

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
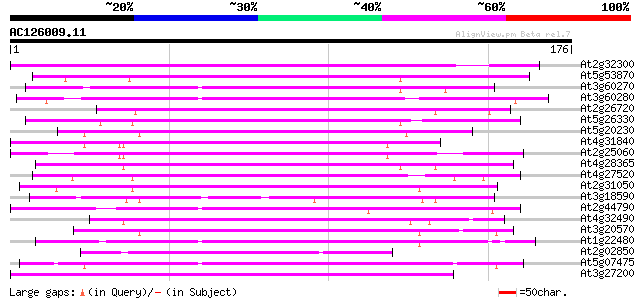
Query= AC126009.11 - phase: 0
(176 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g32300 uclacyanin I - like predicted GPI-anchored protein 96 1e-20
At5g53870 predicted GPI-anchored protein 90 5e-19
At3g60270 stellacyanin (uclacyanin 3) - like predicted GPI-ancho... 90 7e-19
At3g60280 uclacyanin 3 88 3e-18
At2g26720 putative phytocyanin, predicted GPI-anchored protein 85 2e-17
At5g26330 copper binding protein - like, predicted GPI-anchored ... 85 2e-17
At5g20230 blue copper binding protein (bcb) 84 5e-17
At4g31840 predicted GPI-anchored protein 77 5e-15
At2g25060 early nodulin-like predicted GPI-anchored protein 77 5e-15
At4g28365 unknown protein 76 8e-15
At4g27520 early nodulin-like 2 predicted GPI-anchored protein 76 8e-15
At2g31050 phytocyanin - like predicted GPI-anchored protein 76 8e-15
At3g18590 predicted GPI-anchored protein 75 2e-14
At2g44790 phytocyanin, blue copper-binding protein II 73 9e-14
At4g32490 nodulin - like predicted GPI-anchored protein 72 2e-13
At3g20570 predicted GPI-anchored protein 72 2e-13
At1g22480 blue copper protein precursor-like predicted GPI-ancho... 70 4e-13
At2g02850 putative basic blue protein (plantacyanin) 70 6e-13
At5g07475 putative protein 69 1e-12
At3g27200 blue copper protein, putative 69 2e-12
>At2g32300 uclacyanin I - like predicted GPI-anchored protein
Length = 261
Score = 95.5 bits (236), Expect = 1e-20
Identities = 53/166 (31%), Positives = 78/166 (46%), Gaps = 10/166 (6%)
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
MA L + +++A+ + VA D +G GWT +TW A + F +GD L F+Y
Sbjct: 1 MASREMLIIISVLATTLIGLTVATDHTIGGPSGWTVGASLRTWAAGQTFAVGDNLVFSYP 60
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
+VV V +F SC +G + +TT G+R++I + HC G KL +
Sbjct: 61 AAFHDVVEVTKPEFDSCQAVKPLITFANGNSLVPLTTPGKRYFICGMPGHCSQGMKLEVN 120
Query: 121 VQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRSLLP 166
V P P P+ PSL NAP P+S +P + LLP
Sbjct: 121 VVPTATVAPTAPLPNTVPSL----------NAPSPSSVLPIQPLLP 156
>At5g53870 predicted GPI-anchored protein
Length = 370
Score = 90.1 bits (222), Expect = 5e-19
Identities = 57/165 (34%), Positives = 82/165 (49%), Gaps = 9/165 (5%)
Query: 8 FLFALIASI--FSTMAVAKDFVVGDEKGWTT--LFDYQTWTANKVFRLGDTLTFNYVGGK 63
F F ++AS F ++A A F VG W T +Y TW F++ D+L F Y G
Sbjct: 10 FSFLILASFATFFSVADAWRFNVGGNGAWVTNPQENYNTWAERNRFQVNDSLYFKYAKGS 69
Query: 64 DNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV-- 121
D+V +V +DF C+V +G+ + + G ++IS DHC+ GQKL + V
Sbjct: 70 DSVQQVMKADFDGCNVRNPIKNFENGESVVTLDRSGAFYFISGNQDHCQKGQKLIVVVLA 129
Query: 122 ---QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRS 163
QP SPVPS SP+ +P +P + A P+ S P RS
Sbjct: 130 VRNQPSAPAHSPVPSVSPTQPPKSHSPVSPVAPASAPSKSQPPRS 174
Score = 26.2 bits (56), Expect = 9.6
Identities = 13/33 (39%), Positives = 21/33 (63%), Gaps = 1/33 (3%)
Query: 129 SPVPSPSPSPSLDLVTPEAP-PSNAPWPASSVP 160
+P SP+ SPS TP++P PS++P + + P
Sbjct: 260 APSHSPAHSPSHSPATPKSPSPSSSPAQSPATP 292
Score = 26.2 bits (56), Expect = 9.6
Identities = 15/36 (41%), Positives = 18/36 (49%), Gaps = 1/36 (2%)
Query: 132 PSPSPSPSLDLVTPEAPPSNAPWPASS-VPRRSLLP 166
PSPS SP+ TP +P P SS P +S P
Sbjct: 279 PSPSSSPAQSPATPSPMTPQSPSPVSSPSPDQSAAP 314
>At3g60270 stellacyanin (uclacyanin 3) - like predicted GPI-anchored
protein
Length = 187
Score = 89.7 bits (221), Expect = 7e-19
Identities = 55/154 (35%), Positives = 74/154 (47%), Gaps = 10/154 (6%)
Query: 6 ALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDN 65
AL L L+ ++ + AV F VGD GWT +Y +W + K FR+GDTL F Y G +
Sbjct: 8 ALLLLLLLVAVPAVFAVT--FQVGDNDGWTIGVEYTSWVSEKTFRVGDTLEFKY-GPSHS 64
Query: 66 VVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV---- 121
V VN +D+ C + G KI +T G ++ HC G KL + V
Sbjct: 65 VAVVNKADYDGCETSRPTQSFSDGDTKIDLTKVGAIHFLCLTPGHCSLGMKLAVQVLAAV 124
Query: 122 --QPKQDGWSPVPSPS-PSPSLDLVTPEAPPSNA 152
+P +P PSPS PSPS +P P NA
Sbjct: 125 SLEPPPSPSAPSPSPSAPSPSPSAPSPSPSPGNA 158
>At3g60280 uclacyanin 3
Length = 222
Score = 87.8 bits (216), Expect = 3e-18
Identities = 61/173 (35%), Positives = 81/173 (46%), Gaps = 16/173 (9%)
Query: 3 LSRALFLF-ALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVG 61
++ AL LF A + ++F A F VGD GWT+ DY W K FR+GDTL F Y G
Sbjct: 5 VAAALLLFLAAVPAVF-----AATFKVGDISGWTSNLDYTVWLTGKTFRVGDTLEFVY-G 58
Query: 62 GKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV 121
+V V+ + + +C G KI +TT G ++ HC+NG KL + V
Sbjct: 59 LSHSVSVVDKAGYDNCDSSGATQNFADGDTKIDLTTVGTMHFLCPTFGHCKNGMKLAVPV 118
Query: 122 QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPAS-----SVPRRSLLPKKL 169
+P PS SP TP +PPS P+S S P SL P L
Sbjct: 119 LAA----APSPSTPSSPPSTPSTPSSPPSTPSTPSSPPSPPSPPSPSLPPSSL 167
>At2g26720 putative phytocyanin, predicted GPI-anchored protein
Length = 206
Score = 85.1 bits (209), Expect = 2e-17
Identities = 54/144 (37%), Positives = 71/144 (48%), Gaps = 14/144 (9%)
Query: 28 VGDEKGWTTLF-DYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVL 86
VG+ KGWT + DY+ W +++VF++GDTL F Y +V V +DF+ C
Sbjct: 31 VGNTKGWTMIGGDYEAWASSRVFQVGDTLVFAYNKDYHDVTEVTHNDFEMCESSKPLRRY 90
Query: 87 TSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVP------------SP 134
+G D I +T G + +I V HC+ GQKL I V P G VP SP
Sbjct: 91 KTGSDSISLTKPGLQHFICGVPGHCKKGQKLQIHVLPASLGHVAVPVPGPVRSQSSSSSP 150
Query: 135 SPSPSLDLVTPEAPP-SNAPWPAS 157
SPSP +D AP P PAS
Sbjct: 151 SPSPLVDPPVNNAPQYQMGPTPAS 174
>At5g26330 copper binding protein - like, predicted GPI-anchored
protein
Length = 187
Score = 84.7 bits (208), Expect = 2e-17
Identities = 54/165 (32%), Positives = 79/165 (47%), Gaps = 13/165 (7%)
Query: 6 ALFLFALIASIFSTMAVAKDFV--VGDEKGWTTL--FDYQTWTANKVFRLGDTLTFNYVG 61
A + A +A I + +++ V VGD GWTT+ DY+ W + K F +GDT+ F Y
Sbjct: 2 AAIIVAALACIVVMLRLSEAAVYKVGDSAGWTTIANVDYKLWASTKTFHIGDTVLFEYNP 61
Query: 62 GKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV 121
NV+RV ++SC+ T+G D I +T +G ++ V HC GQKL + V
Sbjct: 62 QFHNVMRVTHPMYRSCNTSKPISTFTTGNDSITLTNHGHHFFFCGVPGHCLAGQKLDLHV 121
Query: 122 ------QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVP 160
P D P S S SP + P +P A+S+P
Sbjct: 122 LLPASSTPLSD---PPTSSSSSPPSTTIPAAGVPGPSPSLAASLP 163
>At5g20230 blue copper binding protein (bcb)
Length = 196
Score = 83.6 bits (205), Expect = 5e-17
Identities = 49/140 (35%), Positives = 73/140 (52%), Gaps = 10/140 (7%)
Query: 16 IFSTMAV-AKDFVVGDEKGWTTLFD---YQTWTANKVFRLGDTLTFNYVGGKDNVVRVNG 71
+F+ + V A+D+ VGD+ WT D Y TW K FR+GD L F++ G+ +V V+
Sbjct: 14 VFAAVVVFAEDYDVGDDTEWTRPMDPEFYTTWATGKTFRVGDELEFDFAAGRHDVAVVSE 73
Query: 72 SDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQP------KQ 125
+ F++C +T KI++ T G +++I +V DHC GQKL ITV
Sbjct: 74 AAFENCEKEKPISHMTVPPVKIMLNTTGPQYFICTVGDHCRFGQKLSITVVAAGATGGAT 133
Query: 126 DGWSPVPSPSPSPSLDLVTP 145
G P+P +PS TP
Sbjct: 134 PGAGATPAPGSTPSTGGTTP 153
>At4g31840 predicted GPI-anchored protein
Length = 177
Score = 77.0 bits (188), Expect = 5e-15
Identities = 44/142 (30%), Positives = 70/142 (48%), Gaps = 7/142 (4%)
Query: 1 MALSRALFLFALIASIFSTMAV-AKDFVVGDEKG-W----TTLFDYQTWTANKVFRLGDT 54
MA S L L S+F +V A + VG + G W ++ F + W F++GD
Sbjct: 1 MASSSLLVTIFLCISVFFFSSVNANEVTVGGKSGDWKIPPSSSFSFNEWAQKARFKVGDF 60
Query: 55 LTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENG 114
+ F Y GKD+V++V ++ C+ T G K+ + G +++S HC+ G
Sbjct: 61 IVFKYEAGKDSVLQVTREAYEKCNTTSPKASYTDGNTKVKLDQAGPVYFVSGTEGHCQKG 120
Query: 115 QKL-FITVQPKQDGWSPVPSPS 135
QKL + + P+ +SP PSPS
Sbjct: 121 QKLRLVVITPRNSAFSPGPSPS 142
>At2g25060 early nodulin-like predicted GPI-anchored protein
Length = 176
Score = 77.0 bits (188), Expect = 5e-15
Identities = 51/167 (30%), Positives = 77/167 (45%), Gaps = 22/167 (13%)
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKG-W----TTLFDYQTWTANKVFRLGDTL 55
+A+ +FLF+L A A + VG + G W ++ + + W F++GD +
Sbjct: 8 VAIFSLIFLFSL--------AAANEVTVGGKSGDWKIPPSSSYSFTEWAQKARFKVGDFI 59
Query: 56 TFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQ 115
F Y GKD+V+ V + SC+ T G+ K+ + G ++IS HCE GQ
Sbjct: 60 VFRYESGKDSVLEVTKEAYNSCNTTNPLANYTDGETKVKLDRSGPFYFISGANGHCEKGQ 119
Query: 116 KL-FITVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPR 161
KL + + P+ SP PSP V E P+ AP P S R
Sbjct: 120 KLSLVVISPRHSVISPAPSP--------VEFEDGPALAPAPISGSVR 158
>At4g28365 unknown protein
Length = 199
Score = 76.3 bits (186), Expect = 8e-15
Identities = 47/166 (28%), Positives = 78/166 (46%), Gaps = 16/166 (9%)
Query: 9 LFALIASIFSTMAVAKDFVVGDEKGW--TTLFDYQTWTANKVFRLGDTLTFNYVGGKDNV 66
+F ++ + T++ F VG + GW T DY W+ F++ DTL F Y GKD+V
Sbjct: 12 MFVMLMGLGFTISNGYKFYVGGKDGWVPTPSEDYSHWSHRNRFQVNDTLHFKYAKGKDSV 71
Query: 67 VRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV----- 121
+ V ++ +C+ L+ G +++ G ++IS + +C GQKL + V
Sbjct: 72 LEVTEQEYNTCNTTHPLTSLSDGDSLFLLSHSGSYFFISGNSQNCLKGQKLAVKVLSTVH 131
Query: 122 ---QPKQDGWSPVP------SPSPSPSLDLVTPEAPPSNAPWPASS 158
P+ SP P SP PSP ++ + AP PA++
Sbjct: 132 HSHSPRHTSPSPSPVHQELSSPGPSPGVEPSSDSNSRVPAPGPATA 177
>At4g27520 early nodulin-like 2 predicted GPI-anchored protein
Length = 349
Score = 76.3 bits (186), Expect = 8e-15
Identities = 51/162 (31%), Positives = 83/162 (50%), Gaps = 14/162 (8%)
Query: 8 FLFALIASIFS--TMAVAKDFVVGDEKGWTTL--FDYQTWTANKVFRLGDTLTFNYVGGK 63
F F ++ S+ + T++ A+ F VG W T +Y++W+ F + DTL F+Y G
Sbjct: 11 FFFTILLSLSTLFTISNARKFNVGGSGAWVTNPPENYESWSGKNRFLVHDTLYFSYAKGA 70
Query: 64 DNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQP 123
D+V+ VN +D+ +C+ + G +I + YG ++IS D+C+ GQKL + V
Sbjct: 71 DSVLEVNKADYDACNTKNPIKRVDDGDSEISLDRYGPFYFISGNEDNCKKGQKLNVVVIS 130
Query: 124 KQDGWSPVPSPSPSP---SLDLVTPEA--PPSNAPWPASSVP 160
+ +PS + SP + TP + PP A P SS P
Sbjct: 131 AR-----IPSTAQSPHAAAPGSSTPGSMTPPGGAHSPKSSSP 167
Score = 26.9 bits (58), Expect = 5.7
Identities = 16/35 (45%), Positives = 19/35 (53%), Gaps = 1/35 (2%)
Query: 129 SPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRS 163
SPV SPS +P P AP S++ P SS P S
Sbjct: 225 SPV-SPSSAPMTSPPAPMAPKSSSTIPPSSAPMTS 258
>At2g31050 phytocyanin - like predicted GPI-anchored protein
Length = 200
Score = 76.3 bits (186), Expect = 8e-15
Identities = 49/154 (31%), Positives = 73/154 (46%), Gaps = 4/154 (2%)
Query: 4 SRALFLFALI-ASIFSTMAVAKDFVVGDEKGWTTL-FDYQTWTANKVFRLGDTLTFNYVG 61
S A F LI ++F VGD GWT + +Y+TW + F++GD+L F Y
Sbjct: 6 SNAFFTSLLILVALFGISVGGTVHKVGDSDGWTIMSVNYETWASTITFQVGDSLVFKYNK 65
Query: 62 GKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV 121
+V V +D++ C +G D +I+T G + +I HC+ GQKL I V
Sbjct: 66 DFHDVTEVTHNDYEMCEPSKPLARYETGSDIVILTKPGLQHFICGFPGHCDMGQKLQIHV 125
Query: 122 QPKQDG--WSPVPSPSPSPSLDLVTPEAPPSNAP 153
P G +PVP P PS ++P + +P
Sbjct: 126 LPASLGPVAAPVPGPVRPPSSFSSPSQSPLAESP 159
>At3g18590 predicted GPI-anchored protein
Length = 188
Score = 75.1 bits (183), Expect = 2e-14
Identities = 55/159 (34%), Positives = 83/159 (51%), Gaps = 18/159 (11%)
Query: 7 LFLFALIASIFSTMAVAKDFVVGDEKGWT-----TLFD-YQTWTANKVFRLGDTLTFNYV 60
+ +F + +FS ++ + +F VG E GW TL D + W ++ F++GDTL F Y
Sbjct: 9 IVMFLVTFYMFSCVS-STEFEVGGENGWIVPKSKTLGDAFNQWASDNRFKVGDTLRFKYT 67
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKII--ITTYGRRWYISSVTDHCENGQKLF 118
KD+V+ V+ ++K C T P L S + + + G ++IS V+ HCE GQK+
Sbjct: 68 --KDSVLVVSEEEYKKCKA--TKPQLYSNNEDTVFKLDRPGLFYFISGVSGHCEKGQKMI 123
Query: 119 ITVQPKQDGW-SPVP----SPSPSPSLDLVTPEAPPSNA 152
+ V + SP P S S S SL TP+A SNA
Sbjct: 124 VKVMETESSTESPPPSSSSSSSSSSSLPASTPKAKKSNA 162
>At2g44790 phytocyanin, blue copper-binding protein II
Length = 202
Score = 72.8 bits (177), Expect = 9e-14
Identities = 50/163 (30%), Positives = 70/163 (42%), Gaps = 10/163 (6%)
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
MA++ A L ++ + +AV WTT DY W K FR+GD L F Y
Sbjct: 9 MAVAAATALLLVLTIVPGAVAVTYTIE------WTTGVDYSGWATGKTFRVGDILEFKY- 61
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHC--ENGQKLF 118
G V V+ + + C + + G KI + T G ++I S HC G KL
Sbjct: 62 GSSHTVDVVDKAGYDGCDASSSTENHSDGDTKIDLKTVGINYFICSTPGHCRTNGGMKLA 121
Query: 119 ITVQPKQDGWSPVPSPSPSPSLDLVTPEAPPS-NAPWPASSVP 160
+ V G P+P S TPE+PPS +P P + P
Sbjct: 122 VNVVAGSAGPPATPTPPSSTPGTPTTPESPPSGGSPTPTTPTP 164
>At4g32490 nodulin - like predicted GPI-anchored protein
Length = 221
Score = 71.6 bits (174), Expect = 2e-13
Identities = 47/134 (35%), Positives = 68/134 (50%), Gaps = 5/134 (3%)
Query: 26 FVVGDEKGW--TTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTA 83
F VG GW T DY W+ F++ DTL F YV GKD+V+ V+ ++ +C+
Sbjct: 31 FYVGGRDGWVLTPSEDYSHWSHRNRFQVNDTLYFKYVKGKDSVLEVSEKEYNTCNTTHPL 90
Query: 84 PVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPK-QDGWSP-VPSPSPSPSLD 141
L+ G +++ +++S + C GQKL +TV +P PSPSPSPS
Sbjct: 91 TSLSDGDSLFLLSRSDPFFFVSGNSGSCLKGQKLAVTVMSTGHHSHTPRHPSPSPSPSAS 150
Query: 142 LVTPEAPPSNAPWP 155
V +A S AP P
Sbjct: 151 PVR-KALLSPAPIP 163
>At3g20570 predicted GPI-anchored protein
Length = 203
Score = 71.6 bits (174), Expect = 2e-13
Identities = 45/151 (29%), Positives = 70/151 (45%), Gaps = 14/151 (9%)
Query: 21 AVAKDFVVGDEKGWTTLFD---YQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSC 77
A A++F VG GWT Y W F++GD+L F Y +D+V++V + SC
Sbjct: 25 AYAREFTVGGATGWTVPSGSQVYSQWAEQSRFQIGDSLLFVYQSNQDSVLQVTRDAYDSC 84
Query: 78 SVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDG----WSPVPS 133
+ G+ + + G ++IS D+C+ +KL + V + G S PS
Sbjct: 85 NTDSPTAKFADGKTSVTLNHSGPYYFISGNKDNCKKNEKLVVIVMADRSGNKNTASSPPS 144
Query: 134 PSPSPSLDLVTPEAPPSN------APWPASS 158
P+P+PS + P P S AP P +S
Sbjct: 145 PAPAPSGE-SAPSPPVSGTFEMTPAPTPTTS 174
>At1g22480 blue copper protein precursor-like predicted GPI-anchored
protein
Length = 174
Score = 70.5 bits (171), Expect = 4e-13
Identities = 52/163 (31%), Positives = 75/163 (45%), Gaps = 11/163 (6%)
Query: 9 LFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVR 68
L + IFS +A A + W+ DY T K F +GDT+ FNY G V
Sbjct: 4 LLGCLVLIFSMVAQASSASL--TVNWSLGTDYTPLTTGKTFSVGDTIVFNY-GAGHTVDE 60
Query: 69 VNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDG- 127
V+ +D+KSC++ + +SG I +TT G R++I + HC G KL +TV
Sbjct: 61 VSENDYKSCTLGNSITSDSSGTTTIALTTTGPRYFICGIPGHCAAGMKLAVTVASNSSNG 120
Query: 128 -----WSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRSLL 165
+P P + T +A P A W A S P R+L+
Sbjct: 121 VAGGTTTPTPFTGGGGGYNPTTTQAIPC-AAW-AVSCPLRALV 161
>At2g02850 putative basic blue protein (plantacyanin)
Length = 129
Score = 70.1 bits (170), Expect = 6e-13
Identities = 36/98 (36%), Positives = 54/98 (54%), Gaps = 3/98 (3%)
Query: 23 AKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLT 82
A + VGD WT F+ W K FR GD L FNY NVV+V+ + +C P
Sbjct: 33 AATYTVGDSGIWT--FNAVGWPKGKHFRAGDVLVFNYNPRMHNVVKVDSGSYNNCKTPTG 90
Query: 83 APVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
A TSG+D+I ++ G+ ++I + +HCE+ K+ +T
Sbjct: 91 AKPYTSGKDRITLSK-GQNFFICNFPNHCESDMKIAVT 127
>At5g07475 putative protein
Length = 192
Score = 68.9 bits (167), Expect = 1e-12
Identities = 47/163 (28%), Positives = 76/163 (45%), Gaps = 8/163 (4%)
Query: 4 SRALFLFALIASIFSTMAV----AKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNY 59
S+ FLF L IF + + A + VGD GW D ++WT+ K F GD L F Y
Sbjct: 5 SKIQFLFNLCI-IFGVVVIRRCNATTYFVGDSSGWDISSDLESWTSGKRFSPGDVLMFQY 63
Query: 60 VGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFI 119
+V V ++++C+ T+G + ++ G R+++ HC G +L +
Sbjct: 64 -SSTHSVYEVAKDNYQNCNTTDAIRTFTNGNTTVALSKPGNRFFVCGNRLHCFAGMRLLV 122
Query: 120 TVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSN-APWPASSVPR 161
V+ +PV SP + S ++ P + +N A ASS R
Sbjct: 123 NVEGNGPSQAPVGSPQAATS-GILQPSSKKNNPATGVASSAAR 164
>At3g27200 blue copper protein, putative
Length = 174
Score = 68.6 bits (166), Expect = 2e-12
Identities = 36/139 (25%), Positives = 65/139 (45%)
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
M + L + + A+A V+G +GW D+ +W++++ F++GD + F Y
Sbjct: 1 MKMQAVLVILVFSGLLSVKTALAARHVIGGSQGWEQSVDFDSWSSDQSFKVGDQIVFKYS 60
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
V + + +KSC + + L+SG D + ++ G R++ HCE G K+ +
Sbjct: 61 ELHSVVELGSETAYKSCDLGTSVNSLSSGNDVVKLSKTGTRYFACGTVGHCEQGMKIKVN 120
Query: 121 VQPKQDGWSPVPSPSPSPS 139
V + PS S S S
Sbjct: 121 VVSSDSKSASSPSGSGSGS 139
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,274,556
Number of Sequences: 26719
Number of extensions: 209781
Number of successful extensions: 1921
Number of sequences better than 10.0: 142
Number of HSP's better than 10.0 without gapping: 98
Number of HSP's successfully gapped in prelim test: 46
Number of HSP's that attempted gapping in prelim test: 1277
Number of HSP's gapped (non-prelim): 565
length of query: 176
length of database: 11,318,596
effective HSP length: 93
effective length of query: 83
effective length of database: 8,833,729
effective search space: 733199507
effective search space used: 733199507
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC126009.11


