

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126008.6 + phase: 0
(530 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
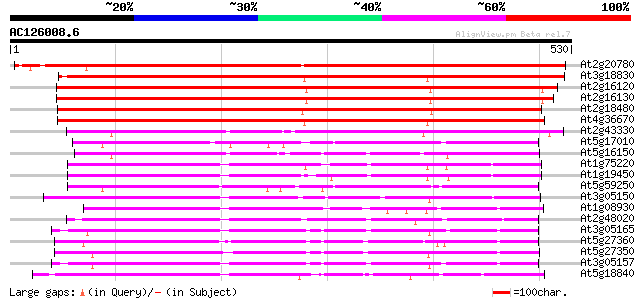
Score E
Sequences producing significant alignments: (bits) Value
At2g20780 putative sugar transporter 705 0.0
At3g18830 sugar transporter like protein 457 e-129
At2g16120 putative sugar transporter 448 e-126
At2g16130 putative sugar transporter 446 e-125
At2g18480 putative sugar transporter 443 e-124
At4g36670 sugar transporter like protein 436 e-122
At2g43330 membrane transporter like protein 263 2e-70
At5g17010 sugar transporter - like protein 243 2e-64
At5g16150 sugar transporter like protein 240 2e-63
At1g75220 integral membrane protein, putative 236 3e-62
At1g19450 integral membrane protein, putative 230 2e-60
At5g59250 D-xylose-H+ symporter - like protein 227 1e-59
At3g05150 putative sugar transporter 223 2e-58
At1g08930 ERD6 protein 222 3e-58
At2g48020 sugar transporter like protein 217 1e-56
At3g05165 sugar transporter like protein 213 2e-55
At5g27360 sugar-porter family protein 2 (SFP2) 213 2e-55
At5g27350 sugar transporter like protein 213 2e-55
At3g05157 sugar transporter like protein 212 4e-55
At5g18840 sugar transporter - like protein 211 6e-55
>At2g20780 putative sugar transporter
Length = 547
Score = 705 bits (1820), Expect = 0.0
Identities = 371/546 (67%), Positives = 433/546 (78%), Gaps = 33/546 (6%)
Query: 5 ENGNGEMGLSGIPL----GTKNKYKRMNSHLVDDNDDVLHQQQLEDKRNSTRKYVIACAI 60
E GNG G SG P KNKY+RM+S D ++ + ++ E + + TRKYV+ACA
Sbjct: 7 EVGNG--GGSGFPAVSVGNKKNKYQRMDS----DAEESQNHREAEARNSRTRKYVMACAF 60
Query: 61 FASLNNVLLGY---------------------DVGVMSGAVIFIKEDLKITEVQVEFLIG 99
FASLNNVLLGY DVGVMSGAV+FI++DLKITEVQ E LIG
Sbjct: 61 FASLNNVLLGYGRFYLYNRILLLLLYFVDLQKDVGVMSGAVLFIQQDLKITEVQTEVLIG 120
Query: 100 ILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGI 159
LSI+SL GSL GGRTSD IGRKWTMALAA+VFQ G M +APS++VLMIGR LAGIGI
Sbjct: 121 SLSIISLFGSLAGGRTSDSIGRKWTMALAALVFQTGAAVMAVAPSFEVLMIGRTLAGIGI 180
Query: 160 GFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAV 219
G GVMI+P+YIAEISP + RG T+FPEIFIN+GI+LGYVSNYAFSGLSVHISWR+MLAV
Sbjct: 181 GLGVMIAPVYIAEISPTVARGFFTSFPEIFINLGILLGYVSNYAFSGLSVHISWRIMLAV 240
Query: 220 GILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFAN 279
GILPSVFIGFAL +IPESPRWLVM+ R++ AR VL+KTNE + E EERLAEIQ AA A+
Sbjct: 241 GILPSVFIGFALCVIPESPRWLVMKGRVDSAREVLMKTNERDDEAEERLAEIQLAA--AH 298
Query: 280 SGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSK 339
+ ED+PVWRELLSP P +R+MLI G GIQCFQQI+GIDATVYYSPEIL AGI+D++K
Sbjct: 299 TEGSEDRPVWRELLSPSPVVRKMLIVGFGIQCFQQITGIDATVYYSPEILKEAGIQDETK 358
Query: 340 LLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLVI 399
LLAATVAVG+TKTVFIL A LID VGRKPLL STIGMT CLFC+ TL+ +G L I
Sbjct: 359 LLAATVAVGVTKTVFILFATFLIDSVGRKPLLYVSTIGMTLCLFCLSFTLTFLGQGTLGI 418
Query: 400 ALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSV 459
L +LFVCGNVAFFS+G+GPVCWVLTSEIFPLR+RAQASALGAV NRVCSGLVAMSFLSV
Sbjct: 419 TLALLFVCGNVAFFSIGMGPVCWVLTSEIFPLRLRAQASALGAVGNRVCSGLVAMSFLSV 478
Query: 460 SDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQGKEMELGDVE 519
S AI+ GGTFF+FS +SAL+++FV+ LVPET GKSLEQIE+MF+ + E+ELGD E
Sbjct: 479 SRAITVGGTFFVFSLVSALSVIFVYVLVPETSGKSLEQIELMFQGGLERKDGEVELGDAE 538
Query: 520 QLVQNK 525
+LV+ +
Sbjct: 539 RLVRKE 544
>At3g18830 sugar transporter like protein
Length = 539
Score = 457 bits (1175), Expect = e-129
Identities = 237/493 (48%), Positives = 329/493 (66%), Gaps = 18/493 (3%)
Query: 47 KRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSL 106
KRN+ Y ACAI AS+ ++LLGYD+GVMSGA+I+IK DLKI ++Q+ L G L+I SL
Sbjct: 31 KRNN---YAFACAILASMTSILLGYDIGVMSGAMIYIKRDLKINDLQIGILAGSLNIYSL 87
Query: 107 LGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMIS 166
+GS GRTSD IGR++T+ LA +F G I M L+P+Y LM GR +AGIG+G+ +MI+
Sbjct: 88 IGSCAAGRTSDWIGRRYTIVLAGAIFFAGAILMGLSPNYAFLMFGRFIAGIGVGYALMIA 147
Query: 167 PIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVF 226
P+Y AE+SP +RG L +FPE+FIN GIMLGYVSN AFS L + + WR+ML +G +PSV
Sbjct: 148 PVYTAEVSPASSRGFLNSFPEVFINAGIMLGYVSNLAFSNLPLKVGWRLMLGIGAVPSVI 207
Query: 227 IGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGF--------- 277
+ + +PESPRWLVMQ R+ +A+ VL KT++ E RL +I+ AAG
Sbjct: 208 LAIGVLAMPESPRWLVMQGRLGDAKRVLDKTSDSPTEATLRLEDIKHAAGIPADCHDDVV 267
Query: 278 -ANSGKYEDKPVWRELL-SPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIE 335
+ + VWRELL P PA+RR++I +GI FQQ SGIDA V +SP I AG++
Sbjct: 268 QVSRRNSHGEGVWRELLIRPTPAVRRVMIAAIGIHFFQQASGIDAVVLFSPRIFKTAGLK 327
Query: 336 DKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE-- 393
+ L ATVAVG+ KT FILVA L+D++GR+PLL+TS GM L +G +L++ +
Sbjct: 328 TDHQQLLATVAVGVVKTSFILVATFLLDRIGRRPLLLTSVGGMVLSLAALGTSLTIIDQS 387
Query: 394 --KGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGL 451
K + + I V VA FS+G GP+ WV +SEIFPLR+R+Q S++G V NRV SG+
Sbjct: 388 EKKVMWAVVVAIATVMTYVATFSIGAGPITWVYSSEIFPLRLRSQGSSMGVVVNRVTSGV 447
Query: 452 VAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQGK 511
+++SFL +S A++ GG F+LF I+ +A VF +T +PET+G+ LE ++ +F K
Sbjct: 448 ISISFLPMSKAMTTGGAFYLFGGIATVAWVFFYTFLPETQGRMLEDMDELFSGFRWRDSK 507
Query: 512 EMELGDVEQLVQN 524
G+ E+ V N
Sbjct: 508 SKPKGNPEKTVPN 520
>At2g16120 putative sugar transporter
Length = 511
Score = 448 bits (1152), Expect = e-126
Identities = 233/494 (47%), Positives = 331/494 (66%), Gaps = 21/494 (4%)
Query: 45 EDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIV 104
E R + +Y ACAI AS+ +++LGYD+GVMSGA IFIK+DLK+++VQ+E L+GIL+I
Sbjct: 16 EPPRGNRSRYAFACAILASMTSIILGYDIGVMSGASIFIKDDLKLSDVQLEILMGILNIY 75
Query: 105 SLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVM 164
SL+GS GRTSD +GR++T+ LA F G + M A +Y +M+GR +AGIG+G+ +M
Sbjct: 76 SLVGSGAAGRTSDWLGRRYTIVLAGAFFFCGALLMGFATNYPFIMVGRFVAGIGVGYAMM 135
Query: 165 ISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPS 224
I+P+Y AE++P +RG LT+FPEIFIN+GI+LGYVSNY FS L H+ WR ML VG +PS
Sbjct: 136 IAPVYTAEVAPASSRGFLTSFPEIFINIGILLGYVSNYFFSKLPEHLGWRFMLGVGAVPS 195
Query: 225 VFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFAN----- 279
VF+ + +PESPRWLV+Q R+ +A VL KT+ ++E RL +I++A G +
Sbjct: 196 VFLAIGVLAMPESPRWLVLQGRLGDAFKVLDKTSNTKEEAISRLDDIKRAVGIPDDMTDD 255
Query: 280 -----SGKYEDKPVWRELL-SPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAG 333
+ K K VW++LL P P++R +LI LGI QQ SGIDA V YSP I AG
Sbjct: 256 VIVVPNKKSAGKGVWKDLLVRPTPSVRHILIACLGIHFAQQASGIDAVVLYSPTIFSKAG 315
Query: 334 IEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE 393
++ K+ L ATVAVG+ KT+FI+V ++D+ GR+ LL+TS GM L +G +L++
Sbjct: 316 LKSKNDQLLATVAVGVVKTLFIVVGTCVVDRFGRRALLLTSMGGMFLSLTALGTSLTVIN 375
Query: 394 KGP-----LVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVC 448
+ P I L + V VA FS+G GPV WV SEIFP+R+RAQ ++LG + NR+
Sbjct: 376 RNPGQTLKWAIGLAVTTVMTFVATFSIGAGPVTWVYCSEIFPVRLRAQGASLGVMLNRLM 435
Query: 449 SGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF-----E 503
SG++ M+FLS+S ++ GG F LF+ ++A A VF FT +PET+G LE++E +F
Sbjct: 436 SGIIGMTFLSLSKGLTIGGAFLLFAGVAAAAWVFFFTFLPETRGIPLEEMETLFGSYTAN 495
Query: 504 NEHGSQGKEMELGD 517
++ S K+ E+ D
Sbjct: 496 KKNNSMSKDNEVVD 509
>At2g16130 putative sugar transporter
Length = 511
Score = 446 bits (1146), Expect = e-125
Identities = 232/492 (47%), Positives = 330/492 (66%), Gaps = 23/492 (4%)
Query: 45 EDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIV 104
E R + ++ ACAI AS+ +++LGYD+GVMSGA IFIK+DLK+++VQ+E L+GIL+I
Sbjct: 16 EPPRGNRSRFAFACAILASMTSIILGYDIGVMSGAAIFIKDDLKLSDVQLEILMGILNIY 75
Query: 105 SLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVM 164
SL+GS GRTSD IGR++T+ LA F G + M A +Y +M+GR +AGIG+G+ +M
Sbjct: 76 SLIGSGAAGRTSDWIGRRYTIVLAGFFFFCGALLMGFATNYPFIMVGRFVAGIGVGYAMM 135
Query: 165 ISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPS 224
I+P+Y E++P +RG L++FPEIFIN+GI+LGYVSNY F+ L HI WR ML +G +PS
Sbjct: 136 IAPVYTTEVAPASSRGFLSSFPEIFINIGILLGYVSNYFFAKLPEHIGWRFMLGIGAVPS 195
Query: 225 VFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFAN----- 279
VF+ + +PESPRWLVMQ R+ +A VL KT+ ++E RL +I++A G +
Sbjct: 196 VFLAIGVLAMPESPRWLVMQGRLGDAFKVLDKTSNTKEEAISRLNDIKRAVGIPDDMTDD 255
Query: 280 -----SGKYEDKPVWRELL-SPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAG 333
+ K K VW++LL P P++R +LI LGI QQ SGIDA V YSP I AG
Sbjct: 256 VIVVPNKKSAGKGVWKDLLVRPTPSVRHILIACLGIHFSQQASGIDAVVLYSPTIFSRAG 315
Query: 334 IEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE 393
++ K+ L ATVAVG+ KT+FI+V L+D+ GR+ LL+TS GM L +G +L++ +
Sbjct: 316 LKSKNDQLLATVAVGVVKTLFIVVGTCLVDRFGRRALLLTSMGGMFFSLTALGTSLTVID 375
Query: 394 KGP-----LVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVC 448
+ P I L + V VA FS+G GPV WV SEIFP+R+RAQ ++LG + NR+
Sbjct: 376 RNPGQTLKWAIGLAVTTVMTFVATFSLGAGPVTWVYASEIFPVRLRAQGASLGVMLNRLM 435
Query: 449 SGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF------ 502
SG++ M+FLS+S ++ GG F LF+ ++ A VF FT +PET+G LE+IE +F
Sbjct: 436 SGIIGMTFLSLSKGLTIGGAFLLFAGVAVAAWVFFFTFLPETRGVPLEEIESLFGSYSAN 495
Query: 503 -ENEHGSQGKEM 513
+N S+GK++
Sbjct: 496 KKNNVMSKGKQV 507
>At2g18480 putative sugar transporter
Length = 508
Score = 443 bits (1139), Expect = e-124
Identities = 226/471 (47%), Positives = 322/471 (67%), Gaps = 14/471 (2%)
Query: 46 DKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVS 105
D K+ CAI AS+ +++ GYD GVMSGA IFI++DLKI + Q+E L GIL++ +
Sbjct: 13 DPNPHMNKFAFGCAIVASIISIIFGYDTGVMSGAQIFIRDDLKINDTQIEVLAGILNLCA 72
Query: 106 LLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMI 165
L+GSL G+TSD+IGR++T+AL+AV+F +G + M P+Y VLM+GR +AG+G+GF +MI
Sbjct: 73 LVGSLTAGKTSDVIGRRYTIALSAVIFLVGSVLMGYGPNYPVLMVGRCIAGVGVGFALMI 132
Query: 166 SPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSV 225
+P+Y AEIS RG LT+ PE+ I++GI+LGYVSNY F L++ + WR+ML + PS+
Sbjct: 133 APVYSAEISSASHRGFLTSLPELCISLGILLGYVSNYCFGKLTLKLGWRLMLGIAAFPSL 192
Query: 226 FIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAA---------- 275
+ F + +PESPRWLVMQ R+EEA+ +++ + E+E EER +I AA
Sbjct: 193 ILAFGITRMPESPRWLVMQGRLEEAKKIMVLVSNTEEEAEERFRDILTAAEVDVTEIKEV 252
Query: 276 GFANSGKYEDKPVWREL-LSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGI 334
G K K VWREL + P PA+R +LI +GI F+ +GI+A V YSP I AG+
Sbjct: 253 GGGVKKKNHGKSVWRELVIKPRPAVRLILIAAVGIHFFEHATGIEAVVLYSPRIFKKAGV 312
Query: 335 EDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEK 394
K KLL ATV VG+TK FI++A L+DKVGR+ LL+TST GM L + V+L++ ++
Sbjct: 313 VSKDKLLLATVGVGLTKAFFIIIATFLLDKVGRRKLLLTSTGGMVFALTSLAVSLTMVQR 372
Query: 395 -GPL--VIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGL 451
G L ++L I+ VAFFS+GLGP+ WV +SEIFPLR+RAQ +++G NR+ +
Sbjct: 373 FGRLAWALSLSIVSTYAFVAFFSIGLGPITWVYSSEIFPLRLRAQGASIGVAVNRIMNAT 432
Query: 452 VAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
V+MSFLS++ AI+ GG FF+F+ I+ A F F ++PETKG LE++E +F
Sbjct: 433 VSMSFLSMTKAITTGGVFFVFAGIAVAAWWFFFFMLPETKGLPLEEMEKLF 483
>At4g36670 sugar transporter like protein
Length = 493
Score = 436 bits (1120), Expect = e-122
Identities = 220/475 (46%), Positives = 317/475 (66%), Gaps = 15/475 (3%)
Query: 46 DKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVS 105
+K ++ + CAI AS+ +++ GYD GVMSGA++FI+EDLK +VQ+E L GIL++ +
Sbjct: 8 EKPAGVNRFALQCAIVASIVSIIFGYDTGVMSGAMVFIEEDLKTNDVQIEVLTGILNLCA 67
Query: 106 LLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMI 165
L+GSL GRTSDIIGR++T+ LA+++F +G I M P+Y VL+ GR AG+G+GF +M+
Sbjct: 68 LVGSLLAGRTSDIIGRRYTIVLASILFMLGSILMGWGPNYPVLLSGRCTAGLGVGFALMV 127
Query: 166 SPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSV 225
+P+Y AEI+ RG L + P + I++GI+LGY+ NY FS L +HI WR+ML + +PS+
Sbjct: 128 APVYSAEIATASHRGLLASLPHLCISIGILLGYIVNYFFSKLPMHIGWRLMLGIAAVPSL 187
Query: 226 FIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGF-------- 277
+ F + +PESPRWL+MQ R++E + +L + +E E R +I+ AAG
Sbjct: 188 VLAFGILKMPESPRWLIMQGRLKEGKEILELVSNSPEEAELRFQDIKAAAGIDPKCVDDV 247
Query: 278 --ANSGKYEDKPVWREL-LSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGI 334
K + VW+EL L P PA+RR+L+T LGI FQ SGI+A + Y P I AGI
Sbjct: 248 VKMEGKKTHGEGVWKELILRPTPAVRRVLLTALGIHFFQHASGIEAVLLYGPRIFKKAGI 307
Query: 335 EDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE- 393
K KL T+ VGI KT FI A +L+DKVGR+ LL+TS GM L +G L++ +
Sbjct: 308 TTKDKLFLVTIGVGIMKTTFIFTATLLLDKVGRRKLLLTSVGGMVIALTMLGFGLTMAQN 367
Query: 394 ---KGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSG 450
K + L I+ VAFFS+GLGP+ WV +SE+FPL++RAQ ++LG NRV +
Sbjct: 368 AGGKLAWALVLSIVAAYSFVAFFSIGLGPITWVYSSEVFPLKLRAQGASLGVAVNRVMNA 427
Query: 451 LVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENE 505
V+MSFLS++ AI+ GG FF+F+ ++A+A F F L+PETKGKSLE+IE +F+ +
Sbjct: 428 TVSMSFLSLTSAITTGGAFFMFAGVAAVAWNFFFFLLPETKGKSLEEIEALFQRD 482
>At2g43330 membrane transporter like protein
Length = 509
Score = 263 bits (672), Expect = 2e-70
Identities = 159/481 (33%), Positives = 253/481 (52%), Gaps = 17/481 (3%)
Query: 54 YVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQV---EFLIGILSIVSLLGSL 110
Y++ + A + +L GYD GV+SGA+++IK+D ++ + E ++ + + +++G+
Sbjct: 30 YILGLTVTAGIGGLLFGYDTGVISGALLYIKDDFEVVKQSSFLQETIVSMALVGAMIGAA 89
Query: 111 GGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYI 170
GG +D GRK A VVF G I M AP VL+ GRLL G+G+G + +P+YI
Sbjct: 90 AGGWINDYYGRKKATLFADVVFAAGAIVMAAAPDPYVLISGRLLVGLGVGVASVTAPVYI 149
Query: 171 AEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFA 230
AE SP+ RG L + + I G L Y+ N AF+ V +WR ML V +P+V
Sbjct: 150 AEASPSEVRGGLVSTNVLMITGGQFLSYLVNSAFT--QVPGTWRWMLGVSGVPAVIQFIL 207
Query: 231 LFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVWR 290
+ +PESPRWL M+NR EA VL +T D +E+ EI + K + V
Sbjct: 208 MLFMPESPRWLFMKNRKAEAIQVLART-YDISRLED---EIDHLSAAEEEEKQRKRTVGY 263
Query: 291 ELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGIT 350
+ LR + G G+Q FQQ +GI+ +YYSP I+ AG L ++ V
Sbjct: 264 LDVFRSKELRLAFLAGAGLQAFQQFTGINTVMYYSPTIVQMAGFHSNQLALFLSLIVAAM 323
Query: 351 KTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTL----SLFEKGPLVIALGILFV 406
+V I ID GRK L ++S G+ L + V+ G L L +L +
Sbjct: 324 NAAGTVVGIYFIDHCGRKKLALSSLFGVIISLLILSVSFFKQSETSSDGGLYGWLAVLGL 383
Query: 407 CGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFG 466
+ FF+ G+GPV W + SEI+P + R + A N + + +VA +FL++++A G
Sbjct: 384 ALYIVFFAPGMGPVPWTVNSEIYPQQYRGICGGMSATVNWISNLIVAQTFLTIAEAAGTG 443
Query: 467 GTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF-ENEHGS---QGKEMELGDVEQLV 522
TF + + I+ LA++FV VPET+G + ++E ++ E +G+ G + ++E L+
Sbjct: 444 MTFLILAGIAVLAVIFVIVFVPETQGLTFSEVEQIWKERAYGNISGWGSSSDSNNMEGLL 503
Query: 523 Q 523
+
Sbjct: 504 E 504
>At5g17010 sugar transporter - like protein
Length = 503
Score = 243 bits (620), Expect = 2e-64
Identities = 148/464 (31%), Positives = 243/464 (51%), Gaps = 40/464 (8%)
Query: 60 IFASLNNVLLGYDVGVMSGAVIFIKED-------LKITEVQVEFLIGILSIVSLLGSLGG 112
+F +L +L GY++G S A I ++ ++ V V + +L GS+
Sbjct: 52 LFPALGGLLYGYEIGATSCATISLQSPSLSGISWYNLSSVDVGLVTSGSLYGALFGSIVA 111
Query: 113 GRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAE 172
+D+IGR+ + LAA+++ +G + LAP+Y VL+IGR++ G+ +G + +P+YIAE
Sbjct: 112 FTIADVIGRRKELILAALLYLVGALVTALAPTYSVLIIGRVIYGVSVGLAMHAAPMYIAE 171
Query: 173 ISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGL--SVHISWRVMLAVGILPSVFIGFA 230
+P+ RG L + E F I+LG V Y L +VH WR M A + +V +G
Sbjct: 172 TAPSPIRGQLVSLKEFF----IVLGMVGGYGIGSLTVNVHSGWRYMYATSVPLAVIMGIG 227
Query: 231 LFIIPESPRWLVM-----QNRIEEARSVLLKT----------NEDEKEVEERLAEIQQAA 275
++ +P SPRWL++ + +E R +K+ + ++V E LAE+
Sbjct: 228 MWWLPASPRWLLLRVIQGKGNVENQREAAIKSLCCLRGPAFVDSAAEQVNEILAELTFVG 287
Query: 276 GFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIE 335
EDK V L L+ ++I G G+ FQQI+G + +YY+P IL AG
Sbjct: 288 --------EDKEVTFGELFQGKCLKALIIGG-GLVLFQQITGQPSVLYYAPSILQTAGFS 338
Query: 336 DKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKG 395
++ +G+ K + VA+V+ID++GR+PLL+ GM LF +G F
Sbjct: 339 AAGDATRVSILLGLLKLIMTGVAVVVIDRLGRRPLLLGGVGGMVVSLFLLGSYYLFFSAS 398
Query: 396 PLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMS 455
P+V + +L G + + GP+ W++ SEIFPL++R + +L + N + LV +
Sbjct: 399 PVVAVVALLLYVG---CYQLSFGPIGWLMISEIFPLKLRGRGLSLAVLVNFGANALVTFA 455
Query: 456 FLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
F + + + G F F I L++VF+F +VPETKG +LE+IE
Sbjct: 456 FSPLKELLGAGILFCGFGVICVLSLVFIFFIVPETKGLTLEEIE 499
>At5g16150 sugar transporter like protein
Length = 546
Score = 240 bits (612), Expect = 2e-63
Identities = 154/444 (34%), Positives = 241/444 (53%), Gaps = 20/444 (4%)
Query: 62 ASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQV--EFLIGILSIVSLLGSLGGGRTSDII 119
A L +L GY +GV++GA+ ++ +DL I E V +++ L + +GS GG +D
Sbjct: 112 ACLGAILFGYHLGVVNGALEYLAKDLGIAENTVLQGWIVSSLLAGATVGSFTGGALADKF 171
Query: 120 GRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAEISPNLTR 179
GR T L A+ +G A S Q +++GRLLAGIGIG I P+YI+EISP R
Sbjct: 172 GRTRTFQLDAIPLAIGAFLCATAQSVQTMIVGRLLAGIGIGISSAIVPLYISEISPTEIR 231
Query: 180 GSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFALFIIPESPR 239
G+L + ++FI +GI+ ++ + + + WR M V ++PSV + + PESPR
Sbjct: 232 GALGSVNQLFICIGILAALIAGLPLA--ANPLWWRTMFGVAVIPSVLLAIGMAFSPESPR 289
Query: 240 WLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVWRELLSPPPAL 299
WLV Q ++ EA +KT +ER+ E+ + + G E + W +L S
Sbjct: 290 WLVQQGKVSEAEKA-IKTLYG----KERVVELVRDLSASGQGSSEPEAGWFDLFS--SRY 342
Query: 300 RRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAI 359
+++ G + FQQ++GI+A VYYS + +AGI+ +AA+ VG + VA
Sbjct: 343 WKVVSVGAALFLFQQLAGINAVVYYSTSVFRSAGIQSD---VAASALVGASNVFGTAVAS 399
Query: 360 VLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLVIALGILFVCGNVAF---FSVG 416
L+DK+GRK LL+TS GM + + ++ F L G L V G V + FS+G
Sbjct: 400 SLMDKMGRKSLLLTSFGGMALSMLLLSLS---FTWKALAAYSGTLAVVGTVLYVLSFSLG 456
Query: 417 LGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAIS 476
GPV +L EIF R+RA+A AL + + + ++ + FLSV + F+ +
Sbjct: 457 AGPVPALLLPEIFASRIRAKAVALSLGMHWISNFVIGLYFLSVVTKFGISSVYLGFAGVC 516
Query: 477 ALAIVFVFTLVPETKGKSLEQIEM 500
LA++++ V ETKG+SLE+IE+
Sbjct: 517 VLAVLYIAGNVVETKGRSLEEIEL 540
>At1g75220 integral membrane protein, putative
Length = 487
Score = 236 bits (601), Expect = 3e-62
Identities = 146/458 (31%), Positives = 243/458 (52%), Gaps = 29/458 (6%)
Query: 55 VIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGSLGGGR 114
V+AC + +L + G+ G S I +DL +T + + ++ +++G++ G+
Sbjct: 48 VLACVLIVALGPIQFGFTCGYSSPTQAAITKDLGLTVSEYSVFGSLSNVGAMVGAIASGQ 107
Query: 115 TSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAEIS 174
++ IGRK ++ +AA+ +G + ++ A L +GRLL G G+G P+YIAEI+
Sbjct: 108 IAEYIGRKGSLMIAAIPNIIGWLCISFAKDTSFLYMGRLLEGFGVGIISYTVPVYIAEIA 167
Query: 175 PNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFALFII 234
P RG L + ++ + +GIML Y+ L + + WR++ +GILP + LF I
Sbjct: 168 PQNMRGGLGSVNQLSVTIGIMLAYL-------LGLFVPWRILAVLGILPCTLLIPGLFFI 220
Query: 235 PESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFA---NSGKYEDKPVWRE 291
PESPRWL +E + L E ++ + EI+++ + N+ ++ D R
Sbjct: 221 PESPRWLAKMGMTDEFETSLQVLRGFETDITVEVNEIKRSVASSTKRNTVRFVDLKRRRY 280
Query: 292 LLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITK 351
L+ G+G+ QQ+ GI+ ++YS I +AG+ + AAT VG +
Sbjct: 281 YFP--------LMVGIGLLVLQQLGGINGVLFYSSTIFESAGVTSSN---AATFGVGAIQ 329
Query: 352 TVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE----KGPLVIALGILFVC 407
V ++ L+DK GR+ LL S++GMT L + L E + L IL V
Sbjct: 330 VVATAISTWLVDKAGRRLLLTISSVGMTISLVIVAAAFYLKEFVSPDSDMYSWLSILSVV 389
Query: 408 GNVA---FFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAIS 464
G VA FFS+G+GP+ W++ SEI P+ ++ A ++ +AN S L+ M+ ++ A S
Sbjct: 390 GVVAMVVFFSLGMGPIPWLIMSEILPVNIKGLAGSIATLANWFFSWLITMT-ANLLLAWS 448
Query: 465 FGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
GGTF L+ + A +VFV VPETKGK+LE+++ +F
Sbjct: 449 SGGTFTLYGLVCAFTVVFVTLWVPETKGKTLEELQSLF 486
>At1g19450 integral membrane protein, putative
Length = 488
Score = 230 bits (586), Expect = 2e-60
Identities = 144/459 (31%), Positives = 245/459 (53%), Gaps = 31/459 (6%)
Query: 55 VIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGSLGGGR 114
V+AC + +L + G+ G S I +DL +T + + ++ +++G++ G+
Sbjct: 49 VLACVLIVALGPIQFGFTCGYSSPTQAAITKDLGLTVSEYSVFGSLSNVGAMVGAIASGQ 108
Query: 115 TSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAEIS 174
++ +GRK ++ +AA+ +G ++++ A L +GRLL G G+G P+YIAEI+
Sbjct: 109 IAEYVGRKGSLMIAAIPNIIGWLSISFAKDTSFLYMGRLLEGFGVGIISYTVPVYIAEIA 168
Query: 175 PNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFALFII 234
P RG+L + ++ + +GIML Y+ L + + WR++ +G+LP + LF I
Sbjct: 169 PQTMRGALGSVNQLSVTIGIMLAYL-------LGLFVPWRILAVLGVLPCTLLIPGLFFI 221
Query: 235 PESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVWRELLS 294
PESPRWL ++ + L E ++ + EI+++ A+S K R +
Sbjct: 222 PESPRWLAKMGLTDDFETSLQVLRGFETDITVEVNEIKRSV--ASSSK-------RSAVR 272
Query: 295 PPPALRRM----LITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGIT 350
RR L+ G+G+ QQ+ GI+ ++YS I +AG+ + AT VG+
Sbjct: 273 FVDLKRRRYYFPLMVGIGLLALQQLGGINGVLFYSSTIFESAGVTSSN---VATFGVGVV 329
Query: 351 KTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE----KGPLVIALGILFV 406
+ V +A L+DK GR+ LL+ S+IGMT L + V L E + L ++ V
Sbjct: 330 QVVATGIATWLVDKAGRRLLLMISSIGMTISLVIVAVAFYLKEFVSPDSNMYNILSMVSV 389
Query: 407 CGNVAFF---SVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAI 463
G VA S+G+GP+ W++ SEI P+ ++ A ++ + N S LV M+ ++ A
Sbjct: 390 VGVVAMVISCSLGMGPIPWLIMSEILPVNIKGLAGSIATLLNWFVSWLVTMT-ANMLLAW 448
Query: 464 SFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
S GGTF L++ + +VFV VPETKGK+LE+I+ +F
Sbjct: 449 SSGGTFTLYALVCGFTVVFVSLWVPETKGKTLEEIQALF 487
>At5g59250 D-xylose-H+ symporter - like protein
Length = 558
Score = 227 bits (579), Expect = 1e-59
Identities = 147/465 (31%), Positives = 240/465 (51%), Gaps = 29/465 (6%)
Query: 55 VIACAIFASLNNVLLGYDVGVMSGAVIFIKED-------LKITEVQVEFLIGILSIVSLL 107
VI IF +L +L GYD+G SGA + ++ + VQ+ ++ +LL
Sbjct: 98 VILPFIFPALGGLLFGYDIGATSGATLSLQSPALSGTTWFNFSPVQLGLVVSGSLYGALL 157
Query: 108 GSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISP 167
GS+ +D +GR+ + +AAV++ +G + AP +L++GRLL G GIG + +P
Sbjct: 158 GSISVYGVADFLGRRRELIIAAVLYLLGSLITGCAPDLNILLVGRLLYGFGIGLAMHGAP 217
Query: 168 IYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFI 227
+YIAE P+ RG+L + E+FI +GI+LG+ + + V WR M G ++ +
Sbjct: 218 LYIAETCPSQIRGTLISLKELFIVLGILLGF--SVGSFQIDVVGGWRYMYGFGTPVALLM 275
Query: 228 GFALFIIPESPRWLV---------MQNRIEEARSVL--LKTNEDEKEVEERLAEIQQAAG 276
G ++ +P SPRWL+ +Q E+A L L+ ++ E+L + A
Sbjct: 276 GLGMWSLPASPRWLLLRAVQGKGQLQEYKEKAMLALSKLRGRPPGDKISEKLVD---DAY 332
Query: 277 FANSGKYEDKPVWRELLS--PPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGI 334
+ YED+ L P L+ + I G G+ FQQI+G + +YY+ IL AG
Sbjct: 333 LSVKTAYEDEKSGGNFLEVFQGPNLKALTIGG-GLVLFQQITGQPSVLYYAGSILQTAGF 391
Query: 335 EDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEK 394
+ +V +G+ K + VA+ +D +GR+PLLI G+ LF +
Sbjct: 392 SAAADATRVSVIIGVFKLLMTWVAVAKVDDLGRRPLLIGGVSGIALSLFLLSAYYKFLGG 451
Query: 395 GPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAM 454
PLV A+G L + V + + GP+ W++ SEIFPLR R + +L + N + +V
Sbjct: 452 FPLV-AVGALLL--YVGCYQISFGPISWLMVSEIFPLRTRGRGISLAVLTNFGSNAIVTF 508
Query: 455 SFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
+F + + + F LF I+ ++++FV +VPETKG SLE+IE
Sbjct: 509 AFSPLKEFLGAENLFLLFGGIALVSLLFVILVVPETKGLSLEEIE 553
>At3g05150 putative sugar transporter
Length = 463
Score = 223 bits (569), Expect = 2e-58
Identities = 148/473 (31%), Positives = 241/473 (50%), Gaps = 21/473 (4%)
Query: 33 DDNDDVLHQQQLEDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEV 92
D ++ +L + D V I A + G VG + I E+L ++
Sbjct: 6 DKSEPLLLPENGSDVSEEASWMVYLSTIIAVCGSYEFGTCVGYSAPTQFGIMEELNLSYS 65
Query: 93 QVEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGR 152
Q IL++ ++LG++ G+ SD IGRK M L++V+ +G + + LA L GR
Sbjct: 66 QFSVFGSILNMGAVLGAITSGKISDFIGRKGAMRLSSVISAIGWLIIYLAKGDVPLDFGR 125
Query: 153 LLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHIS 212
L G G G + P++IAEISP RG+L T ++FI +G+ ++ + ++
Sbjct: 126 FLTGYGCGTLSFVVPVFIAEISPRKLRGALATLNQLFIVIGLASMFL-------IGAVVN 178
Query: 213 WRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQ 272
WR + G+ P V + F + IPESPRWL M R + L K + + EIQ
Sbjct: 179 WRTLALTGVAPCVVLFFGTWFIPESPRWLEMVGRHSDFEIALQKLRGPQANITREAGEIQ 238
Query: 273 QAAGFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAA 332
+ + S + K +L+ R +I G+G+ FQQ GI+ ++Y+ +I ++A
Sbjct: 239 E---YLASLAHLPKATLMDLIDKKNI--RFVIVGVGLMFFQQFVGINGVIFYAQQIFVSA 293
Query: 333 GIEDK-SKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSL 391
G +L + V +T L A +LID++GR+PLL+ S +GM +G + L
Sbjct: 294 GASPTLGSILYSIEQVVLT----ALGATLLIDRLGRRPLLMASAVGMLIGCLLIGNSFLL 349
Query: 392 FEKG---PLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVC 448
G ++ AL + V + FS+G+G + WV+ SEIFP+ ++ A L V N +
Sbjct: 350 KAHGLALDIIPALAVSGVLVYIGSFSIGMGAIPWVIMSEIFPINLKGTAGGLVTVVNWLS 409
Query: 449 SGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMM 501
S LV+ +F + S GTF+++ + LAI+F+ LVPETKG++LE+I+ M
Sbjct: 410 SWLVSFTF-NFLMIWSPHGTFYVYGGVCVLAIIFIAKLVPETKGRTLEEIQAM 461
>At1g08930 ERD6 protein
Length = 496
Score = 222 bits (566), Expect = 3e-58
Identities = 139/445 (31%), Positives = 230/445 (51%), Gaps = 34/445 (7%)
Query: 70 GYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAA 129
G VG SGA I +DL ++ + IL++ L+G++ G+ +D++GRK TM
Sbjct: 73 GCGVGFSSGAQAGITKDLSLSVAEYSMFGSILTLGGLIGAVFSGKVADVLGRKRTMLFCE 132
Query: 130 VVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIF 189
G + + LA + L GRLL GIG+G + P+YIAEI+P RGS ++
Sbjct: 133 FFCITGWLCVALAQNAMWLDCGRLLLGIGVGIFSYVIPVYIAEIAPKHVRGSFVFANQLM 192
Query: 190 INVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEE 249
N GI L ++ + I WR++ VG++P VF F LF IPESPRWL R +E
Sbjct: 193 QNCGISLFFI-------IGNFIPWRLLTVVGLVPCVFHVFCLFFIPESPRWLAKLGRDKE 245
Query: 250 ARSVLLKTNEDEKEVEERLAEIQQAAGFA-NSGKYEDKPVWRELLSPPPALRRMLITGLG 308
RS L + + ++ I+ N G+ + +++ + P LI G+G
Sbjct: 246 CRSSLQRLRGSDVDISREANTIRDTIDMTENGGETKMSELFQRRYAYP------LIIGVG 299
Query: 309 IQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFI---LVAIVLIDKV 365
+ QQ+ G YY+ + G + + + T+ + ++A VL+DK+
Sbjct: 300 LMFLQQLCGSSGVTYYASSLFNKGG-------FPSAIGTSVIATIMVPKAMLATVLVDKM 352
Query: 366 GRKPLLIT--STIGMTACLFCMGVTLSLF----EKGPLVIALGILFVCGNVAFFSVGLGP 419
GR+ LL+ S +G++A L + F E P+ +G+L G++ F++G+G
Sbjct: 353 GRRTLLMASCSAMGLSALLLSVSYGFQSFGILPELTPIFTCIGVL---GHIVSFAMGMGG 409
Query: 420 VCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALA 479
+ W++ +EIFP+ V+ A L V N + ++ +F + + + G F +FS +SA +
Sbjct: 410 LPWIIMAEIFPMNVKVSAGTLVTVTNWLFGWIITYTFNFMLE-WNASGMFLIFSMVSASS 468
Query: 480 IVFVFTLVPETKGKSLEQIEMMFEN 504
IVF++ LVPETKG+SLE+I+ + N
Sbjct: 469 IVFIYFLVPETKGRSLEEIQALLNN 493
>At2g48020 sugar transporter like protein
Length = 463
Score = 217 bits (553), Expect = 1e-56
Identities = 141/452 (31%), Positives = 225/452 (49%), Gaps = 32/452 (7%)
Query: 54 YVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGSLGGG 113
+V C FA G G S A I+ DL +T + +L+ +++G++ G
Sbjct: 33 FVAVCGSFA------FGSCAGYSSPAQAAIRNDLSLTIAEFSLFGSLLTFGAMIGAITSG 86
Query: 114 RTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAEI 173
+D++GRK M +++ +G + + A L +GRL G G+G + PI+IAEI
Sbjct: 87 PIADLVGRKGAMRVSSAFCVVGWLAIIFAKGVVALDLGRLATGYGMGAFSYVVPIFIAEI 146
Query: 174 SPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFALFI 233
+P RG+LTT +I I G+ + ++ + ++WRV+ +GI+P LF
Sbjct: 147 APKTFRGALTTLNQILICTGVSVSFI-------IGTLVTWRVLALIGIIPCAASFLGLFF 199
Query: 234 IPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVWRELL 293
IPESPRWL R E + L K + ++ E AEIQ E P + L
Sbjct: 200 IPESPRWLAKVGRDTEFEAALRKLRGKKADISEEAAEIQDYI-----ETLERLPKAKMLD 254
Query: 294 SPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTV 353
R ++ G+ FQQ GI+ +Y+ I AG + + + + V
Sbjct: 255 LFQRRYIRSVLIAFGLMVFQQFGGINGICFYTSSIFEQAGFPTR----LGMIIYAVLQVV 310
Query: 354 FILVAIVLIDKVGRKPLLITSTIGMT-ACL-----FCMGVTLSLFEKGPLVIALGILFVC 407
+ ++D+ GRKPLL+ S G+ CL F + V E P++ +GI+
Sbjct: 311 ITALNAPIVDRAGRKPLLLVSATGLVIGCLIAAVSFYLKVHDMAHEAVPVLAVVGIMVYI 370
Query: 408 GNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGG 467
G+ FS G+G + WV+ SEIFP+ ++ A + + N + V+ +F + S+ G
Sbjct: 371 GS---FSAGMGAMPWVVMSEIFPINIKGVAGGMATLVNWFGAWAVSYTFNFLMSWSSY-G 426
Query: 468 TFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
TF +++AI+ALAIVFV +VPETKGK+LEQI+
Sbjct: 427 TFLIYAAINALAIVFVIAIVPETKGKTLEQIQ 458
>At3g05165 sugar transporter like protein
Length = 467
Score = 213 bits (543), Expect = 2e-55
Identities = 138/470 (29%), Positives = 234/470 (49%), Gaps = 36/470 (7%)
Query: 40 HQQQLEDKRNSTRKYVIACAIFASLNNVLLGYD----VGVMSGAVIFIKEDLKITEVQVE 95
HQ +D+R + AC I ++ V + G SGA I ++L ++ Q
Sbjct: 17 HQNDRDDRR------ITACVILSTFVAVCSAFSYGCAAGYTSGAETAIMKELDLSMAQFS 70
Query: 96 FLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLA 155
L++ +G+L G+ + I+GR+ T+ G +++ A + L +GR+
Sbjct: 71 AFGSFLNVGGAVGALFSGQLAVILGRRRTLWACDFFCVFGWLSIAFAKNVFWLDLGRISL 130
Query: 156 GIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRV 215
GIG+G + P+YIAEI+P RG+ T ++ N G+ L Y I+WRV
Sbjct: 131 GIGVGLISYVVPVYIAEITPKHVRGAFTASNQLLQNSGVSLIYF-------FGTVINWRV 183
Query: 216 MLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAA 275
M +G +P + +F IPESPRWL +E S L + + +V AEIQ
Sbjct: 184 MAVIGAIPCILQTIGIFFIPESPRWLAKIRLSKEVESSLHRLRGKDTDVSGEAAEIQVMT 243
Query: 276 GFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIE 335
+ K + ++ RR L+ G+G+ QQ+SG YYS I AG
Sbjct: 244 KMLEE---DSKSSFSDMFQ--KKYRRTLVVGIGLMLIQQLSGASGITYYSNAIFRKAGFS 298
Query: 336 DKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKG 395
++ ++ G+ LV ++L+D+ GR+PLL+ S +GM+ +GV+ +L +
Sbjct: 299 ER----LGSMIFGVFVIPKALVGLILVDRWGRRPLLLASAVGMSIGSLLIGVSFTLQQMN 354
Query: 396 ------PLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCS 449
P+ + + IL G AF G+G + WV+ SEIFP+ ++ A + A+ +
Sbjct: 355 VLPELIPIFVFVNILVYFGCFAF---GIGGLPWVIMSEIFPINIKVSAGTIVALTSWTSG 411
Query: 450 GLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
V+ +F + + S GTF++F+A+ ++ +F++ LVPETKG+SLE+++
Sbjct: 412 WFVSYAFNFMFE-WSAQGTFYIFAAVGGMSFIFIWMLVPETKGQSLEELQ 460
>At5g27360 sugar-porter family protein 2 (SFP2)
Length = 478
Score = 213 bits (542), Expect = 2e-55
Identities = 139/465 (29%), Positives = 243/465 (51%), Gaps = 26/465 (5%)
Query: 43 QLEDKRNSTRKYVIACAIFASLNNVL----LGYDVGVMSGAVIFIKEDLKITEVQVEFLI 98
QL+++ + + + AC I ++ V G +G SGA I I +DL ++ Q
Sbjct: 19 QLKNQNDDSECRITACVILSTFIAVCGSFSFGVSLGYTSGAEIGIMKDLDLSIAQFSAFA 78
Query: 99 GILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIG 158
+ ++ + +G+L G+ + I+GR+ TM ++ ++ +G ++ A L GR+ +GIG
Sbjct: 79 SLSTLGAAIGALFSGKMAIILGRRKTMWVSDLLCIIGWFSIAFAKDVMWLNFGRISSGIG 138
Query: 159 IGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLA 218
+G + P+YIAEISP RG+ T ++ N G+ + Y FSG ++WR++
Sbjct: 139 LGLISYVVPVYIAEISPKHVRGTFTFTNQLLQNSGLAMVY-----FSG--NFLNWRILAL 191
Query: 219 VGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFA 278
+G LP LF +PESPRWL +E + LL+ ++ ++I+
Sbjct: 192 LGALPCFIQVIGLFFVPESPRWLAKVGSDKELENSLLRLRGGNADISREASDIEVMTKMV 251
Query: 279 NSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKS 338
+ + K + +L R L+ G+G+ QQ SG A + Y+ IL AG
Sbjct: 252 EN---DSKSSFCDLFQ--RKYRYTLVVGIGLMLIQQFSGSSAVLSYASTILRKAGF---- 302
Query: 339 KLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLV 398
+ + +G+ ++ ++L+DK GR+PLL+TS GM +GV +L +K L+
Sbjct: 303 SVTIGSTLLGLFMIPKAMIGVILVDKWGRRPLLLTSVSGMCITSMLIGVAFTL-QKMQLL 361
Query: 399 IALG--ILFVCGN--VAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAM 454
L F+C + +++GLG + WV+ SEIFP+ ++ A ++ + + S +V
Sbjct: 362 PELTPVFTFICVTLYIGTYAIGLGGLPWVIMSEIFPMNIKVTAGSIVTLVSWSSSSIVTY 421
Query: 455 SFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
+F + + S GTF++F A+ LA++F++ LVPETKG SLE+I+
Sbjct: 422 AFNFLLE-WSTQGTFYVFGAVGGLALLFIWLLVPETKGLSLEEIQ 465
>At5g27350 sugar transporter like protein
Length = 474
Score = 213 bits (542), Expect = 2e-55
Identities = 137/466 (29%), Positives = 237/466 (50%), Gaps = 26/466 (5%)
Query: 43 QLEDKRNSTRKYVIACAIFASLNNVLLGYDVGVM----SGAVIFIKEDLKITEVQVEFLI 98
QL++K + + + AC I ++ V + GV SGA + +DL ++ Q
Sbjct: 15 QLKNKNDDSECRITACVILSTFVAVCGSFSFGVATGYTSGAETGVMKDLDLSIAQFSAFG 74
Query: 99 GILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIG 158
++ + +G+L G + +IGR+ TM ++ + G +++ A +L GR+++GIG
Sbjct: 75 SFATLGAAIGALFCGNLAMVIGRRGTMWVSDFLCITGWLSIAFAKEVVLLNFGRIISGIG 134
Query: 159 IGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGI-MLGYVSNYAFSGLSVHISWRVML 217
G + P+YIAEI+P RG+ T ++ N G+ M+ + N+ I+WR +
Sbjct: 135 FGLTSYVVPVYIAEITPKHVRGTFTFSNQLLQNAGLAMIYFCGNF--------ITWRTLA 186
Query: 218 AVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGF 277
+G LP LF +PESPRWL +E + L + + ++ +EIQ
Sbjct: 187 LLGALPCFIQVIGLFFVPESPRWLAKVGSDKELENSLFRLRGRDADISREASEIQVMTKM 246
Query: 278 ANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDK 337
+ + K + +L R L+ G+G+ QQ SG A + Y+ I AG
Sbjct: 247 VEN---DSKSSFSDLFQ--RKYRYTLVVGIGLMLIQQFSGSAAVISYASTIFRKAGF--- 298
Query: 338 SKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEK--- 394
+ T +GI ++ ++L+DK GR+PLL+TS GM+ +GV +L +
Sbjct: 299 -SVAIGTTMLGIFVIPKAMIGLILVDKWGRRPLLMTSAFGMSMTCMLLGVAFTLQKMQLL 357
Query: 395 GPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAM 454
L L + V +A +++GLG + WV+ SEIFP+ ++ A ++ + + S +V
Sbjct: 358 SELTPILSFICVMMYIATYAIGLGGLPWVIMSEIFPINIKVTAGSIVTLVSFSSSSIVTY 417
Query: 455 SFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEM 500
+F + + S GTFF+F+ I A++F++ LVPETKG SLE+I++
Sbjct: 418 AFNFLFE-WSTQGTFFIFAGIGGAALLFIWLLVPETKGLSLEEIQV 462
>At3g05157 sugar transporter like protein
Length = 458
Score = 212 bits (540), Expect = 4e-55
Identities = 135/470 (28%), Positives = 237/470 (49%), Gaps = 36/470 (7%)
Query: 40 HQQQLEDKRNSTRKYVIACAIFASLNNVLLGYDVGVM----SGAVIFIKEDLKITEVQVE 95
H+ +D+R + AC I ++ V + G SGA I ++L ++ Q
Sbjct: 8 HENDRDDRR------ITACVILSTFVAVCSSFSYGCANGYTSGAETAIMKELDLSMAQFS 61
Query: 96 FLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLA 155
L++ +G+L G+ + I+GR+ T+ + G +++ A + L +GR+
Sbjct: 62 AFGSFLNLGGAVGALFSGQLAVILGRRRTLWACDLFCIFGWLSIAFAKNVLWLDLGRISL 121
Query: 156 GIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRV 215
GIG+G + P+YIAEI+P RG+ + + N GI L Y I+WRV
Sbjct: 122 GIGVGLTSYVVPVYIAEITPKHVRGAFSASTLLLQNSGISLIYF-------FGTVINWRV 174
Query: 216 MLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAA 275
+ +G LP ++ IPESPRWL ++E + L + + +V + AEIQ
Sbjct: 175 LAVIGALPCFIPVIGIYFIPESPRWLAKIGSVKEVENSLHRLRGKDADVSDEAAEIQVMT 234
Query: 276 GFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIE 335
+ K + ++ RR L+ G+G+ QQ+SG YYS I AG
Sbjct: 235 KMLEE---DSKSSFCDMFQ--KKYRRTLVVGIGLMLIQQLSGASGITYYSNAIFRKAGFS 289
Query: 336 DKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKG 395
++ ++ G+ LV ++L+D+ GR+PLL+ S +GM+ +GV+ +L E
Sbjct: 290 ER----LGSMIFGVFVIPKALVGLILVDRWGRRPLLLASAVGMSIGSLLIGVSFTLQEMN 345
Query: 396 ------PLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCS 449
P+ + + IL G FF++G+G + W++ SEIFP+ ++ A ++ A+ +
Sbjct: 346 LFPEFIPVFVFINILVYFG---FFAIGIGGLPWIIMSEIFPINIKVSAGSIVALTSWTTG 402
Query: 450 GLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
V+ F + + S GTF++F+ + L+++F++ LVPETKG+SLE+++
Sbjct: 403 WFVSYGFNFMFE-WSAQGTFYIFAMVGGLSLLFIWMLVPETKGQSLEELQ 451
>At5g18840 sugar transporter - like protein
Length = 482
Score = 211 bits (538), Expect = 6e-55
Identities = 154/494 (31%), Positives = 239/494 (48%), Gaps = 38/494 (7%)
Query: 22 NKYKRMNSHLVDDNDDVLHQQQLEDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVI 81
NK + + + DD + E + N + V+ A + G VG +
Sbjct: 16 NKVEDLGKPFLTHEDD-----EKESENNESYLMVLFSTFVAVCGSFEFGSCVGYSAPTQS 70
Query: 82 FIKEDLKITEVQVEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTL 141
I++DL ++ + IL+I ++LG++ G+ SD GRK M +A G + +
Sbjct: 71 SIRQDLNLSLAEFSMFGSILTIGAMLGAVMSGKISDFSGRKGAMRTSACFCITGWLAVFF 130
Query: 142 APSYQVLMIGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSN 201
+L +GR G GIG + P+YIAEISP RG LTT ++ I +G + ++
Sbjct: 131 TKGALLLDVGRFFTGYGIGVFSYVVPVYIAEISPKNLRGGLTTLNQLMIVIGSSVSFL-- 188
Query: 202 YAFSGLSVHISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDE 261
+ ISW+ + G+ P + + F L IPESPRWL +E R L K +
Sbjct: 189 -----IGSLISWKTLALTGLAPCIVLLFGLCFIPESPRWLAKAGHEKEFRVALQKLRGKD 243
Query: 262 KEVEERLAEIQ---QAAGFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGI 318
++ IQ QA + +D L+S R +I G+ + FQQ GI
Sbjct: 244 ADITNEADGIQVSIQALEILPKARIQD------LVS--KKYGRSVIIGVSLMVFQQFVGI 295
Query: 319 DATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIG- 377
+ +Y+ E + AG KL T+A+ + ++ +LIDK GR+PL++ S G
Sbjct: 296 NGIGFYASETFVKAGF-TSGKL--GTIAIACVQVPITVLGTILIDKSGRRPLIMISAGGI 352
Query: 378 -----MTACLFCMGVTLSLFEKGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLR 432
+T F + L E P + G+L VA FS+G+GPV WV+ SEIFP+
Sbjct: 353 FLGCILTGTSFLLKGQSLLLEWVPSLAVGGVLIY---VAAFSIGMGPVPWVIMSEIFPIN 409
Query: 433 VRAQASALGAVANRVCSGLVAMSF-LSVSDAISFGGTFFLFSAISALAIVFVFTLVPETK 491
V+ A +L + N SG A+S+ + + S GTF+L+SA +A I+FV +VPETK
Sbjct: 410 VKGIAGSLVVLVN--WSGAWAVSYTFNFLMSWSSPGTFYLYSAFAAATIIFVAKMVPETK 467
Query: 492 GKSLEQIEMMFENE 505
GK+LE+I+ E
Sbjct: 468 GKTLEEIQACIRRE 481
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.140 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,882,411
Number of Sequences: 26719
Number of extensions: 454380
Number of successful extensions: 1680
Number of sequences better than 10.0: 96
Number of HSP's better than 10.0 without gapping: 65
Number of HSP's successfully gapped in prelim test: 31
Number of HSP's that attempted gapping in prelim test: 1306
Number of HSP's gapped (non-prelim): 122
length of query: 530
length of database: 11,318,596
effective HSP length: 104
effective length of query: 426
effective length of database: 8,539,820
effective search space: 3637963320
effective search space used: 3637963320
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC126008.6


