNQGDLFLKCVLHNL--QSDSGE-KYLQARSSL 150
>At2g18630 unknown protein
Length = 392
Score = 33.1 bits (74), Expect = 0.28
Identities = 36/155 (23%), Positives = 64/155 (41%), Gaps = 16/155 (10%)
Query: 265 SVLLFVMWALVAAIPCQDRGLNVHFSIPRSYT--WAIPLLLLHERIMEESKKRDRKNACG 322
SVL+F + A A P + ++P W L +E+++ K+ G
Sbjct: 233 SVLIFSVVAAAVAAPPVVAAIAGALAVPVGSVGKWCNTLWTKYEKVVRGQKEIITSIRIG 292
Query: 323 L---LREIQQIEKCVRVMS----DLVDSAHFPLTEEKEGEVRQKVHEVSKVCDALKDGLD 375
++E+ I VR + L+ A F +TEEK EVR + E+ K D + ++
Sbjct: 293 TYISVKEMDNISILVRKVEVEIESLLKKAEFAITEEK--EVRLAIDEIKKKLDVFTETIE 350
Query: 376 PLERQVREVFHRIVRSKTEGLDSL-----GRPNNE 405
L + + +++T L + G P +E
Sbjct: 351 ELGEHAGKYCSDVTKARTVILQRIIRYPAGSPKDE 385
Score = 32.0 bits (71), Expect = 0.62
Identities = 29/123 (23%), Positives = 58/123 (46%), Gaps = 4/123 (3%)
Query: 42 DSFQQHVTDRFVD-LSSVPHDELLSLKWVGKMLDCFL-ICQEEFKAILHTHKGQVVRPPL 99
DS T+R ++ L+S + LS + ++ C L + Q+ K IL + L
Sbjct: 50 DSALHERTNRVINKLASGVEIKSLSFDSLREVTQCLLDMNQDVVKVILQDKEDIWNNQDL 109
Query: 100 DRMVSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKAL 159
+V+ YFE + K +D C+ + + + + R Q +++ + + + E R+ +K L
Sbjct: 110 FSLVNLYFESTAKTMDFCSELENCLNRARRSQVIIQFAVNQFEEENEDKEN--RKYEKTL 167
Query: 160 IDL 162
+L
Sbjct: 168 EEL 170
>At5g66670 At14a protein-like
Length = 408
Score = 32.3 bits (72), Expect = 0.48
Identities = 17/87 (19%), Positives = 42/87 (47%), Gaps = 3/87 (3%)
Query: 80 QEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLY 139
Q+ + I+ + + + L +V YF+ + K LD CN + V++ + Q ++ +
Sbjct: 99 QDTVRVIIESKEDVLKNNDLKALVDVYFKSTSKTLDFCNTVEKCVKKAEISQLIIRFAVK 158
Query: 140 ALDHQ---RSIGEGQFRRAKKALIDLS 163
+ + +GE + ++ K L +++
Sbjct: 159 QFETETVDTDLGESKKKKYVKTLEEMN 185
>At4g28990 unknown protein
Length = 347
Score = 32.3 bits (72), Expect = 0.48
Identities = 20/85 (23%), Positives = 40/85 (46%), Gaps = 6/85 (7%)
Query: 179 HRNRSFGRSNGGRDRDHHQHGNSHSNNNIN------TYQHRSMGQFRSLSWSVSRTWSAA 232
HR R+ S G R + H+HG+ +++ + + + + G+ R S ++R + A
Sbjct: 65 HRTRASSSSPGRRGYEDHKHGSDLNHSGVPPRGRELSSRREAPGRHRDYSPPLARGGAGA 124
Query: 233 RQLQAIGNNINPPRANDLMATNGLA 257
R + + PP D M+ N ++
Sbjct: 125 RPYRRGLDGPEPPHGRDGMSRNNIS 149
>At3g15095 unknown protein, 3' partial
Length = 417
Score = 32.3 bits (72), Expect = 0.48
Identities = 20/56 (35%), Positives = 28/56 (49%), Gaps = 2/56 (3%)
Query: 155 AKKALIDLSIS--MLDDGKESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNIN 208
A+ A + S+S + G NAS N +NGGR R + NS++NNN N
Sbjct: 52 ARSACLTTSLSRRLRTSGSLKNASAGVLNSPMFGANGGRKRSGSGYENSNNNNNNN 107
>At3g20460 sugar transporter, putative
Length = 488
Score = 32.0 bits (71), Expect = 0.62
Identities = 28/111 (25%), Positives = 52/111 (46%), Gaps = 8/111 (7%)
Query: 248 NDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHFSIPRSYTWAIPLLLLHER 307
N L+ +A++ Y + SV+ + AL++ +PC + + F IP S W L + R
Sbjct: 184 NSLVMCASVAVT-YLLGSVISWQKLALISTVPCVFEFVGLFF-IPESPRW----LSRNGR 237
Query: 308 IMEE--SKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFPLTEEKEGEV 356
+ E S +R R N + +E +I+K + + + + F L + V
Sbjct: 238 VKESEVSLQRLRGNNTDITKEAAEIKKYMDNLQEFKEDGFFDLFNPRYSRV 288
>At3g49930 zinc-finger-like protein
Length = 215
Score = 30.8 bits (68), Expect = 1.4
Identities = 13/37 (35%), Positives = 21/37 (56%)
Query: 170 GKESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNN 206
GK N S+ ++ G++ GG R H+ GN +SN +
Sbjct: 146 GKTHNCSICFKSFPSGQALGGHKRCHYDGGNGNSNGD 182
>At3g12080 unknown protein
Length = 663
Score = 30.8 bits (68), Expect = 1.4
Identities = 24/77 (31%), Positives = 37/77 (47%), Gaps = 14/77 (18%)
Query: 9 SSPSSLTNF--GRSILSFRREQIHS----------MEGSTLEIELD--SFQQHVTDRFVD 54
SSPSS ++ S+LS+ + HS ++GS+ E ELD F Q+ D F D
Sbjct: 34 SSPSSSSSIIPSLSVLSYTHQHPHSSRFPFLVAATLDGSSAEEELDFEEFDQYAEDNFAD 93
Query: 55 LSSVPHDELLSLKWVGK 71
S D+ + + + K
Sbjct: 94 DYSDDEDDSIDISVLEK 110
>At2g47500 kinesin heavy chain like protein
Length = 983
Score = 30.8 bits (68), Expect = 1.4
Identities = 34/128 (26%), Positives = 61/128 (47%), Gaps = 19/128 (14%)
Query: 9 SSPSSLTNFGRSILSFRR-EQIHSMEGSTLEIELDSFQQHVTDRFVDLSSVPHDELLS-- 65
SSPSSL+ R++LS ++ E + + S L ++ F+ VT+++ + + P + S
Sbjct: 225 SSPSSLSTLVRAVLSDKKPEDVPKLIESLLSKVVEEFENRVTNQYELVRAAPRESTSSQN 284
Query: 66 ----LKWVGKMLDCFLICQEEFKAI-LHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAI 120
LK +G+ ++ FKAI H Q+ LD + R K L + N
Sbjct: 285 NRSFLKPLGERER----EEKSFKAIKKDDHNSQI----LDEKMK---TRQFKQLTIFNQQ 333
Query: 121 RDGVEQIR 128
++ +E +R
Sbjct: 334 QEDIEGLR 341
>At1g36620 hypothetical protein
Length = 1152
Score = 30.4 bits (67), Expect = 1.8
Identities = 20/55 (36%), Positives = 25/55 (45%), Gaps = 2/55 (3%)
Query: 182 RSFGRSNGGRDRDHHQHGNSHSNNNINT-YQHRSMGQFRSLSWSVSRTWSAARQL 235
R GRS+ R R HGNS+ NN T + S +F + WSA R L
Sbjct: 309 RGRGRSSTPRGRGRG-HGNSYKANNAQTSHPSSSASEFSDIPGVSKEAWSAIRNL 362
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,058,994
Number of Sequences: 26719
Number of extensions: 372873
Number of successful extensions: 1375
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 1303
Number of HSP's gapped (non-prelim): 51
length of query: 405
length of database: 11,318,596
effective HSP length: 102
effective length of query: 303
effective length of database: 8,593,258
effective search space: 2603757174
effective search space used: 2603757174
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126014.1


