

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126010.8 - phase: 0
(727 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
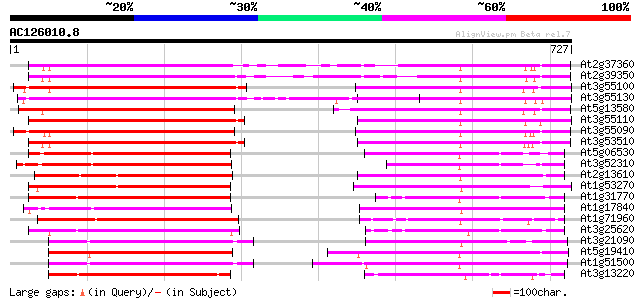
Score E
Sequences producing significant alignments: (bits) Value
At2g37360 ABC transporter like protein 394 e-110
At2g39350 ABC transporter like protein 369 e-102
At3g55100 ABC transporter - like protein 277 1e-74
At3g55130 ABC transporter - like protein 276 2e-74
At5g13580 ABC transporter-like protein 273 2e-73
At3g55110 ABC transporter - like protein 265 5e-71
At3g55090 ABC transporter - like protein 262 4e-70
At3g53510 ABC transporter -like protein 258 1e-68
At5g06530 ABC transporter like protein 200 3e-51
At3g52310 ABC transporter-like protein 199 5e-51
At2g13610 putative ABC transporter 196 3e-50
At1g53270 putative ABC transporter emb|AAD22683.1 195 7e-50
At1g31770 unknown protein 191 2e-48
At1g17840 putative ABC transporter (At1g17840) 189 5e-48
At1g71960 putative ABC transporter 187 2e-47
At3g25620 membrane transporter, putative 184 2e-46
At3g21090 ABC transporter, putative 183 3e-46
At5g19410 membrane transporter - like protein 175 9e-44
At1g51500 ATP-dependent transmembrane transporter, putative 175 9e-44
At3g13220 ABC transporter 174 1e-43
>At2g37360 ABC transporter like protein
Length = 755
Score = 394 bits (1013), Expect = e-110
Identities = 265/750 (35%), Positives = 387/750 (51%), Gaps = 141/750 (18%)
Query: 25 LEFESLTYTVTKKKKVDG----KWSNEDVD-----LLHDITGYAPKGCITAVMGPSGAGK 75
L F LTY+V +KK + + S D LL+ I+G A +G + AV+G SG+GK
Sbjct: 98 LSFTDLTYSVKIQKKFNPLACCRRSGNDSSVNTKILLNGISGEAREGEMMAVLGASGSGK 157
Query: 76 STLLDGLAGRIASGSLKGKVSLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYETLMFAAD 135
STL+D LA RIA SL+G ++L+G + +S+ K SAY+MQ+D LFPMLTV ETLMF+A+
Sbjct: 158 STLIDALANRIAKDSLRGSITLNGEVLESSMQKVISAYVMQDDLLFPMLTVEETLMFSAE 217
Query: 136 FRLG-PLSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVDIIHGPS 194
FRL LS K+ RV+ LI+QLGL S+ T IGDEG RGVSGGERRRVSIG DIIH P
Sbjct: 218 FRLPRSLSKKKKKARVQALIDQLGLRSAAKTVIGDEGHRGVSGGERRRVSIGNDIIHDPI 277
Query: 195 LLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRIQLLLDHLIILARGQLM 254
+LFLDEPTSGLDSTSA VI+ L IA++GS VI++IHQPS RI LLD LI L++G +
Sbjct: 278 ILFLDEPTSGLDSTSAYMVIKVLQRIAQSGSIVIMSIHQPSYRIMGLLDQLIFLSKGNTV 337
Query: 255 FQGSLKDVGHHLNRMGRKIPKGENPIENLIDVIQEYDQCDFVGVEVLAEFARTGMKPPLL 314
+ GS + + IP+ EN E +D+I+E E++ G KP
Sbjct: 338 YSGSPTHLPQFFSEFKHPIPENENKTEFALDLIRE------------LEYSTEGTKP--- 382
Query: 315 SDMEEIISYTNSIAPSPSPLHRGSKYEEKSQDFSYSSQISRRSLNDEFDHSIRSPYNNTP 374
++ + +P SY++ R + +I + +
Sbjct: 383 -----LVEFHKQWRAKQAP--------------SYNNNNKRNTNVSSLKEAITASISRGK 423
Query: 375 MSWSASNSAAFLKFTPSRLKNENKVQKPPSHASPGIYTYSSEILPATPTPHSSDYVVDEN 434
+ A+N+ + S+ +P T+++ P + +V
Sbjct: 424 LVSGATNNNS-------------------SNLTPSFQTFAN--------PFWIEMIVIGK 456
Query: 435 DYLTPTNSSQEHLGPKFANSYIGETWILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATM 494
+ + E LG + + T I++ FTN+ +P
Sbjct: 457 RAILNSRRQPELLGMRLGAVMV--TGIILATMFTNLDNSP-------------------- 494
Query: 495 FHNPKNTLQGITNRLSFFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIA 554
+G RL FF F + F++ +A+P F+QER+IF+RET++NAYR S Y ++
Sbjct: 495 --------KGAQERLGFFAFAMSTTFYTCAEAIPVFLQERYIFMRETAYNAYRRSSYVLS 546
Query: 555 SLITHMPFLALQALAYAAIVWFALELRG---PFIYFFLVLFISLLSTNSFVVFVSSIVPN 611
I +P L + + ++AA ++A+ L G F +F+ + S + +SFV F+S ++PN
Sbjct: 547 QSIISIPALIVLSASFAATTFWAVGLDGGANGFFFFYFTILASFWAGSSFVTFLSGVIPN 606
Query: 612 YILGYAAVIAFTALFFLFCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEYQT----- 666
+LG+ V+A A F LF G+F+S + IP+YW W + +S + YPYEG+L NE+Q
Sbjct: 607 VMLGFTVVVAILAYFLLFSGFFISRDRIPVYWLWFHYISLVKYPYEGVLQNEFQNPTRCF 666
Query: 667 --------NETFGS--ND-------------GVSI-------TGFDILKSLHIGTEEIKK 696
N G ND G ++ TG DILK G +I K
Sbjct: 667 ARGVQLFDNSPLGEFPNDVKVNLLKSMSGVLGTNVTAETCVTTGIDILKQQ--GITDISK 724
Query: 697 RNNVLIMLGWAVLYRILFYIILRFASKNQR 726
N + I + W +R+LFY L SKN+R
Sbjct: 725 WNCLWITVAWGFFFRVLFYFTLLIGSKNKR 754
>At2g39350 ABC transporter like protein
Length = 740
Score = 369 bits (948), Expect = e-102
Identities = 264/758 (34%), Positives = 377/758 (48%), Gaps = 141/758 (18%)
Query: 25 LEFESLTYTVTKKKKVD-----GKWSNEDVD-----------LLHDITGYAPKGCITAVM 68
L F++LTY V+ + K+D + ED + LL++I+G G I AV+
Sbjct: 67 LSFDNLTYNVSVRPKLDFRNLFPRRRTEDPEIAQTARPKTKTLLNNISGETRDGEIMAVL 126
Query: 69 GPSGAGKSTLLDGLAGRIASGSLKGKVSLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYE 128
G SG+GKSTL+D LA RIA GSLKG V L+G ++ + ++K SAY+MQ+D LFPMLTV E
Sbjct: 127 GASGSGKSTLIDALANRIAKGSLKGTVKLNGETLQSRMLKVISAYVMQDDLLFPMLTVEE 186
Query: 129 TLMFAADFRLG-PLSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGV 187
TLMFAA+FRL L K+ RV+ LI+QLG+ ++ T IGDEG RG+SGGERRRVSIG+
Sbjct: 187 TLMFAAEFRLPRSLPKSKKKLRVQALIDQLGIRNAAKTIIGDEGHRGISGGERRRVSIGI 246
Query: 188 DIIHGPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRIQLLLDHLII 247
DIIH P LLFLDEPTSGLDSTSA V++ L IA++GS VI++IHQPS R+ LLD LI
Sbjct: 247 DIIHDPILLFLDEPTSGLDSTSAFMVVKVLKRIAQSGSIVIMSIHQPSHRVLGLLDRLIF 306
Query: 248 LARGQLMFQGSLKDVGHHLNRMGRKIPKGENPIENLIDVIQEYDQCDFVGVEVLAEFART 307
L+RG ++ GS + G IP+ EN E +D+I+E + G L EF +
Sbjct: 307 LSRGHTVYSGSPASLPRFFTEFGSPIPENENRTEFALDLIRELEG-SAGGTRGLIEFNK- 364
Query: 308 GMKPPLLSDMEEIISYTNSIAPSPSPLHRGSKYEEKSQDFSYSSQISRRSLNDEFDHSIR 367
+E+ +N P P
Sbjct: 365 --------KWQEMKKQSNRQPPLTPP---------------------------------S 383
Query: 368 SPYNNTPMSWSASNSAAFLKFTPSRLKNENKVQKPPSHASPGIYTYSSEI-LPATPTPHS 426
SPY N + + + S + V S A G T ++ + +PA P
Sbjct: 384 SPYPNLTLKEAIAAS----------ISRGKLVSGGESVAHGGATTNTTTLAVPAFANPMW 433
Query: 427 SDYVVDENDYLTPTNSSQEHLGPKFANSYIGETWILMRRNFTNIRRTPELFLSRLMVLTF 486
+ + + E G + A+ I T ++ F + +P+ R L F
Sbjct: 434 IEIKTLSKRSMLNSRRQPELFGIRIASVVI--TGFILATVFWRLDNSPKGVQER---LGF 488
Query: 487 MGVMMATMFHNPKNTLQGITNRLSFFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAY 546
M+TMF+ + L P F+QER+IF+RET++NAY
Sbjct: 489 FAFAMSTMFYTCADAL-------------------------PVFLQERYIFMRETAYNAY 523
Query: 547 RASCYTIASLITHMPFLALQALAYAAIVWFALELRG---PFIYFFLVLFISLLSTNSFVV 603
R S Y ++ I P L ++A+AA ++A+ L G +++ L++ S S +SFV
Sbjct: 524 RRSSYVLSHAIVSFPSLIFLSVAFAATTYWAVGLDGGLTGLLFYCLIILASFWSGSSFVT 583
Query: 604 FVSSIVPNYILGYAAVIAFTALFFLFCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNE 663
F+S +VP+ +LGY V+A A F LF G+F++ IP YW W + +S + YPYE +L NE
Sbjct: 584 FLSGVVPSVMLGYTIVVAILAYFLLFSGFFINRNRIPDYWIWFHYMSLVKYPYEAVLQNE 643
Query: 664 YQTNE----------------------------TFGSNDGVSI-------TGFDILKSLH 688
+ T + GV+I TG DIL+
Sbjct: 644 FSDATKCFVRGVQIFDNTPLGELPEVMKLKLLGTVSKSLGVTISSTTCLTTGSDILRQQ- 702
Query: 689 IGTEEIKKRNNVLIMLGWAVLYRILFYIILRFASKNQR 726
G ++ K N + I + + +RILFY L SKN+R
Sbjct: 703 -GVVQLSKWNCLFITVAFGFFFRILFYFTLLLGSKNKR 739
>At3g55100 ABC transporter - like protein
Length = 662
Score = 277 bits (709), Expect = 1e-74
Identities = 161/308 (52%), Positives = 209/308 (67%), Gaps = 10/308 (3%)
Query: 6 LELETVIDIKHK---PVSFTGGLEFESLTYTVTKKKKVDGKWSNEDVD---LLHDITGYA 59
L+ + VID+ P+ F L F LTY VT +++ ++ + LL+ ITG A
Sbjct: 2 LQRDAVIDVDESEIPPIPFV--LAFNDLTYNVTLQQRFGLRFGHSPAKIKTLLNGITGEA 59
Query: 60 PKGCITAVMGPSGAGKSTLLDGLAGRIASGSLKGKVSLDGNSVNASLIKRTSAYIMQEDR 119
+G I A++G SGAGKSTL+D LAG+IA GSLKG V+L+G ++ + L++ SAY+MQED
Sbjct: 60 KEGEILAILGASGAGKSTLIDALAGQIAEGSLKGTVTLNGEALQSRLLRVISAYVMQEDL 119
Query: 120 LFPMLTVYETLMFAADFRLG-PLSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGG 178
LFPMLTV ETLMFAA+FRL LS KR RVE LI+QLGL++ +NT IGDEG RGVSGG
Sbjct: 120 LFPMLTVEETLMFAAEFRLPRSLSKSKKRNRVETLIDQLGLTTVKNTVIGDEGHRGVSGG 179
Query: 179 ERRRVSIGVDIIHGPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRI 238
ERRRVSIG DIIH P +LFLDEPTSGLDSTSA V++ L IAR+GS VI++IHQPS RI
Sbjct: 180 ERRRVSIGTDIIHDPIVLFLDEPTSGLDSTSAFMVVQVLKKIARSGSIVIMSIHQPSGRI 239
Query: 239 QLLLDHLIILARGQLMFQGSLKDVGHHLNRMGRKIPKGENPIENLIDVIQEYDQCDFVGV 298
LD +I+L+ GQ++F S + + G IP+ EN E +D+I++ + G
Sbjct: 240 MEFLDRVIVLSSGQIVFSDSPATLPLFFSEFGSPIPEKENIAEFTLDLIKDLEGSP-EGT 298
Query: 299 EVLAEFAR 306
L EF R
Sbjct: 299 RGLVEFNR 306
Score = 194 bits (494), Expect = 1e-49
Identities = 109/317 (34%), Positives = 172/317 (53%), Gaps = 40/317 (12%)
Query: 449 PKFANSYIGETWILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNR 508
P + N + ET IL +R N RTPEL +R+ ++ G ++AT++ ++ +G+ R
Sbjct: 348 PSYVNPWWVETVILAKRYMINWTRTPELIGTRVFIVMMTGFLLATVYWKVDDSPRGVQER 407
Query: 509 LSFFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQAL 568
LSFF F + F+S D +PAFIQER+IF+RET+HNAYR S Y I+ + +P L ++
Sbjct: 408 LSFFSFAMATMFYSCADGLPAFIQERYIFLRETAHNAYRRSSYVISHSLVTLPHLFALSI 467
Query: 569 AYAAIVWFALELRG---PFIYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTAL 625
+AA ++ + L G FIY+ +++F S S SFV FVS ++PN ++ Y + +
Sbjct: 468 GFAATTFWFVGLNGGLAGFIYYLMIIFASFWSGCSFVTFVSGVIPNVMMSYMVTFGYLSY 527
Query: 626 FFLFCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEY--------QTNETFGSN--DG 675
LF G++++ + I LYW W++ +S + YPYE +L NE+ + N+ F + +G
Sbjct: 528 CLLFSGFYVNRDRIHLYWIWIHYISLLKYPYEAVLHNEFDDPSRCFVRGNQVFDNTIMEG 587
Query: 676 VS-------------------------ITGFDILKSLHIGTEEIKKRNNVLIMLGWAVLY 710
VS TG D+LK G E++ K + + L W +
Sbjct: 588 VSETTKAKLLETMSGYLGMELTESTCLTTGSDLLK--QHGIEQLDKWGCLWVTLAWGFFF 645
Query: 711 RILFYIILRFASKNQRS 727
RILFY L SKN+R+
Sbjct: 646 RILFYFSLLLGSKNKRA 662
>At3g55130 ABC transporter - like protein
Length = 725
Score = 276 bits (707), Expect = 2e-74
Identities = 206/534 (38%), Positives = 290/534 (53%), Gaps = 34/534 (6%)
Query: 11 VIDIKH---KPVSFTGGLEFESLTYTVTKKKKVDGKWSNEDVDLLHDITGYAPKGCITAV 67
+ID++ KPV + L F +L Y VT +++ N LL D++G A G I AV
Sbjct: 58 IIDVEALYVKPVPYV--LNFNNLQYDVTLRRRFGFSRQNGVKTLLDDVSGEASDGDILAV 115
Query: 68 MGPSGAGKSTLLDGLAGRIASGSLKGKVSLDGNSVNAS-LIKRTSAYIMQEDRLFPMLTV 126
+G SGAGKSTL+D LAGR+A GSL+G V+L+G V S L+K SAY+MQ+D LFPMLTV
Sbjct: 116 LGASGAGKSTLIDALAGRVAEGSLRGSVTLNGEKVLQSRLLKVISAYVMQDDLLFPMLTV 175
Query: 127 YETLMFAADFRLG-PLSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSI 185
ETLMFA++FRL LS K +RVE LI+QLGL ++ NT IGDEG RGVSGGERRRVSI
Sbjct: 176 KETLMFASEFRLPRSLSKSKKMERVEALIDQLGLRNAANTVIGDEGHRGVSGGERRRVSI 235
Query: 186 GVDIIHGPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRIQLLLDHL 245
G+DIIH P +LFLDEPTSGLDST+A V++ L IA++GS VI++IHQPS+RI LLD L
Sbjct: 236 GIDIIHDPIVLFLDEPTSGLDSTNAFMVVQVLKRIAQSGSIVIMSIHQPSARIVELLDRL 295
Query: 246 IILARGQLMFQGSLKDVGHHLNRMGRKIPKGENPIENLIDVIQEYDQCDFVGVEVLAEFA 305
IIL+RG+ +F GS + + GR IP+ EN E +D+++E + + G + L +F
Sbjct: 296 IILSRGKSVFNGSPASLPGFFSDFGRPIPEKENISEFALDLVRELEGSN-EGTKALVDFN 354
Query: 306 RTGMKPPLLSDMEEIISYTNSIAPSPSPLHRGSKYEEKSQDFSYSSQISRRSLNDEFDHS 365
+ IS S AP + L + K + ++ +SR L S
Sbjct: 355 EKW--------QQNKISLIQS-APQTNKLDQDRSLSLKE---AINASVSRGKL---VSGS 399
Query: 366 IRSPYNNTPMSWSASNSAAFLKFTPSRLKNENKVQKPPSHASPGIYTYSSEILPAT---- 421
RS + S +N + F F ++ +N ++ P + + L AT
Sbjct: 400 SRSNPTSMETVSSYANPSLFETFILAKRYMKNWIRMPELVGTRIATVMVTGCLLATVYWK 459
Query: 422 --PTPHSSDYVVDENDYLTPTNSSQEHLGPKFANSYIGETWILMRRNFTNIRRTPELFLS 479
TP + + ++ PT + +I E +I +R N RT +S
Sbjct: 460 LDHTPRGAQERLTLFAFVVPT---MFYCCLDNVPVFIQERYIFLRETTHNAYRTSSYVIS 516
Query: 480 RLMVLTFMGVMMATMFHNPKNTLQGITNRLSFFIFTVCLFF--FSSNDAVPAFI 531
+V + + +F G++ L F+F L + F S +V FI
Sbjct: 517 HSLVSLPQLLAPSLVFSAITFWTVGLSGGLEGFVFYCLLIYASFWSGSSVVTFI 570
Score = 172 bits (437), Expect = 5e-43
Identities = 104/313 (33%), Positives = 165/313 (52%), Gaps = 36/313 (11%)
Query: 451 FANSYIGETWILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLS 510
+AN + ET+IL +R N R PEL +R+ + G ++AT++ +T +G RL+
Sbjct: 413 YANPSLFETFILAKRYMKNWIRMPELVGTRIATVMVTGCLLATVYWKLDHTPRGAQERLT 472
Query: 511 FFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAY 570
F F V F+ D VP FIQER+IF+RET+HNAYR S Y I+ + +P L +L +
Sbjct: 473 LFAFVVPTMFYCCLDNVPVFIQERYIFLRETTHNAYRTSSYVISHSLVSLPQLLAPSLVF 532
Query: 571 AAIVWFALELRG---PFIYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFF 627
+AI ++ + L G F+++ L+++ S S +S V F+S +VPN +L Y I + A
Sbjct: 533 SAITFWTVGLSGGLEGFVFYCLLIYASFWSGSSVVTFISGVVPNIMLCYMVSITYLAYCL 592
Query: 628 LFCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEYQTN--------ETFGSN--DGVS 677
L G++++ + IP YW W + +S + YPYE +L+NE+ + F S GVS
Sbjct: 593 LLSGFYVNRDRIPFYWTWFHYISILKYPYEAVLINEFDDPSRCFVRGVQVFDSTLLGGVS 652
Query: 678 ITG-----FDILKSLHI------------------GTEEIKKRNNVLIMLGWAVLYRILF 714
+G + KSL G ++ K + + I + +RILF
Sbjct: 653 DSGKVKLLETLSKSLRTKITESTCLRTGSDLLAQQGITQLSKWDCLWITFASGLFFRILF 712
Query: 715 YIILRFASKNQRS 727
Y L F S+N+R+
Sbjct: 713 YFALLFGSRNKRT 725
>At5g13580 ABC transporter-like protein
Length = 727
Score = 273 bits (698), Expect = 2e-73
Identities = 157/295 (53%), Positives = 199/295 (67%), Gaps = 15/295 (5%)
Query: 12 IDIKHKPVSFTGGLEFESLTYTVTKKKKV--------------DGKWSNEDVDLLHDITG 57
+D+ S L F LTY+V ++K +G +S++ LL+ ITG
Sbjct: 55 VDLASPDQSVPFVLSFTDLTYSVKVRRKFTWRRSVSSDPGAPSEGIFSSKTKTLLNGITG 114
Query: 58 YAPKGCITAVMGPSGAGKSTLLDGLAGRIASGSLKGKVSLDGNSVNASLIKRTSAYIMQE 117
A G I AV+G SG+GKSTL+D LA RIA GSLKG V+L+G +N+ + K SAY+MQ+
Sbjct: 115 EARDGEILAVLGASGSGKSTLIDALANRIAKGSLKGNVTLNGEVLNSKMQKAISAYVMQD 174
Query: 118 DRLFPMLTVYETLMFAADFRLG-PLSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVS 176
D LFPMLTV ETLMFAA+FRL LS K RV+ LI+QLGL ++ NT IGDEG RG+S
Sbjct: 175 DLLFPMLTVEETLMFAAEFRLPRSLSKSKKSLRVQALIDQLGLRNAANTVIGDEGHRGIS 234
Query: 177 GGERRRVSIGVDIIHGPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSS 236
GGERRRVSIG+DIIH P LLFLDEPTSGLDSTSALSVI+ L IA++GS VI+T+HQPS
Sbjct: 235 GGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSALSVIKVLKRIAQSGSMVIMTLHQPSY 294
Query: 237 RIQLLLDHLIILARGQLMFQGSLKDVGHHLNRMGRKIPKGENPIENLIDVIQEYD 291
R+ LLD L+ L+RGQ +F GS + G IP+ EN E +D+I+E +
Sbjct: 295 RLLRLLDRLLFLSRGQTVFSGSPAMLPRFFAEFGHPIPEHENRTEFALDLIRELE 349
Score = 205 bits (522), Expect = 6e-53
Identities = 123/345 (35%), Positives = 184/345 (52%), Gaps = 53/345 (15%)
Query: 420 ATPTPHSSDYVVDENDYLTPTNSSQEHLGPKFANSYIGETWILMRRNFTNIRRTPELFLS 479
AT T HSS SS P FAN + E +L +R+ TN RR PELF
Sbjct: 397 ATTTTHSS-------------GSSPVSTIPTFANPFWVELAVLAKRSMTNSRRQPELFGI 443
Query: 480 RLMVLTFMGVMMATMFHNPKNTLQGITNRLSFFIFTVCLFFFSSNDAVPAFIQERFIFIR 539
RL + G ++ATMF N+ +G+ RL F F + F++ DA+P F+QERFIF+R
Sbjct: 444 RLGAVLVTGFILATMFWQLDNSPKGVQERLGCFAFAMSTTFYTCADALPVFLQERFIFMR 503
Query: 540 ETSHNAYRASCYTIASLITHMPFLALQALAYAAIVWFALELRG---PFIYFFLVLFISLL 596
ET++NAYR S Y ++ + +P L + +LA+AAI ++ + L G F+++FLV+ S
Sbjct: 504 ETAYNAYRRSSYVLSHSLVALPSLIILSLAFAAITFWGVGLDGGLMGFLFYFLVILASFW 563
Query: 597 STNSFVVFVSSIVPNYILGYAAVIAFTALFFLFCGYFLSSEDIPLYWRWMNKVSTMTYPY 656
+ +SFV F+S +VP+ +LGY V+A A F LF G+F++ + IP YW W + +S + YPY
Sbjct: 564 AGSSFVTFLSGVVPHVMLGYTIVVAILAYFLLFSGFFINRDRIPGYWIWFHYISLVKYPY 623
Query: 657 EGLLMNEY----------------------------QTNETFGSNDGVSI-------TGF 681
E +L+NE+ + T + G+ I TG+
Sbjct: 624 EAVLLNEFGDPTKCFVRGVQIFDNTPLVAVPQGMKVRLLATMSKSLGMRITSSTCLTTGY 683
Query: 682 DILKSLHIGTEEIKKRNNVLIMLGWAVLYRILFYIILRFASKNQR 726
DIL+ G ++ K N + + + W +RILFY L SKN+R
Sbjct: 684 DILQQQ--GVTDLTKWNCLWVTVAWGFFFRILFYFSLLLGSKNKR 726
>At3g55110 ABC transporter - like protein
Length = 708
Score = 265 bits (678), Expect = 5e-71
Identities = 157/284 (55%), Positives = 197/284 (69%), Gaps = 5/284 (1%)
Query: 25 LEFESLTYTVTKKKKVD--GKWSNEDVDLLHDITGYAPKGCITAVMGPSGAGKSTLLDGL 82
L F +L+Y V +++ D + + LL DITG A G I AV+G SGAGKSTL+D L
Sbjct: 63 LSFNNLSYNVVLRRRFDFSRRKTASVKTLLDDITGEARDGEILAVLGGSGAGKSTLIDAL 122
Query: 83 AGRIASGSLKGKVSLDGNSVNAS-LIKRTSAYIMQEDRLFPMLTVYETLMFAADFRLG-P 140
AGR+A SLKG V+L+G V S L+K SAY+MQ+D LFPMLTV ETLMFA++FRL
Sbjct: 123 AGRVAEDSLKGTVTLNGEKVLQSRLLKVISAYVMQDDLLFPMLTVKETLMFASEFRLPRS 182
Query: 141 LSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVDIIHGPSLLFLDE 200
L K +RVE LI+QLGL ++ +T IGDEG RGVSGGERRRVSIG+DIIH P LLFLDE
Sbjct: 183 LPKSKKMERVETLIDQLGLRNAADTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDE 242
Query: 201 PTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRIQLLLDHLIILARGQLMFQGSLK 260
PTSGLDST+A V++ L IA++GS VI++IHQPS+RI LLD LIIL+ G+ +F GS
Sbjct: 243 PTSGLDSTNAFMVVQVLKRIAQSGSVVIMSIHQPSARIIGLLDRLIILSHGKSVFNGSPV 302
Query: 261 DVGHHLNRMGRKIPKGENPIENLIDVIQEYDQCDFVGVEVLAEF 304
+ + GR IP+ EN E +DVI+E + G L EF
Sbjct: 303 SLPSFFSSFGRPIPEKENITEFALDVIRELEGSS-EGTRDLVEF 345
Score = 189 bits (481), Expect = 4e-48
Identities = 102/313 (32%), Positives = 168/313 (53%), Gaps = 36/313 (11%)
Query: 451 FANSYIGETWILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLS 510
+AN + ET+IL +R N RTPEL R+ + G+++AT++ NT +G R+
Sbjct: 396 YANPPLAETFILAKRYIKNWIRTPELIGMRIGTVMVTGLLLATVYWRLDNTPRGAQERMG 455
Query: 511 FFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAY 570
FF F + F+ D +P FIQER+IF+RET+HNAYR S Y I+ + +P L ++A+
Sbjct: 456 FFAFGMSTMFYCCADNIPVFIQERYIFLRETTHNAYRTSSYVISHALVSLPQLLALSIAF 515
Query: 571 AAIVWFALELRG---PFIYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFF 627
AA ++ + L G F Y+ L+++ + S +S V F+S ++PN ++ Y IA+ +
Sbjct: 516 AATTFWTVGLSGGLESFFYYCLIIYAAFWSGSSIVTFISGLIPNVMMSYMVTIAYLSYCL 575
Query: 628 LFCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEYQTN-------------------- 667
L G++++ + IPLYW W + +S + YPYE +L+NE+
Sbjct: 576 LLGGFYINRDRIPLYWIWFHYISLLKYPYEAVLINEFDDPSRCFVKGVQVFDGTLLAEVS 635
Query: 668 --------ETFGSNDGVSITGFDILKS-----LHIGTEEIKKRNNVLIMLGWAVLYRILF 714
+T + G IT L++ + G ++ K + + I L W + +RILF
Sbjct: 636 HVMKVKLLDTLSGSLGTKITESTCLRTGPDLLMQQGITQLSKWDCLWITLAWGLFFRILF 695
Query: 715 YIILRFASKNQRS 727
Y+ L F SKN+R+
Sbjct: 696 YLSLLFGSKNKRT 708
>At3g55090 ABC transporter - like protein
Length = 720
Score = 262 bits (670), Expect = 4e-70
Identities = 150/296 (50%), Positives = 196/296 (65%), Gaps = 11/296 (3%)
Query: 5 GLELETVIDIKHKPVSFTGGLEFESLTYTVTKKKKVDGK----WSNEDVD----LLHDIT 56
G L+ D +PV F L F +LTY V+ ++K+D W LL +I+
Sbjct: 39 GSSLDGDNDHLMRPVPFV--LSFNNLTYNVSVRRKLDFHDLVPWRRTSFSKTKTLLDNIS 96
Query: 57 GYAPKGCITAVMGPSGAGKSTLLDGLAGRIASGSLKGKVSLDGNSVNASLIKRTSAYIMQ 116
G G I AV+G SG+GKSTL+D LA RIA GSLKG V+L+G ++ + ++K SAY+MQ
Sbjct: 97 GETRDGEILAVLGASGSGKSTLIDALANRIAKGSLKGTVTLNGEALQSRMLKVISAYVMQ 156
Query: 117 EDRLFPMLTVYETLMFAADFRLG-PLSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGV 175
+D LFPMLTV ETLMFAA+FRL L K+ RV+ LI+QLG+ ++ T IGDEG RG+
Sbjct: 157 DDLLFPMLTVEETLMFAAEFRLPRSLPKSKKKLRVQALIDQLGIRNAAKTIIGDEGHRGI 216
Query: 176 SGGERRRVSIGVDIIHGPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPS 235
SGGERRRVSIG+DIIH P +LFLDEPTSGLDSTSA V++ L IA +GS +I++IHQPS
Sbjct: 217 SGGERRRVSIGIDIIHDPIVLFLDEPTSGLDSTSAFMVVKVLKRIAESGSIIIMSIHQPS 276
Query: 236 SRIQLLLDHLIILARGQLMFQGSLKDVGHHLNRMGRKIPKGENPIENLIDVIQEYD 291
R+ LLD LI L+RG +F GS + G IP+ EN E +D+I+E +
Sbjct: 277 HRVLSLLDRLIFLSRGHTVFSGSPASLPSFFAGFGNPIPENENQTEFALDLIRELE 332
Score = 201 bits (511), Expect = 1e-51
Identities = 117/316 (37%), Positives = 172/316 (54%), Gaps = 40/316 (12%)
Query: 449 PKFANSYIGETWILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNR 508
P FAN + E L RR+ N RR PEL RL + G ++AT+F N+ +G+ R
Sbjct: 406 PAFANPFWIEIKTLTRRSILNSRRQPELLGMRLATVIVTGFILATVFWRLDNSPKGVQER 465
Query: 509 LSFFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQAL 568
L FF F + F++ DA+P F+QER+IF+RET++NAYR S Y ++ I P L +L
Sbjct: 466 LGFFAFAMSTMFYTCADALPVFLQERYIFMRETAYNAYRRSSYVLSHAIVTFPSLIFLSL 525
Query: 569 AYAAIVWFALELRG---PFIYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTAL 625
A+A ++A+ L G F+++ L++ S S +SFV F+S +VP+ +LGY V+A A
Sbjct: 526 AFAVTTFWAVGLEGGLMGFLFYCLIILASFWSGSSFVTFLSGVVPHVMLGYTIVVAILAY 585
Query: 626 FFLFCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEYQT------------------N 667
F LF G+F++ + IP YW W + +S + YPYE +L NE+
Sbjct: 586 FLLFSGFFINRDRIPQYWIWFHYLSLVKYPYEAVLQNEFSDPTECFVRGVQLFDNSPLGE 645
Query: 668 ETFGSN----DGVS-------------ITGFDILKSLHIGTEEIKKRNNVLIMLGWAVLY 710
T+G D VS TG D+LK G ++ K N +LI +G+ L+
Sbjct: 646 LTYGMKLRLLDSVSRSIGMRISSSTCLTTGADVLKQQ--GVTQLSKWNCLLITVGFGFLF 703
Query: 711 RILFYIILRFASKNQR 726
RILFY+ L SKN+R
Sbjct: 704 RILFYLCLLLGSKNKR 719
>At3g53510 ABC transporter -like protein
Length = 739
Score = 258 bits (658), Expect = 1e-68
Identities = 150/293 (51%), Positives = 197/293 (67%), Gaps = 14/293 (4%)
Query: 25 LEFESLTYTVTKKKKV-------DGKWSNEDVD-----LLHDITGYAPKGCITAVMGPSG 72
L F+ LTY+V KKK + + D++ LL+ I+G A +G + AV+G SG
Sbjct: 88 LSFKDLTYSVKIKKKFKPFPCCGNSPFDGNDMEMNTKVLLNGISGEAREGEMMAVLGASG 147
Query: 73 AGKSTLLDGLAGRIASGSLKGKVSLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYETLMF 132
+GKSTL+D LA RI+ SL+G ++L+G + +SL K SAY+MQ+D LFPMLTV ETLMF
Sbjct: 148 SGKSTLIDALANRISKESLRGDITLNGEVLESSLHKVISAYVMQDDLLFPMLTVEETLMF 207
Query: 133 AADFRL-GPLSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVDIIH 191
+A+FRL LS K+ RV+ LI+QLGL ++ T IGDEG RGVSGGERRRVSIG DIIH
Sbjct: 208 SAEFRLPSSLSKKKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGTDIIH 267
Query: 192 GPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRIQLLLDHLIILARG 251
P +LFLDEPTSGLDSTSA V++ L IA++GS VI++IHQPS RI LLD LI L+RG
Sbjct: 268 DPIILFLDEPTSGLDSTSAYMVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDKLIFLSRG 327
Query: 252 QLMFQGSLKDVGHHLNRMGRKIPKGENPIENLIDVIQEYDQCDFVGVEVLAEF 304
++ GS + + G IP+ EN E +D+I+E + G + L EF
Sbjct: 328 NTVYSGSPTHLPQFFSEFGHPIPENENKPEFALDLIRELEDSP-EGTKSLVEF 379
Score = 198 bits (504), Expect = 8e-51
Identities = 115/314 (36%), Positives = 171/314 (53%), Gaps = 40/314 (12%)
Query: 451 FANSYIGETWILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLS 510
FAN + E ++ +R+ N RR PELF RL + G+++AT+F N+ +GI RL
Sbjct: 427 FANPFWTEMLVIGKRSILNSRRQPELFGIRLGAVLVTGMILATIFWKLDNSPRGIQERLG 486
Query: 511 FFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAY 570
FF F + F++ +A+P F+QER+IF+RET++NAYR S Y +A I +P L + + A+
Sbjct: 487 FFAFAMSTTFYTCAEAIPVFLQERYIFMRETAYNAYRRSSYVLAHTIISIPALIILSAAF 546
Query: 571 AAIVWFALELRG---PFIYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFF 627
AA + A+ L G F++FF + + + +SFV F+S +V + ++G+ V+A A F
Sbjct: 547 AASTFSAVGLAGGSEGFLFFFFTILTAFWAGSSFVTFLSGVVSHVMIGFTVVVAILAYFL 606
Query: 628 LFCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEYQ-------------TNETFG--- 671
LF G+F+S + IPLYW W + +S + YPYEG+L NE++ N G
Sbjct: 607 LFSGFFISRDRIPLYWIWFHYLSLVKYPYEGVLQNEFEDPTKCFVRGIQMFDNSPLGQVP 666
Query: 672 ---------SNDGV----------SITGFDILKSLHIGTEEIKKRNNVLIMLGWAVLYRI 712
S GV TG DILK G EI K N + I + W +R+
Sbjct: 667 TAVKISLLKSMSGVLGINVTAETCVTTGIDILKQQ--GITEISKWNCLWITVAWGFFFRV 724
Query: 713 LFYIILRFASKNQR 726
LFY L SKN+R
Sbjct: 725 LFYFTLLIGSKNKR 738
>At5g06530 ABC transporter like protein
Length = 627
Score = 200 bits (508), Expect = 3e-51
Identities = 123/263 (46%), Positives = 162/263 (60%), Gaps = 6/263 (2%)
Query: 25 LEFESLTYTVTKKKKVDGKWSNEDVDLLHDITGYAPKGCITAVMGPSGAGKSTLLDGLAG 84
L+F +TY V KK S+ + ++L I+G G + A+MGPSG+GK+TLL LAG
Sbjct: 33 LKFRDVTYKVVIKKLT----SSVEKEILTGISGSVNPGEVLALMGPSGSGKTTLLSLLAG 88
Query: 85 RIASGSLKGKVSLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYETLMFAADFRLGP-LSA 143
RI+ S G V+ + + L K ++ Q+D LFP LTV ETL +AA RL L+
Sbjct: 89 RISQSSTGGSVTYNDKPYSKYL-KSKIGFVTQDDVLFPHLTVKETLTYAARLRLPKTLTR 147
Query: 144 VDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVDIIHGPSLLFLDEPTS 203
K+QR +I++LGL ++T IG RGVSGGER+RVSIG +II PSLL LDEPTS
Sbjct: 148 EQKKQRALDVIQELGLERCQDTMIGGAFVRGVSGGERKRVSIGNEIIINPSLLLLDEPTS 207
Query: 204 GLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRIQLLLDHLIILARGQLMFQGSLKDVG 263
GLDST+AL I LHDIA G TVI TIHQPSSR+ D LI+L RG L++ G +
Sbjct: 208 GLDSTTALRTILMLHDIAEAGKTVITTIHQPSSRLFHRFDKLILLGRGSLLYFGKSSEAL 267
Query: 264 HHLNRMGRKIPKGENPIENLIDV 286
+ + +G NP E L+D+
Sbjct: 268 DYFSSIGCSPLIAMNPAEFLLDL 290
Score = 56.2 bits (134), Expect = 6e-08
Identities = 59/263 (22%), Positives = 107/263 (40%), Gaps = 23/263 (8%)
Query: 461 ILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLSFFIFTVCLF- 519
IL R R +L VL+ ++ + + T G+ ++ F +
Sbjct: 378 ILFCRGLKERRHEYFSWLRVTQVLSTAVILGLLWWQSDIRTPMGLQDQAGLLFFIAVFWG 437
Query: 520 FFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAYAAIVWFALE 579
FF A+ AF QER + +E + + YR S Y +A + +P + + +V+F
Sbjct: 438 FFPVFTAIFAFPQERAMLNKERAADMYRLSAYFLARTTSDLPLDFILPSLFLLVVYFMTG 497
Query: 580 LR---GPFIYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFFLFCGYFLSS 636
LR PF L +F+ +++ + + +I+ + F L G+F+
Sbjct: 498 LRISPYPFFLSMLTVFLCIIAAQGLGLAIGAILMDLKKATTLASVTVMTFMLAGGFFV-- 555
Query: 637 EDIPLYWRWMNKVSTMTYPYEGLLMNEYQTNETFGSNDGVSITGFDILKSLHIGTEEIKK 696
+ +P++ W+ +S + Y+ LL +YQ + VSI G I L
Sbjct: 556 KKVPVFISWIRYLSFNYHTYKLLLKVQYQ-------DFAVSINGMRIDNGL--------- 599
Query: 697 RNNVLIMLGWAVLYRILFYIILR 719
V ++ YR+L Y+ LR
Sbjct: 600 -TEVAALVVMIFGYRLLAYLSLR 621
>At3g52310 ABC transporter-like protein
Length = 737
Score = 199 bits (506), Expect = 5e-51
Identities = 117/279 (41%), Positives = 172/279 (60%), Gaps = 7/279 (2%)
Query: 10 TVIDIKHKPVSFTGGLEFESLTYTVTKKKKVDGKWSNEDVDLLHDITGYAPKGCITAVMG 69
+V+ + +P +F L+F +TY VT K G S+ + +L+ I+G A G + A+MG
Sbjct: 131 SVVKFQAEP-TFPIYLKFIDITYKVTTK----GMTSSSEKSILNGISGSAYPGELLALMG 185
Query: 70 PSGAGKSTLLDGLAGRIASGSLKGKVSLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYET 129
PSG+GK+TLL+ L GR ++ G VS + + L K ++ Q+D LFP LTV ET
Sbjct: 186 PSGSGKTTLLNALGGRFNQQNIGGSVSYNDKPYSKHL-KTRIGFVTQDDVLFPHLTVKET 244
Query: 130 LMFAADFRLGP-LSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVD 188
L + A RL L+ +K QR +I++LGL ++T IG RGVSGGER+RV IG +
Sbjct: 245 LTYTALLRLPKTLTEQEKEQRAASVIQELGLERCQDTMIGGSFVRGVSGGERKRVCIGNE 304
Query: 189 IIHGPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRIQLLLDHLIIL 248
I+ PSLL LDEPTS LDST+AL +++ LH IA+ G T++ TIHQPSSR+ D L++L
Sbjct: 305 IMTNPSLLLLDEPTSSLDSTTALKIVQMLHCIAKAGKTIVTTIHQPSSRLFHRFDKLVVL 364
Query: 249 ARGQLMFQGSLKDVGHHLNRMGRKIPKGENPIENLIDVI 287
+RG L++ G + + + +G NP E L+D++
Sbjct: 365 SRGSLLYFGKASEAMSYFSSIGCSPLLAMNPAEFLLDLV 403
Score = 51.2 bits (121), Expect = 2e-06
Identities = 48/234 (20%), Positives = 95/234 (40%), Gaps = 22/234 (9%)
Query: 489 VMMATMFHNPKNTLQGITNRLSFFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRA 548
+++ ++ T Q T F V FF A+ F QER + +E N YR
Sbjct: 515 IILGLLWWQSDITSQRPTRSGLLFFIAVFWGFFPVFTAIFTFPQERAMLSKERESNMYRL 574
Query: 549 SCYTIASLITHMPFLALQALAYAAIVWFALELR---GPFIYFFLVLFISLLSTNSFVVFV 605
S Y +A + +P + + + +V+F LR F L +F+ +++ + +
Sbjct: 575 SAYFVARTTSDLPLDLILPVLFLVVVYFMAGLRLRAESFFLSVLTVFLCIVAAQGLGLAI 634
Query: 606 SSIVPNYILGYAAVIAFTALFFLFCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEYQ 665
+ + + F L GYF+ + +P + W+ +S + Y+ L+ +Y+
Sbjct: 635 GASLMDLKKATTLASVTVMTFMLAGGYFV--KKVPFFIAWIRFMSFNYHTYKLLVKVQYE 692
Query: 666 TNETFGSNDGVSITGFDILKSLHIGTEEIKKRNNVLIMLGWAVLYRILFYIILR 719
+I++S++ G E V ++ + YR++ Y LR
Sbjct: 693 ----------------EIMESVN-GEEIESGLKEVSALVAMIIGYRLVAYFSLR 729
>At2g13610 putative ABC transporter
Length = 649
Score = 196 bits (499), Expect = 3e-50
Identities = 114/257 (44%), Positives = 163/257 (63%), Gaps = 4/257 (1%)
Query: 33 TVTKKKKVDGKWSNEDVDLLHDITGYAPKGCITAVMGPSGAGKSTLLDGLAGRIASGSLK 92
T + K++ + N+ +L +T A I A++GPSGAGKS+LL+ LA R+ +
Sbjct: 44 TEEESLKLEDETGNKVKHVLKGVTCRAKPWEILAIVGPSGAGKSSLLEILAARLIPQT-- 101
Query: 93 GKVSLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYETLMFAADFRLGPLSAVDKRQRVEK 152
G V ++ V+ + K+ S Y+ Q+D LFP+LTV ETL+F+A RL L A + R RV+
Sbjct: 102 GSVYVNKRPVDRANFKKISGYVTQKDTLFPLLTVEETLLFSAKLRL-KLPADELRSRVKS 160
Query: 153 LIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVDIIHGPSLLFLDEPTSGLDSTSALS 212
L+ +LGL + +GD+ RG+SGGERRRVSIGV++IH P +L LDEPTSGLDSTSAL
Sbjct: 161 LVHELGLEAVATARVGDDSVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALL 220
Query: 213 VIEKLHDIAR-NGSTVILTIHQPSSRIQLLLDHLIILARGQLMFQGSLKDVGHHLNRMGR 271
+I+ L +A G T+ILTIHQP RI + +++LA G + QGS+ +G +L G
Sbjct: 221 IIDMLKHMAETRGRTIILTIHQPGFRIVKQFNSVLLLANGSTLKQGSVDQLGVYLRSNGL 280
Query: 272 KIPKGENPIENLIDVIQ 288
P EN +E I+ I+
Sbjct: 281 HPPLHENIVEFAIESIE 297
Score = 188 bits (477), Expect = 1e-47
Identities = 104/274 (37%), Positives = 167/274 (59%), Gaps = 7/274 (2%)
Query: 451 FANSYIGETWILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLS 510
FANS + ET IL R NI RT ELF R + + G+++ +FHN K+ L+G R+
Sbjct: 365 FANSRLEETMILTHRFSKNIFRTKELFACRTVQMLGSGIVLGLIFHNLKDDLKGARERVG 424
Query: 511 FFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAY 570
F F + S+ +A+P F+QER I ++ETS +YR S Y +A+ + ++PFL + A+ +
Sbjct: 425 LFAFILTFLLTSTIEALPIFLQEREILMKETSSGSYRVSSYAVANGLVYLPFLLILAILF 484
Query: 571 AAIVWFALELRG---PFIYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFF 627
+ V++ + L F++F L++++ L + NS VV S++VPN+I+G + + FF
Sbjct: 485 STPVYWLVGLNPSFMAFLHFSLLIWLILYTANSVVVCFSALVPNFIVGNSVISGVMGSFF 544
Query: 628 LFCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEY-QTNETFGSNDG-VSITGFDILK 685
LF GYF+S+ +IP YW +M+ +S YP+EG L+NE+ ++N+ G +T D+LK
Sbjct: 545 LFSGYFISNHEIPGYWIFMHYISLFKYPFEGFLINEFSKSNKCLEYGFGKCLVTEEDLLK 604
Query: 686 SLHIGTEEIKKRNNVLIMLGWAVLYRILFYIILR 719
G E + NV+IML + +LYR + Y+ILR
Sbjct: 605 EERYGEE--SRWRNVVIMLCFVLLYRFISYVILR 636
>At1g53270 putative ABC transporter emb|AAD22683.1
Length = 590
Score = 195 bits (496), Expect = 7e-50
Identities = 109/268 (40%), Positives = 169/268 (62%), Gaps = 7/268 (2%)
Query: 25 LEFESLTYTV----TKKKKVDGKWSN-EDVDLLHDITGYAPKGCITAVMGPSGAGKSTLL 79
LE ++L+Y + K + G S E+ +L D++ A ITA+ GPSGAGK+TLL
Sbjct: 19 LETKNLSYRIGGNTPKFSNLCGLLSEKEEKVILKDVSCDARSAEITAIAGPSGAGKTTLL 78
Query: 80 DGLAGRIASGSLKGKVSLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYETLMFAADFRLG 139
+ LAG+++ G + G+V ++G ++ +R S ++ QED LFP LTV ETL ++A RL
Sbjct: 79 EILAGKVSHGKVSGQVLVNGRPMDGPEYRRVSGFVPQEDALFPFLTVQETLTYSALLRL- 137
Query: 140 PLSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVDIIHGPSLLFLD 199
D +V++LI++LGL ++ IG G+SGGERRRVSIGV+++H P+++ +D
Sbjct: 138 KTKRKDAAAKVKRLIQELGLEHVADSRIGQGSRSGISGGERRRVSIGVELVHDPNVILID 197
Query: 200 EPTSGLDSTSALSVIEKLHDIA-RNGSTVILTIHQPSSRIQLLLDHLIILARGQLMFQGS 258
EPTSGLDS SAL V+ L D+ + G T++LTIHQP RI +D +++L+ G ++ GS
Sbjct: 198 EPTSGLDSASALQVVTLLKDMTIKQGKTIVLTIHQPGFRILEQIDRIVLLSNGMVVQNGS 257
Query: 259 LKDVGHHLNRMGRKIPKGENPIENLIDV 286
+ + + G +IP+ N +E ID+
Sbjct: 258 VYSLHQKIKFSGHQIPRRVNVLEYAIDI 285
Score = 164 bits (415), Expect = 2e-40
Identities = 95/286 (33%), Positives = 157/286 (54%), Gaps = 20/286 (6%)
Query: 446 HLGPKFANSYIGETWILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTL-QG 504
H +NS + E IL +R+ NI RT +LF +R + + G+++ +++ N N +
Sbjct: 321 HQSDSHSNSVLEEVQILGQRSCKNIFRTKQLFTTRALQASIAGLILGSIYLNVGNQKKEA 380
Query: 505 ITNRLSFFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLA 564
R FF F + S+ + +P F+Q+R I +RETS AYR Y +A + +PFL
Sbjct: 381 KVLRTGFFAFILTFLLSSTTEGLPIFLQDRRILMRETSRRAYRVLSYVLADTLIFIPFLL 440
Query: 565 LQALAYAAIVWFALELRGP---FIYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIA 621
+ ++ +A V++ + LR F+YF LV++I LL +NSFV S++VPN+I+G + +
Sbjct: 441 IISMLFATPVYWLVGLRRELDGFLYFSLVIWIVLLMSNSFVACFSALVPNFIMGTSVISG 500
Query: 622 FTALFFLFCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEYQTNETFGSNDGVSITGF 681
FFLF GYF++ + IP+YW +M+ +S YP+E L++NEY+ + D
Sbjct: 501 LMGSFFLFSGYFIAKDRIPVYWEFMHYLSLFKYPFECLMINEYRGDVFLKQQD------- 553
Query: 682 DILKSLHIGTEEIKKRNNVLIMLGWAVLYRILFYIILRFASKNQRS 727
+E +K +N+ IM + V YR+L + IL + RS
Sbjct: 554 ---------LKESQKWSNLGIMASFIVGYRVLGFFILWYRCYRTRS 590
>At1g31770 unknown protein
Length = 648
Score = 191 bits (484), Expect = 2e-48
Identities = 106/265 (40%), Positives = 164/265 (61%), Gaps = 5/265 (1%)
Query: 25 LEFESLTYTVT--KKKKVDGKWSNEDVDLLHDITGYAPKGCITAVMGPSGAGKSTLLDGL 82
L+FE + Y V + + G W +++ +L+ ITG G A++GPSG+GK+TLL L
Sbjct: 53 LKFEEVVYKVKIEQTSQCMGSWKSKEKTILNGITGMVCPGEFLAMLGPSGSGKTTLLSAL 112
Query: 83 AGRIASGSLKGKVSLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYETLMFAADFRL-GPL 141
GR+ S + GKV +G + IKR + ++ Q+D L+P LTV+ETL F A RL L
Sbjct: 113 GGRL-SKTFSGKVMYNGQPFSGC-IKRRTGFVAQDDVLYPHLTVWETLFFTALLRLPSSL 170
Query: 142 SAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVDIIHGPSLLFLDEP 201
+ +K + V+++I +LGL+ N+ IG RG+SGGE++RVSIG +++ PSLL LDEP
Sbjct: 171 TRDEKAEHVDRVIAELGLNRCTNSMIGGPLFRGISGGEKKRVSIGQEMLINPSLLLLDEP 230
Query: 202 TSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRIQLLLDHLIILARGQLMFQGSLKD 261
TSGLDST+A ++ + +A G TV+ TIHQPSSRI + D +++L+ G ++ G+
Sbjct: 231 TSGLDSTTAHRIVTTIKRLASGGRTVVTTIHQPSSRIYHMFDKVVLLSEGSPIYYGAASS 290
Query: 262 VGHHLNRMGRKIPKGENPIENLIDV 286
+ + +G NP + L+D+
Sbjct: 291 AVEYFSSLGFSTSLTVNPADLLLDL 315
Score = 68.6 bits (166), Expect = 1e-11
Identities = 63/249 (25%), Positives = 118/249 (47%), Gaps = 15/249 (6%)
Query: 475 ELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLSFFIFTVCLFFFSSNDAVPAFIQER 534
+L + +++ + F+G ++ +H PK+ +Q T L F F+V F+ +AV F QE+
Sbjct: 404 KLRIFQVISVAFLGGLL--WWHTPKSHIQDRTALL--FFFSVFWGFYPLYNAVFTFPQEK 459
Query: 535 FIFIRETSHNAYRASCYTIASLITHMPFLALQALAYAAIVWFALELR---GPFIYFFLVL 591
+ I+E S YR S Y +A + +P A+ I+++ L+ FI LV+
Sbjct: 460 RMLIKERSSGMYRLSSYFMARNVGDLPLELALPTAFVFIIYWMGGLKPDPTTFILSLLVV 519
Query: 592 FISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFFLFCGYFLSSEDIPLYWRWMNKVST 651
S+L + +++ N T +F + GY++ + IP + W+ +S
Sbjct: 520 LYSVLVAQGLGLAFGALLMNIKQATTLASVTTLVFLIAGGYYV--QQIPPFIVWLKYLSY 577
Query: 652 MTYPYEGLLMNEYQTNETFGSNDGV--SITGFDILKSLHIGTEEIKKRNNVLIMLGWAVL 709
Y Y+ LL +Y ++ + + GV + F +KS+ + I +V +M V
Sbjct: 578 SYYCYKLLLGIQYTDDDYYECSKGVWCRVGDFPAIKSMGLNNLWI----DVFVMGVMLVG 633
Query: 710 YRILFYIIL 718
YR++ Y+ L
Sbjct: 634 YRLMAYMAL 642
>At1g17840 putative ABC transporter (At1g17840)
Length = 703
Score = 189 bits (480), Expect = 5e-48
Identities = 117/276 (42%), Positives = 163/276 (58%), Gaps = 15/276 (5%)
Query: 18 PVSFTGG----LEFESLTYTVTKKKKVDGKWSNEDVDLLHDITGYAPKGCITAVMGPSGA 73
P F G L ++ LT VT DG+ N +L +TGYA G +TA+MGPSG+
Sbjct: 39 PTEFVGDVSARLTWQDLTVMVTMG---DGETQN----VLEGLTGYAEPGSLTALMGPSGS 91
Query: 74 GKSTLLDGLAGRIASGS-LKGKVSLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYETLMF 132
GKST+LD LA R+A+ + L G V L+G S T+AY+ Q+D L LTV ET+ +
Sbjct: 92 GKSTMLDALASRLAANAFLSGTVLLNGRKTKLSF--GTAAYVTQDDNLIGTLTVRETIWY 149
Query: 133 AADFRL-GPLSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVDIIH 191
+A RL + +KR VE+ I ++GL +T IG+ RG+SGGE+RRVSI ++I+
Sbjct: 150 SARVRLPDKMLRSEKRALVERTIIEMGLQDCADTVIGNWHLRGISGGEKRRVSIALEILM 209
Query: 192 GPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRIQLLLDHLIILARG 251
P LLFLDEPTSGLDS SA V + L ++R+G TVI +IHQPSS + L D L +L+ G
Sbjct: 210 RPRLLFLDEPTSGLDSASAFFVTQTLRALSRDGRTVIASIHQPSSEVFELFDRLYLLSGG 269
Query: 252 QLMFQGSLKDVGHHLNRMGRKIPKGENPIENLIDVI 287
+ ++ G D + G P NP ++ + I
Sbjct: 270 KTVYFGQASDAYEFFAQAGFPCPALRNPSDHFLRCI 305
Score = 105 bits (262), Expect = 9e-23
Identities = 68/271 (25%), Positives = 132/271 (48%), Gaps = 5/271 (1%)
Query: 454 SYIGETWILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLSFFI 513
S++ +T+ L +R+F N+ R + RL++ + V + T++ N + I R S
Sbjct: 379 SFLLQTYTLTKRSFINMSRDFGYYWLRLLIYILVTVCIGTIYLNVGTSYSAILARGSCAS 438
Query: 514 FTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAYAAI 573
F F S P+F+++ +F RE + Y + + IA+ ++ PFL + I
Sbjct: 439 FVFGFVTFMSIGGFPSFVEDMKVFQRERLNGHYGVAAFVIANTLSATPFLIMITFISGTI 498
Query: 574 VWFALELRGPF---IYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFFLFC 630
+F + L F ++F L L+ S+ S ++ ++SIVPN+++G +F L
Sbjct: 499 CYFMVGLHPGFTHYLFFVLCLYASVTVVESLMMAIASIVPNFLMGIIIGAGIQGIFMLVS 558
Query: 631 GYFLSSEDIPL-YWRW-MNKVSTMTYPYEGLLMNEYQTNETFGSNDGVSITGFDILKSLH 688
G+F DIP +WR+ M+ +S + +G N+ + I G +L+++
Sbjct: 559 GFFRLPNDIPKPFWRYPMSYISFHFWALQGQYQNDLRGLTFDSQGSAFKIPGEYVLENVF 618
Query: 689 IGTEEIKKRNNVLIMLGWAVLYRILFYIILR 719
K N+ ++L ++YRI+F+I+++
Sbjct: 619 QIDLHRSKWINLSVILSMIIIYRIIFFIMIK 649
>At1g71960 putative ABC transporter
Length = 662
Score = 187 bits (475), Expect = 2e-47
Identities = 107/263 (40%), Positives = 160/263 (60%), Gaps = 3/263 (1%)
Query: 36 KKKKVDGKWSNEDVDLLHDITGYAPKGCITAVMGPSGAGKSTLLDGLAGRIASGSLKGKV 95
K+K D S E+ +L +TG G AV+GPSG+GKSTLL+ +AGR+ +L GK+
Sbjct: 68 KQKPSDETRSTEERTILSGVTGMISPGEFMAVLGPSGSGKSTLLNAVAGRLHGSNLTGKI 127
Query: 96 SLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYETLMFAADFRLGPLSAVDKRQRV-EKLI 154
++ + +KRT ++ Q+D L+P LTV ETL+F A RL D + R E +I
Sbjct: 128 LINDGKITKQTLKRTG-FVAQDDLLYPHLTVRETLVFVALLRLPRSLTRDVKLRAAESVI 186
Query: 155 EQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVDIIHGPSLLFLDEPTSGLDSTSALSVI 214
+LGL+ NT +G+ RG+SGGER+RVSI +++ PSLL LDEPTSGLD+T+AL ++
Sbjct: 187 SELGLTKCENTVVGNTFIRGISGGERKRVSIAHELLINPSLLVLDEPTSGLDATAALRLV 246
Query: 215 EKLHDIAR-NGSTVILTIHQPSSRIQLLLDHLIILARGQLMFQGSLKDVGHHLNRMGRKI 273
+ L +A G TV+ +IHQPSSR+ + D +++L+ G+ +F G +D + +G
Sbjct: 247 QTLAGLAHGKGKTVVTSIHQPSSRVFQMFDTVLLLSEGKCLFVGKGRDAMAYFESVGFSP 306
Query: 274 PKGENPIENLIDVIQEYDQCDFV 296
NP + L+D+ Q D V
Sbjct: 307 AFPMNPADFLLDLANGVCQTDGV 329
Score = 51.2 bits (121), Expect = 2e-06
Identities = 63/283 (22%), Positives = 119/283 (41%), Gaps = 26/283 (9%)
Query: 454 SYIGETWILMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLSFFI 513
++ + IL+ R RR L R+ + ++ M+ + + + + +RL
Sbjct: 385 TWFSQLCILLHRLLKE-RRHESFDLLRIFQVVAASILCGLMWWH--SDYRDVHDRLGLLF 441
Query: 514 FTVCLFF--FSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAYA 571
F + +F+ S +AV F QER IF RE + Y S Y +A ++ + + ++
Sbjct: 442 F-ISIFWGVLPSFNAVFTFPQERAIFTRERASGMYTLSSYFMAHVLGSLSMELVLPASFL 500
Query: 572 AIVWFALELRG---PFIYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFFL 628
++ + LR PF+ VL + +L++ + + + + + V F L
Sbjct: 501 TFTYWMVYLRPGIVPFLLTLSVLLLYVLASQGLGLALGAAIMDAKKASTIVTVTMLAFVL 560
Query: 629 FCGYFLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEYQTNETF------------GSNDGV 676
GY+++ +P WM VST Y Y L+ +Y + E G++
Sbjct: 561 TGGYYVNK--VPSGMVWMKYVSTTFYCYRLLVAIQYGSGEEILRMLGCDSKGKQGASAAT 618
Query: 677 SITGFDILKSLHIGTEEIKKRNNVLIMLGWAVLYRILFYIILR 719
S G ++ IG + VL ++ + YR+L Y+ LR
Sbjct: 619 S-AGCRFVEEEVIGDVGMWTSVGVLFLMFFG--YRVLAYLALR 658
>At3g25620 membrane transporter, putative
Length = 646
Score = 184 bits (467), Expect = 2e-46
Identities = 110/287 (38%), Positives = 169/287 (58%), Gaps = 16/287 (5%)
Query: 25 LEFESLTYTVTKKKKVDGKWSNEDVD-----LLHDITGYAPKGCITAVMGPSGAGKSTLL 79
L+FE LTY++ + W +L ++G G + A++GPSG+GK+TL+
Sbjct: 67 LKFEELTYSIKSQTGKGSYWFGSQEPKPNRLVLKCVSGIVKPGELLAMLGPSGSGKTTLV 126
Query: 80 DGLAGRIASGSLKGKVSLDGNSVNASLIKRTSAYIMQEDRLFPMLTVYETLMFAADFRLG 139
LAGR+ G L G VS +G +S +KR + ++ Q+D L+P LTV ETL + A RL
Sbjct: 127 TALAGRL-QGKLSGTVSYNGEPFTSS-VKRKTGFVTQDDVLYPHLTVMETLTYTALLRLP 184
Query: 140 P-LSAVDKRQRVEKLIEQLGLSSSRNTYIGDEGTRGVSGGERRRVSIGVDIIHGPSLLFL 198
L+ +K ++VE ++ LGL+ N+ IG RG+SGGER+RVSIG +++ PSLL L
Sbjct: 185 KELTRKEKLEQVEMVVSDLGLTRCCNSVIGGGLIRGISGGERKRVSIGQEMLVNPSLLLL 244
Query: 199 DEPTSGLDSTSALSVIEKLHDIARNGSTVILTIHQPSSRIQLLLDHLIILARGQLMFQGS 258
DEPTSGLDST+A ++ L +AR G TV+ TIHQPSSR+ + D +++L+ G ++ G
Sbjct: 245 DEPTSGLDSTTAARIVATLRSLARGGRTVVTTIHQPSSRLYRMFDKVLVLSEGCPIYSGD 304
Query: 259 LKDVGHHLNRMGRKIPKG-ENPIENLIDV-------IQEYDQCDFVG 297
V + +G + NP + ++D+ ++YDQ + G
Sbjct: 305 SGRVMEYFGSIGYQPGSSFVNPADFVLDLANGITSDTKQYDQIETNG 351
Score = 67.4 bits (163), Expect = 3e-11
Identities = 67/263 (25%), Positives = 118/263 (44%), Gaps = 18/263 (6%)
Query: 462 LMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLSFFIFTVCLFFF 521
L ++ TN R P + + VL G++ +H+ LQ L F F++ FF
Sbjct: 395 LRKKAITN--RWPTSWWMQFSVLLKRGLLW---WHSRVAHLQDQVGLL--FFFSIFWGFF 447
Query: 522 SSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAYAAIVWFALELR 581
+A+ F QER + I+E S YR S Y IA + +P + + I ++ L+
Sbjct: 448 PLFNAIFTFPQERPMLIKERSSGIYRLSSYYIARTVGDLPMELILPTIFVTITYWMGGLK 507
Query: 582 GPFIYFFLVLFI---SLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFFLFCGYFLSSED 638
F + L I ++L + + +I+ + +F L GY++ +
Sbjct: 508 PSLTTFIMTLMIVLYNVLVAQGVGLALGAILMDAKKAATLSSVLMLVFLLAGGYYI--QH 565
Query: 639 IPLYWRWMNKVSTMTYPYEGLLMNEYQTNETFGSNDGV--SITGFDILKSLHIGTEEIKK 696
IP + W+ VS Y Y+ L+ +Y +E + G+ S+ ++ +K+L IG
Sbjct: 566 IPGFIAWLKYVSFSHYCYKLLVGVQYTWDEVYECGSGLHCSVMDYEGIKNLRIG----NM 621
Query: 697 RNNVLIMLGWAVLYRILFYIILR 719
+VL + +LYR+L Y+ LR
Sbjct: 622 MWDVLALAVMLLLYRVLAYLALR 644
>At3g21090 ABC transporter, putative
Length = 680
Score = 183 bits (464), Expect = 3e-46
Identities = 115/268 (42%), Positives = 154/268 (56%), Gaps = 8/268 (2%)
Query: 51 LLHDITGYAPKGCITAVMGPSGAGKSTLLDGLAGRIASGS-LKGKVSLDGNSVNASLIKR 109
LL + GYA G I A+MGPSG+GKSTLLD LAGR+A + G + L+G A L
Sbjct: 45 LLQRLNGYAEPGRIMAIMGPSGSGKSTLLDSLAGRLARNVVMTGNLLLNGKK--ARLDYG 102
Query: 110 TSAYIMQEDRLFPMLTVYETLMFAADFRL-GPLSAVDKRQRVEKLIEQLGLSSSRNTYIG 168
AY+ QED L LTV ET+ ++A RL +S + VE I +LGL + IG
Sbjct: 103 LVAYVTQEDVLLGTLTVRETITYSAHLRLPSDMSKEEVSDIVEGTIMELGLQDCSDRVIG 162
Query: 169 DEGTRGVSGGERRRVSIGVDIIHGPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVI 228
+ RGVSGGER+RVSI ++I+ P +LFLDEPTSGLDS SA VI+ L +IAR+G TVI
Sbjct: 163 NWHARGVSGGERKRVSIALEILTRPQILFLDEPTSGLDSASAFFVIQALRNIARDGRTVI 222
Query: 229 LTIHQPSSRIQLLLDHLIILARGQLMFQGSLKDVGHHLNRMGRKIPKGENPIENLIDVIQ 288
++HQPSS + L D L +L+ G+ ++ G K G PK NP ++ + I
Sbjct: 223 SSVHQPSSEVFALFDDLFLLSSGESVYFGEAKSAVEFFAESGFPCPKKRNPSDHFLRCIN 282
Query: 289 EYDQCDFVGVEVLAEFARTGMKPPLLSD 316
DF V + ++ + P SD
Sbjct: 283 S----DFDTVTATLKGSQRIQETPATSD 306
Score = 83.6 bits (205), Expect = 4e-16
Identities = 59/266 (22%), Positives = 123/266 (46%), Gaps = 13/266 (4%)
Query: 462 LMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLSFFIFTVCLFFF 521
L R+F N+ R + +R++ + + + T+F++ + I R+S F F
Sbjct: 365 LTARSFINMCRDVGYYWTRIISYIVVSISVGTIFYDVGYSYTSILARVSCGGFITGFMTF 424
Query: 522 SSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAYAAIVWFALELR 581
S P+F++E +F +E Y S Y +++ I+ PFL ++ I + ++ R
Sbjct: 425 MSIGGFPSFLEEMKVFYKERLSGYYGVSVYILSNYISSFPFLVAISVITGTITYNLVKFR 484
Query: 582 GPF---IYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFFLFCGYFLSSED 638
F +F L +F S+ S ++ V+S+VPN+++G + + G+F D
Sbjct: 485 PGFSHYAFFCLNIFFSVSVIESLMMVVASVVPNFLMGLITGAGLIGIIMMTSGFFRLLPD 544
Query: 639 IP-LYWRWMNKVSTMTYPYEGLLMNEYQTNETFGSNDGVSITGFDILKSLHIGTEEIKKR 697
+P ++WR+ VS ++Y + E +TG ++++ + K
Sbjct: 545 LPKIFWRY--PVSYISYGSWAIQFEPLFPGEP-------KMTGEEVIEKVFGVKVTYSKW 595
Query: 698 NNVLIMLGWAVLYRILFYIILRFASK 723
++ ++ V YR+LF+++L+ +
Sbjct: 596 WDLAAVVAILVCYRLLFFVVLKLRER 621
>At5g19410 membrane transporter - like protein
Length = 624
Score = 175 bits (443), Expect = 9e-44
Identities = 99/244 (40%), Positives = 160/244 (65%), Gaps = 6/244 (2%)
Query: 51 LLHDITGYAPKGCITAVMGPSGAGKSTLLDGLAGRIASGSLKGKVSLDGNS---VNASLI 107
+L+ ++ A I AV+GPSG GKSTLL ++GR+ +L ++ N+ + + +
Sbjct: 66 ILNSVSLAAESSKILAVVGPSGTGKSTLLKIISGRVNHKALDPSSAVLMNNRKITDYNQL 125
Query: 108 KRTSAYIMQEDRLFPMLTVYETLMFAADFRLGPLSAVDKRQRVEKLIEQLGLSSSRNTYI 167
+R ++ Q+D L P+LTV ETLM++A F L +A ++ +RVE L+ LGL +++++
Sbjct: 126 RRLCGFVPQDDDLLPLLTVKETLMYSAKFSLRDSTAKEREERVESLLSDLGLVLVQDSFV 185
Query: 168 G--DEGTRGVSGGERRRVSIGVDIIHGPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGS 225
G DE RGVSGGER+RVSI V++I P +L LDEPTSGLDS ++L V+E L +A++
Sbjct: 186 GEGDEEDRGVSGGERKRVSIAVEMIRDPPILLLDEPTSGLDSRNSLQVVELLATMAKSKQ 245
Query: 226 -TVILTIHQPSSRIQLLLDHLIILARGQLMFQGSLKDVGHHLNRMGRKIPKGENPIENLI 284
TV+ +IHQPS RI + +IL+RG ++ GSL+ + + ++G +IP+ NPIE +
Sbjct: 246 RTVLFSIHQPSYRILDYISDYLILSRGSVIHLGSLEHLEDSIAKLGFQIPEQLNPIEFAM 305
Query: 285 DVIQ 288
++++
Sbjct: 306 EIVE 309
Score = 157 bits (398), Expect = 2e-38
Identities = 105/322 (32%), Positives = 175/322 (53%), Gaps = 12/322 (3%)
Query: 413 YSSEILPATPTPHSSDYVVDENDYLTPTNSSQEHLGPK---FANSYIGETWILMRRNFTN 469
++ EI+ + T + V E+ + P N+ + + K F + E L R
Sbjct: 303 FAMEIVESLRTFKPNSVAVVESSSMWPENNENDGIISKKEAFRVLDVTEISYLCSRFCKI 362
Query: 470 IRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGITNRLSFFIFTVCLFFFSSNDAVPA 529
I RT +LFL+R M G+ + +++ K +G+ RL F F++ S+ +A+P
Sbjct: 363 IYRTKQLFLARTMQAVVAGLGLGSVYTRLKRDEEGVAERLGLFAFSLSFLLSSTVEALPI 422
Query: 530 FIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAYAAIVWFALELR---GPFIY 586
+++ER + ++E+S +YR S Y IA+ I +PFL + +L ++ V++ + L F +
Sbjct: 423 YLRERRVLMKESSRGSYRISSYMIANTIAFVPFLFVVSLLFSIPVYWIVGLNPSIQAFSF 482
Query: 587 FFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFFLFCGYFLSSEDIPLYWRWM 646
F L +++ +L +S V+F+S++ P++I G + + FFLF GYF+ E IP W +M
Sbjct: 483 FVLCVWLIILMASSLVLFLSAVSPDFISGNSLICTVLGAFFLFSGYFIPKEKIPKPWMFM 542
Query: 647 NKVSTMTYPYEGLLMNEY--QTNETFGS-NDGVSITGFDILKSLHIGTEEIKKRNNVLIM 703
VS YP E +++NEY E F S N G +TG D+LK G ++ + NV IM
Sbjct: 543 YYVSLYRYPLESMVVNEYWSMREECFSSGNMGCLMTGEDVLKER--GLDKDTRWINVGIM 600
Query: 704 LGWAVLYRILFY-IILRFASKN 724
L + V YRIL + I+LR ASK+
Sbjct: 601 LAFFVFYRILCWGILLRKASKS 622
>At1g51500 ATP-dependent transmembrane transporter, putative
Length = 687
Score = 175 bits (443), Expect = 9e-44
Identities = 113/269 (42%), Positives = 155/269 (57%), Gaps = 9/269 (3%)
Query: 51 LLHDITGYAPKGCITAVMGPSGAGKSTLLDGLAGRIASGSLK-GKVSLDGNSVNASLIKR 109
LL + G+A G I A+MGPSG+GKSTLLD LAGR+A + G + L+G A L
Sbjct: 44 LLDGLNGHAEPGRIMAIMGPSGSGKSTLLDSLAGRLARNVIMTGNLLLNGKK--ARLDYG 101
Query: 110 TSAYIMQEDRLFPMLTVYETLMFAADFRLGP-LSAVDKRQRVEKLIEQLGLSSSRNTYIG 168
AY+ QED L LTV ET+ ++A RL L+ + VE I +LGL + IG
Sbjct: 102 LVAYVTQEDILMGTLTVRETITYSAHLRLSSDLTKEEVNDIVEGTIIELGLQDCADRVIG 161
Query: 169 DEGTRGVSGGERRRVSIGVDIIHGPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGS-TV 227
+ +RGVSGGER+RVS+ ++I+ P +LFLDEPTSGLDS SA VI+ L +IAR+G TV
Sbjct: 162 NWHSRGVSGGERKRVSVALEILTRPQILFLDEPTSGLDSASAFFVIQALRNIARDGGRTV 221
Query: 228 ILTIHQPSSRIQLLLDHLIILARGQLMFQGSLKDVGHHLNRMGRKIPKGENPIENLIDVI 287
+ +IHQPSS + L D L +L+ G+ ++ G K G PK NP ++ + I
Sbjct: 222 VSSIHQPSSEVFALFDDLFLLSSGETVYFGESKFAVEFFAEAGFPCPKKRNPSDHFLRCI 281
Query: 288 QEYDQCDFVGVEVLAEFARTGMKPPLLSD 316
DF V + ++ + P SD
Sbjct: 282 NS----DFDTVTATLKGSQRIRETPATSD 306
Score = 92.4 bits (228), Expect = 8e-19
Identities = 79/343 (23%), Positives = 156/343 (45%), Gaps = 13/343 (3%)
Query: 393 LKNENKVQKPPSHASPGIYTYSSEILPATPTPHS-SDYVVDENDYLTPTNSSQEHLGPKF 451
LK ++++ P+ + P + +SEI + S Y + S + H G +
Sbjct: 292 LKGSQRIRETPATSDPLMNLATSEIKARLVENYRRSVYAKSAKSRIRELASIEGHHGMEV 351
Query: 452 ANSYIGETWI-----LMRRNFTNIRRTPELFLSRLMVLTFMGVMMATMFHNPKNTLQGIT 506
TW L +R+F N+ R + SR+++ + + T+F++ ++ I
Sbjct: 352 RKGSEA-TWFKQLRTLTKRSFVNMCRDIGYYWSRIVIYIVVSFCVGTIFYDVGHSYTSIL 410
Query: 507 NRLSFFIFTVCLFFFSSNDAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQ 566
R+S F F S P+FI+E +F +E Y S Y I++ ++ PFL
Sbjct: 411 ARVSCGGFITGFMTFMSIGGFPSFIEEMKVFYKERLSGYYGVSVYIISNYVSSFPFLVAI 470
Query: 567 ALAYAAIVWFALELR---GPFIYFFLVLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFT 623
AL +I + ++ R + +F L +F S+ S ++ V+S+VPN+++G
Sbjct: 471 ALITGSITYNMVKFRPGVSHWAFFCLNIFFSVSVIESLMMVVASLVPNFLMGLITGAGII 530
Query: 624 ALFFLFCGYFLSSEDIP-LYWRW-MNKVSTMTYPYEGLLMNEYQTNETFGSNDG-VSITG 680
+ + G+F D+P ++WR+ ++ +S ++ +G N++ E G +TG
Sbjct: 531 GIIMMTSGFFRLLPDLPKVFWRYPISFMSYGSWAIQGAYKNDFLGLEFDPMFAGEPKMTG 590
Query: 681 FDILKSLHIGTEEIKKRNNVLIMLGWAVLYRILFYIILRFASK 723
++ + K ++ ++ V YRILF+I+L+ +
Sbjct: 591 EQVINKIFGVQVTHSKWWDLSAIVLILVCYRILFFIVLKLKER 633
>At3g13220 ABC transporter
Length = 647
Score = 174 bits (442), Expect = 1e-43
Identities = 103/238 (43%), Positives = 150/238 (62%), Gaps = 5/238 (2%)
Query: 51 LLHDITGYAPKGCITAVMGPSGAGKSTLLDGLAGRIASGSLKGKVSLDGNSVNASLIKRT 110
+L ITG G I A+MGPSG+GK+TLL + GR+ ++KGK++ + + S +KR
Sbjct: 68 ILKGITGSTGPGEILALMGPSGSGKTTLLKIMGGRLTD-NVKGKLTYNDIPYSPS-VKRR 125
Query: 111 SAYIMQEDRLFPMLTVYETLMFAADFRL-GPLSAVDKRQRVEKLIEQLGLSSSRNTYIGD 169
++ Q+D L P LTV ETL FAA RL +S K ++E +I++LGL R T +G
Sbjct: 126 IGFVTQDDVLLPQLTVEETLAFAAFLRLPSSMSKEQKYAKIEMIIKELGLERCRRTRVGG 185
Query: 170 EGTRGVSGGERRRVSIGVDIIHGPSLLFLDEPTSGLDSTSALSVIEKLHDIARNGSTVIL 229
+G+SGGER+R SI +I+ PSLL LDEPTSGLDSTSA ++ L +A+ G TVI
Sbjct: 186 GFVKGISGGERKRASIAYEILVDPSLLLLDEPTSGLDSTSATKLLHILQGVAKAGRTVIT 245
Query: 230 TIHQPSSRIQLLLDHLIILARGQLMFQGSLKDVGHHLNRMGRKIPK-GENPIENLIDV 286
TIHQPSSR+ + D L++++ G F G ++ + + + R +P+ NP E L+D+
Sbjct: 246 TIHQPSSRMFHMFDKLLLISEGHPAFYGKARESMEYFSSL-RILPEIAMNPAEFLLDL 302
Score = 67.8 bits (164), Expect = 2e-11
Identities = 70/273 (25%), Positives = 120/273 (43%), Gaps = 27/273 (9%)
Query: 461 ILMRRNFTNIRRTPELFLSRLMVLTFMGV--MMATMFHNPKNTLQGITNRLSFFIFTVCL 518
IL RR F RR + +L ++ +GV ++ ++ K + +F +C+
Sbjct: 380 ILSRRTFRERRRD---YFDKLRLVQSLGVAVVLGLLWWKSKTDTEAHLRDQVGLMFYICI 436
Query: 519 FFFSSN--DAVPAFIQERFIFIRETSHNAYRASCYTIASLITHMPFLALQALAYAAIVWF 576
F+ SS+ AV F E+ ++E YR S Y + S + M L + IV+F
Sbjct: 437 FWTSSSLFGAVYVFPFEKIYLVKERKAEMYRLSVYYVCSTLCDMVAHVLYPTFFMIIVYF 496
Query: 577 ALELRGPFIYFFL----VLFISLLSTNSFVVFVSSIVPNYILGYAAVIAFTALFFLFCGY 632
E F +L I++ S + +S++ G A + LF L GY
Sbjct: 497 MAEFNRNIPCFLFTVLTILLIAITSQGAGEFLGASVLSIKRAGMIASLVL-MLFLLTGGY 555
Query: 633 FLSSEDIPLYWRWMNKVSTMTYPYEGLLMNEYQTNETF--GSNDGV----SITGFDILKS 686
++ + IP + +W+ +S M Y + LL +Y ++ F GS G S + FD + +
Sbjct: 556 YV--QHIPKFMQWLKYLSFMHYGFRLLLKVQYSADQLFECGSKGGCRTLQSSSSFDTI-N 612
Query: 687 LHIGTEEIKKRNNVLIMLGWAVLYRILFYIILR 719
L+ G +E+ ++L A YR+ Y LR
Sbjct: 613 LNGGLQEL------WVLLAMAFGYRLCAYFCLR 639
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.137 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,131,521
Number of Sequences: 26719
Number of extensions: 716880
Number of successful extensions: 2638
Number of sequences better than 10.0: 143
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 33
Number of HSP's that attempted gapping in prelim test: 2143
Number of HSP's gapped (non-prelim): 291
length of query: 727
length of database: 11,318,596
effective HSP length: 107
effective length of query: 620
effective length of database: 8,459,663
effective search space: 5244991060
effective search space used: 5244991060
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC126010.8


