

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
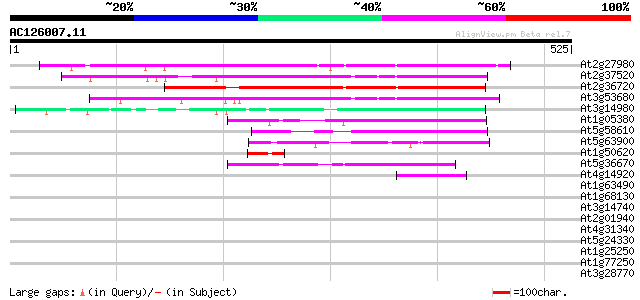
Query= AC126007.11 + phase: 0
(525 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g27980 hypothetical protein 317 1e-86
At2g37520 unknown protein 273 2e-73
At2g36720 putative PHD-type zinc finger protein 238 5e-63
At3g53680 putative protein 234 1e-61
At3g14980 PHD-finger protein, putative 95 1e-19
At1g05380 unknown protein 89 6e-18
At5g58610 putative protein 87 3e-17
At5g63900 putative protein 79 6e-15
At1g50620 hypothetical protein 46 4e-05
At5g36670 putative protein 43 4e-04
At4g14920 hypothetical protein 42 6e-04
At1g63490 RB-binding protein -like 41 0.001
At1g68130 putative C2H2-type zinc finger protein 41 0.002
At3g14740 unknown protein 39 0.005
At2g01940 putative C2H2-type zinc finger protein 39 0.009
At4g31340 unknown protein 37 0.021
At5g24330 putative protein 37 0.027
At1g25250 Indeterminate 1 like Zn finger containing protein 37 0.027
At1g77250 unknown protein 36 0.046
At3g28770 hypothetical protein 36 0.060
>At2g27980 hypothetical protein
Length = 1008
Score = 317 bits (811), Expect = 1e-86
Identities = 188/471 (39%), Positives = 274/471 (57%), Gaps = 38/471 (8%)
Query: 29 KNKKKKKNKNKKESNLRSNESKNLQPSP-----KHTTQTSNSHASPTTINYRDQCLHKLV 83
++K + N++E R + + P K+++ SNSH T +D LHKLV
Sbjct: 522 RSKSSPRQSNRREQPTRKSTEPGVVPGTILSESKNSSIKSNSHGKLTR---KDLRLHKLV 578
Query: 84 FQENVLEDGAAVGYFVYEEVSPSKFEAHAGWASRRKPWPLII--------EFLFPIVMNT 135
F++++L DG VGYFV EVSPS FEAHAG ASRRKP+ I E + M+
Sbjct: 579 FEDDILPDGTEVGYFVAGEVSPSTFEAHAGCASRRKPFQHIYTTNGVSLHELSVALSMDQ 638
Query: 136 AVSANKEP-------------VESPPKEKWYCEYCRNKLQKDKNVEHKENVVTTQKIIES 182
S ++ + S P E+W C+YC N ++++K V+ N + ++
Sbjct: 639 RFSIHENDDLCSICRDGVCASLPSLPSERWSCKYCVNMVEREKFVDSNLNAIAAGRVQGV 698
Query: 183 DPSEQIAKICTLSVKHKEVEHSS-CALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKD 241
D +I C V E S C LC F F+ TV+ICDQCEK++HVGCLK+
Sbjct: 699 DAIAEITNRCIRIVSSFVTELPSVCVLCRGHSFCRLGFNARTVIICDQCEKEFHVGCLKE 758
Query: 242 HNMANLKKVPKHYWFCGVDCYDIHMKLKNFMARGDVLLSDSLLSLIKNKKEQKGLET--- 298
++A+LK++P+ WFC + C +I+ L N + RG+ LS+++L+ ++ KKEQ E
Sbjct: 759 RDIADLKELPEEKWFCSLGCEEINTTLGNLIVRGEEKLSNNILNFLR-KKEQPNEENCPD 817
Query: 299 -EFGLDIKWKVFNRQLIVSKIITSSLLSDVVTIFHEQFDSIVVTGTKIDLIPAMVKGRKI 357
+ DI+W+V + +L S T LL+ ++I HE+FD I +GTK DLIPAMV GR+
Sbjct: 818 YKTTPDIRWRVLSGKL-TSSDDTKILLAKALSILHERFDPISESGTKGDLIPAMVYGRQT 876
Query: 358 KDKYYFGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIEN 417
K + F GMYC +L V++V+VS GIFRVFG E+AEL L+AT + Q QG+F+CL +CIE
Sbjct: 877 KAQD-FSGMYCTMLAVDEVIVSVGIFRVFGSELAELPLVATSKDCQGQGYFQCLFACIER 935
Query: 418 VLKELKVERLVLPAAHEAESMWIDKFGFTEPNQGLGRRYYRRSWSFHLNKG 468
+L L V+ +VLPAA EA+S+W DKFGFT+ + YR+ +S + G
Sbjct: 936 LLGFLNVKHIVLPAADEAKSIWTDKFGFTKMTDEEVKE-YRKDYSVMIFHG 985
>At2g37520 unknown protein
Length = 825
Score = 273 bits (698), Expect = 2e-73
Identities = 164/447 (36%), Positives = 238/447 (52%), Gaps = 64/447 (14%)
Query: 49 SKNLQPSPKHTTQTSNSHASPTTINY---------RDQCLHKLVFQENVLEDGAAVGYFV 99
+K+ PK + SH S T + RD LH+L+F N L DG + Y+V
Sbjct: 353 AKDTLDEPKRIAKKLTSHVSGTGCHKKVSEGSNRKRDNDLHRLLFMPNGLPDGTELAYYV 412
Query: 100 YEEVSPSKFEAHAGWASRRKPWPLIIEF----LFPIVMNTA-----VSANKEPV------ 144
++SPS+FEAHAG A+RR+P+ I L I M+ A + + + +
Sbjct: 413 KTQISPSQFEAHAGMAARRQPYRHIFISSGLSLHDIAMSLANGHVITTGDSDDMCSICGD 472
Query: 145 ---------------------ESPPKEKWYCEYCRNKLQKDKNVEHKENVVTTQKIIESD 183
+S P+ WYC C + ++++K +D
Sbjct: 473 GGDLLLCAGCPQAFHTACLKFQSMPEGTWYCSSCN------------DGPISSKKATTTD 520
Query: 184 PSEQIAKIC---TLSVKHKEVEHSSCALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLK 240
PS I + VK E + C C F+ G+F TV++CDQCEK+YHVGCL+
Sbjct: 521 PSGNARPIVIRLSRVVKAPESDIGGCVFCRSHDFSIGKFDDRTVILCDQCEKEYHVGCLR 580
Query: 241 DHNMANLKKVPKHYWFCGVDCYDIHMKLKNFMARGDVLLSDSLLSLIKNKKEQKGLETEF 300
++ +LK++P+ WFC +C IH ++N ++ G L LL +I K +KG+ T+
Sbjct: 581 ENGFCDLKEIPQEKWFCCSNCSRIHTAVQNSVSCGPQTLPTPLLDMICRKDREKGIFTDI 640
Query: 301 GLDIKWKVFNRQLIVSKIITSSLLSDVVTIFHEQFDSIVVTGTKIDLIPAMVKGRKIKDK 360
G ++W++ + + + + LLS IF E FD IV + DLIP MV GR I +
Sbjct: 641 GDTVEWRILSGKSRYPEHLP--LLSRAAVIFRECFDPIVAKSGR-DLIPVMVYGRNISGQ 697
Query: 361 YYFGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLK 420
FGGMYC VLIVN +VVSA + R+FG+EVAEL ++AT EYQ +G+F+ L +C+EN+L
Sbjct: 698 E-FGGMYCLVLIVNSLVVSAALLRIFGQEVAELPIVATSREYQGRGYFQGLYACVENLLS 756
Query: 421 ELKVERLVLPAAHEAESMWIDKFGFTE 447
L VE LVLPAA EAES+W KFGFT+
Sbjct: 757 SLNVENLVLPAAEEAESIWTKKFGFTK 783
>At2g36720 putative PHD-type zinc finger protein
Length = 958
Score = 238 bits (608), Expect = 5e-63
Identities = 126/300 (42%), Positives = 185/300 (61%), Gaps = 14/300 (4%)
Query: 146 SPPKEKWYCEYCRNKLQKDKNVEHKENVVTTQKIIESDPSEQIAKICTLSVKHKEVEHSS 205
S P+ W+C+YC NK + E+ N ++ DP +Q+A C VK+ E E
Sbjct: 608 SIPRGNWHCKYCENKFTSEIAGEYNVNSSAVGQLEGVDPVDQLAGRCIRVVKNMEAET-- 665
Query: 206 CALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYDIH 265
N F P T++ICDQCEK+YH+GCL N+ +LK++PK WFC +DC I+
Sbjct: 666 ---------NGSGFGPRTIIICDQCEKEYHIGCLSSQNIVDLKELPKGNWFCSMDCTRIN 716
Query: 266 MKLKNFMARGDVLLSDSLLSLIKNKKEQKGLETEFGLDIKWKVFNRQLIVSKIITSSLLS 325
L+ + G LSDS L +I+ K+E+ + + LDI+W++ + + V+ + LLS
Sbjct: 717 STLQKLLLGGAEKLSDSSLGIIQTKQERNDVYSISDLDIRWRLISGK--VTSPESRMLLS 774
Query: 326 DVVTIFHEQFDSIVVTGTKIDLIPAMVKGRKIKDKYYFGGMYCAVLIVNQVVVSAGIFRV 385
+ IFH+ FD IV + +LIP MV G+ ++ + Y GG+ CAVL VN VVSAG+ RV
Sbjct: 775 QALAIFHDCFDPIVDPLSGSNLIPRMVYGKTMQGQDY-GGICCAVLTVNATVVSAGLLRV 833
Query: 386 FGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKFGF 445
FG+EVAEL L+AT+ +++G+F+ L SCIE +L L VE +V+PAA EAE +W++KFGF
Sbjct: 834 FGREVAELPLVATRMCSREKGYFQLLFSCIEKLLSSLNVESIVVPAAEEAEPLWMNKFGF 893
Score = 29.6 bits (65), Expect = 4.3
Identities = 12/24 (50%), Positives = 16/24 (66%)
Query: 75 RDQCLHKLVFQENVLEDGAAVGYF 98
+DQ LHKLVF L +G +GY+
Sbjct: 492 KDQGLHKLVFDRGGLPEGTELGYY 515
Score = 29.3 bits (64), Expect = 5.6
Identities = 9/34 (26%), Positives = 19/34 (55%), Gaps = 6/34 (17%)
Query: 224 VMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFC 257
+++CD C + +H+ C+ +L +P+ W C
Sbjct: 589 LLLCDSCPRAFHIECV------SLPSIPRGNWHC 616
>At3g53680 putative protein
Length = 839
Score = 234 bits (596), Expect = 1e-61
Identities = 156/428 (36%), Positives = 227/428 (52%), Gaps = 48/428 (11%)
Query: 75 RDQCLHKLVFQENVLEDGAAVGYFVYEE--------------------VSPSKFEAHAGW 114
RD LH+L+F N L DG + Y+V + +SPS+FEAHAG
Sbjct: 389 RDNDLHRLLFLPNGLPDGTELAYYVKSQKLLQGYKQGSGIVCSCCDTKISPSQFEAHAGM 448
Query: 115 ASRRKPWPLIIEFLFPIVMNTAVS-ANKEPVESPPKEKWYCEYCRN-----------KLQ 162
A RR+P+ I + + AVS A+ V + C C N +
Sbjct: 449 AGRRQPYRRIHISSGLSLHDIAVSLADGGHVITTGDSDDMCSICGNGGDLLLCAGCPQAF 508
Query: 163 KDKNVEHKENVVTTQKIIESDPSEQIAKICTLSVKHKE--------VEHSSCALC--SER 212
++ + T + KI T S + V HS+ +L S+R
Sbjct: 509 HTACLKFQSMPEGTWYCSSCNDGPTSCKIATASWLYTYFNLNANILVLHSAYSLSPISDR 568
Query: 213 H--FNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYDIHMKLKN 270
F+ G+F TV++CDQCEK+YHVGCL+++ + +LK +P+ WFC DC IH L++
Sbjct: 569 SHDFSIGKFDDRTVILCDQCEKEYHVGCLRENELCDLKGIPQDKWFCCSDCSRIHRVLQS 628
Query: 271 FMARGDVLLSDSLLSLIKNKKEQKGLETEFGLDIKWKVFNRQLIVSKIITSSLLSDVVTI 330
+ G + LL I K +KG+ + G ++W++ + + + + LLS TI
Sbjct: 629 SASCGPQTIPTLLLDTISRKYREKGIYIDNGNTVEWRMLSGKSRYPEHLP--LLSRAATI 686
Query: 331 FHEQFDSIVVTGTKIDLIPAMVKGRKIKDKYYFGGMYCAVLIVNQVVVSAGIFRVFGKEV 390
F E FD IV + DLIP MV GR I + FGGMYC VL+VN +VVSA + R+FG++V
Sbjct: 687 FRECFDPIVAKSGR-DLIPVMVYGRNISGQE-FGGMYCLVLMVNSLVVSAALLRIFGQKV 744
Query: 391 AELSLIATKAEYQKQGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKFGFTEPNQ 450
AEL ++AT EYQ +G+F+ L +C+EN+L L VE L+LPAA EAES+W +KFGFT+ +
Sbjct: 745 AELPIVATSREYQGRGYFQGLFACVENLLSSLNVENLLLPAAEEAESIWTNKFGFTKMTE 804
Query: 451 GLGRRYYR 458
+RY R
Sbjct: 805 HRLQRYQR 812
>At3g14980 PHD-finger protein, putative
Length = 1189
Score = 94.7 bits (234), Expect = 1e-19
Identities = 116/484 (23%), Positives = 191/484 (38%), Gaps = 88/484 (18%)
Query: 6 EELKDNIVVEAVAKDGDDVEKKMKNKKK---KKNKNKKESNLRSNESKNLQPSPKHTTQT 62
++L + + K +KK K K KK N+ L S N++ H Q
Sbjct: 555 DDLMGSTITRNKGKFSRSSQKKKTQKPKARTKKRNNRGGCRLLPRSSSNVE---NHFFQG 611
Query: 63 SNSHASPTT----------------INYRDQCLHKLVFQENVLEDGAAVGYFVYEEVSPS 106
+ S P T I RD +V V +DG V + VS S
Sbjct: 612 NWSILGPRTVLSWLIATKVISRDEVIQLRDPDDDTVVKTGLVTKDGV-VCTCCNKTVSLS 670
Query: 107 KFEAHAGWASRRKPWPLIIEFLFPIVMNTAVSANKEPVESPPKEKWYCEYCRNKLQKDKN 166
+F+ HAG+ + P + F+ + +P S E W EY
Sbjct: 671 EFKNHAGF---NQNCPCLNLFM----------GSGKPFASCQLEAWSAEY---------- 707
Query: 167 VEHKENVVTTQKIIESDPSEQIAKIC-----------TLSVKHKEV-------------E 202
+ + N +K + DP++ +C S H+
Sbjct: 708 -KARRNGWRLEKASDDDPNDDSCGVCGDGGELICCDNCPSTFHQACLSMQVLPEGSWYCS 766
Query: 203 HSSCALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCY 262
+C +CSE +N E S C QC YH CL+ ++ +K+ +FCG +C
Sbjct: 767 SCTCWICSELVSDNAERSQ--DFKCSQCAHKYHGTCLQ--GISKRRKLFPETYFCGKNCE 822
Query: 263 DIHMKLKNFMARGDVLLSDSLLSLIKNKKEQKGLETEFGLDIKWKVFNRQLIVSKIITSS 322
++ L + + + S++K +E V + + + K +S
Sbjct: 823 KVYNGLSSRVGIINPNADGLSWSILKCFQEDG------------MVHSARRLALKAECNS 870
Query: 323 LLSDVVTIFHEQFDSIVVTGTKIDLIP-AMVKGRKIKDKYYFGGMYCAVLIVNQVVVSAG 381
L+ ++I E F S+V T ID+IP + + F G Y V+ + V++S
Sbjct: 871 KLAVALSIMEESFLSMVDPRTGIDMIPHVLYNWGSTFARLDFDGFYTVVVEKDDVMISVA 930
Query: 382 IFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWID 441
RV G +AE+ L+AT ++Y++QG + L++ IE +L LKVE+LV+ A W +
Sbjct: 931 SIRVHGVTIAEMPLVATCSKYRRQGMCRILVAAIEEMLMSLKVEKLVVAALPSLVETWTE 990
Query: 442 KFGF 445
FGF
Sbjct: 991 GFGF 994
>At1g05380 unknown protein
Length = 1138
Score = 89.0 bits (219), Expect = 6e-18
Identities = 66/255 (25%), Positives = 113/255 (43%), Gaps = 38/255 (14%)
Query: 205 SCALCSERHFNNGEFSPW-TVMICDQCEKDYHVGCLKD--HNMANLKKVPKHYWFCGVDC 261
+C C + G+ + +++ C CE+ YH CL D H + + FCG C
Sbjct: 668 TCKFCDAAVASGGKDGNFISLLSCGMCERRYHQLCLNDEAHKVQSFGSASS---FCGPKC 724
Query: 262 YDIHMKLKNFMARGDVLLSDSLLSLIKNKKEQKGLETEFGLDIKWKVFNR---------Q 312
++ KL+ ++ G++TE W + +R Q
Sbjct: 725 LELFEKLQKYL----------------------GVKTEIEGGYSWSLIHRVDTDSDTNSQ 762
Query: 313 LIVSKIITSSLLSDVVTIFHEQFDSIVVTGTKIDLIPAMVKGRKIK-DKYYFGGMYCAVL 371
+ +I +S L+ + I E F IV + +DLI ++ ++ + G Y A+L
Sbjct: 763 MSAQRIENNSKLAVGLAIMDECFLPIVDRRSGVDLIRNVLYNCGSNFNRINYTGFYTAIL 822
Query: 372 IVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKELKVERLVLPA 431
++SA R G ++AE+ I T+ Y++QG + L IE+ ++ LKVE+LV+PA
Sbjct: 823 ERGDEIISAASLRFHGMQLAEMPFIGTRHIYRRQGMCRRLFDAIESAMRSLKVEKLVIPA 882
Query: 432 AHEAESMWIDKFGFT 446
+ W FGFT
Sbjct: 883 IPDFLHAWTGNFGFT 897
>At5g58610 putative protein
Length = 1065
Score = 86.7 bits (213), Expect = 3e-17
Identities = 65/223 (29%), Positives = 97/223 (43%), Gaps = 39/223 (17%)
Query: 227 CDQCEKDYHVGCLK-DHNMANLKKVPKHYWFCGVDCYDIHMKLKNFMARGDVLLSDSLLS 285
C QCE YH CL+ D +L K+ WFC DC + S
Sbjct: 761 CKQCELKYHPSCLRYDGACDSLDKILGEKWFCSKDCEE---------------------S 799
Query: 286 LIKNKKEQKGLETEFGLDIKWKVFNRQLIVSKIITSSLLSDVVTIFHEQFDSIVVTGTKI 345
L N + E + +LS + + HE F+ +
Sbjct: 800 LEPNMYGDDASKIEAAAE----------------NHCILSVALDVMHELFEPVKRPHGGR 843
Query: 346 DLIPAMVKGRKIKDKYY-FGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQK 404
DL ++ R K K F G Y +L N +VS R+ GK+VAE+ I T+ ++++
Sbjct: 844 DLAEDVIFSRWSKFKRLNFSGFYTVLLERNNELVSVATVRILGKKVAEMPFIGTRFQHRQ 903
Query: 405 QGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKFGFTE 447
+G + L++ +E VL +L VERLVLPA + WI+ FGFT+
Sbjct: 904 RGMCRVLINELEKVLIDLGVERLVLPAVPCVLNTWINSFGFTK 946
>At5g63900 putative protein
Length = 557
Score = 79.0 bits (193), Expect = 6e-15
Identities = 62/230 (26%), Positives = 107/230 (45%), Gaps = 33/230 (14%)
Query: 224 VMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYDIHMKLKNFMARGDVLLSDSL 283
+M C+QC++ +H+ CLK+ + V WFC C + L+N + + +D
Sbjct: 315 LMACEQCQRRFHLTCLKEDSCI----VSSRGWFCSSQCNRVFSALENLLGSKIAVGNDGD 370
Query: 284 L--SLIKNKKEQKGLETEFGLDIKWKVFNRQLIVSKIITSSLLSDVVTIFHEQFDSIVVT 341
L +L++ E + + E S L V I H+ F+
Sbjct: 371 LVWTLMRAPNEGEHYDDE--------------------QISKLESAVEILHQGFEPTNDV 410
Query: 342 GTKIDLIPAMVKGRKIKDKYYFGGMYCAVLIV--NQVVVSAGIFRVFGKEVAELSLIATK 399
+ DL+ ++ KD+ G + VLI N+ + A + RV K+V E+ L+AT
Sbjct: 411 FSGRDLVEELIYR---KDRTGVGRGFYTVLIERKNEPITVAAV-RV-DKDVVEIPLVATL 465
Query: 400 AEYQKQGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKFGFTEPN 449
+ Y++ G + L+ +E + ++ V RLVLPAA E + W ++FGF+ N
Sbjct: 466 SSYRRSGMCRVLMDELEKQMSQMGVCRLVLPAAKEVVTTWTERFGFSVMN 515
>At1g50620 hypothetical protein
Length = 629
Score = 46.2 bits (108), Expect = 4e-05
Identities = 20/35 (57%), Positives = 23/35 (65%), Gaps = 3/35 (8%)
Query: 223 TVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFC 257
TV+ICD CEK YH+ CL H N+K VPK W C
Sbjct: 336 TVLICDACEKGYHLKCLHAH---NIKGVPKSEWHC 367
>At5g36670 putative protein
Length = 1030
Score = 43.1 bits (100), Expect = 4e-04
Identities = 45/216 (20%), Positives = 90/216 (40%), Gaps = 19/216 (8%)
Query: 205 SCALCSERHFNNGEFSPW-TVMICDQCEKDYHVGCL-KDHNMANLKKVPKHYWFCGVDCY 262
SC C + E S ++ C CE+ YH C+ +D + + FCG C
Sbjct: 540 SCKFCEKDEAAKHETSTLPSLSSCRLCEEKYHQACINQDGTVPGERSTDS---FCGKYCQ 596
Query: 263 DIHMKLKNFMARGDVLLSDSLLSLIKNKKEQKGLETEFGLDIKWKVFNRQLIVSKIITSS 322
++ +L+ F+ L S ++ ++ +V + I KI ++
Sbjct: 597 ELFEELQLFIGVKHPLPEGFSWSFLRR------------FELPSEVADCD-ISEKIAYNA 643
Query: 323 LLSDVVTIFHEQFDSIVVTGTKIDLIPAMVKGRKIK-DKYYFGGMYCAVLIVNQVVVSAG 381
++ ++ E F +V + ++L+ +V + F AVL +++
Sbjct: 644 KMAVAFSVMDECFSPLVDHRSGVNLLQNIVYNFGSNFHRLDFSSFLTAVLERGDEIIAVA 703
Query: 382 IFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIEN 417
R+ G ++AE+ I T+ Y++QG + L+ IE+
Sbjct: 704 SIRIHGNQLAEMPFIGTRYMYRRQGMCRRLMDGIES 739
>At4g14920 hypothetical protein
Length = 1040
Score = 42.4 bits (98), Expect = 6e-04
Identities = 22/65 (33%), Positives = 34/65 (51%)
Query: 363 FGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKEL 422
FGG Y A+L +V++ R G +AE+ I T+ Y+ QG + L S +E+V
Sbjct: 829 FGGFYTALLERGDEIVASASIRFHGNRLAEMPFIGTRHVYRHQGMCRRLFSVVESVSSTA 888
Query: 423 KVERL 427
V +L
Sbjct: 889 DVAKL 893
>At1g63490 RB-binding protein -like
Length = 1458
Score = 41.2 bits (95), Expect = 0.001
Identities = 33/131 (25%), Positives = 61/131 (46%), Gaps = 16/131 (12%)
Query: 155 EYCRNKLQKDKNVEHKENVVTTQKIIESDPSEQI---AKICTLSVKHKEVEHSSCALC-S 210
E N++ D + K + ++ ES S++ A + V+++E +C C S
Sbjct: 184 ENYHNRMNVDASKSCKRKINGIRRCSESRSSKRRKRNADVKNPKVENEEGVDQACEQCKS 243
Query: 211 ERHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYDIHMKLKN 270
++H GE +++CD C K +H+ CL LK +P W+C ++C + +
Sbjct: 244 DKH---GE----VMLLCDSCNKGWHIYCLS----PPLKHIPLGNWYC-LECLNTDEETFG 291
Query: 271 FMARGDVLLSD 281
F+ +LL D
Sbjct: 292 FVPGKCLLLED 302
>At1g68130 putative C2H2-type zinc finger protein
Length = 419
Score = 40.8 bits (94), Expect = 0.002
Identities = 29/133 (21%), Positives = 61/133 (45%), Gaps = 12/133 (9%)
Query: 150 EKWYCEYCRNKLQKDKNVE-HKENVVTTQKIIESDPSEQIAKICTLSVKHKEVEHSSCAL 208
+++ CE C Q+D+N++ H+ K+++ + +E++ K + + + H+ C
Sbjct: 68 DRYVCEICNQGFQRDQNLQMHRRRHKVPWKLLKRETNEEVRKRVYVCPEPTCLHHNPCHA 127
Query: 209 CSE-----RHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYD 263
+ +HF + S IC++C K Y V + A+LK C DC
Sbjct: 128 LGDLVGIKKHFRR-KHSNHKQWICERCSKGYAV---QSDYKAHLKTCGTRGHSC--DCGR 181
Query: 264 IHMKLKNFMARGD 276
+ ++++F+ D
Sbjct: 182 VFSRVESFIEHQD 194
>At3g14740 unknown protein
Length = 341
Score = 39.3 bits (90), Expect = 0.005
Identities = 27/101 (26%), Positives = 49/101 (47%), Gaps = 11/101 (10%)
Query: 163 KDKNVEHKE--NVVTTQKIIESDPSEQI---AKICTLSVKHKEVEHSSCALCSERHFNNG 217
++K++E K NV ++ ++ E D E I K L + +EVE +C+ +G
Sbjct: 102 EEKSIEKKSTLNVESSLEVEEDDDKENIDPLGKGKALDLSDREVEDEDGIMCAVCQSTDG 161
Query: 218 E-FSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFC 257
+ +P ++ CD C+ H C + +K +P+ WFC
Sbjct: 162 DPLNP--IVFCDGCDLMVHASC---YGNPLVKAIPEGDWFC 197
>At2g01940 putative C2H2-type zinc finger protein
Length = 439
Score = 38.5 bits (88), Expect = 0.009
Identities = 28/133 (21%), Positives = 60/133 (45%), Gaps = 12/133 (9%)
Query: 150 EKWYCEYCRNKLQKDKNVE-HKENVVTTQKIIESDPSEQIAKICTLSVKHKEVEHSSCAL 208
+++ CE C Q+D+N++ H+ K+++ D + ++ K + + + H+ C
Sbjct: 65 DRYICEICNQGFQRDQNLQMHRRRHKVPWKLLKRDNNIEVKKRVYVCPEPTCLHHNPCHA 124
Query: 209 CSE-----RHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYD 263
+ +HF + S +C++C K Y V + A+LK C DC
Sbjct: 125 LGDLVGIKKHFRR-KHSNHKQWVCERCSKGYAV---QSDYKAHLKTCGTRGHSC--DCGR 178
Query: 264 IHMKLKNFMARGD 276
+ ++++F+ D
Sbjct: 179 VFSRVESFIEHQD 191
>At4g31340 unknown protein
Length = 437
Score = 37.4 bits (85), Expect = 0.021
Identities = 53/204 (25%), Positives = 77/204 (36%), Gaps = 18/204 (8%)
Query: 22 DDVEKK---MKNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSNSHASPTTINYRDQC 78
D++EK+ +KN ++KNK K + R+NE++ K S+ T +
Sbjct: 115 DELEKQVEVLKNFLEQKNKEKDSTEARTNEAEK-----KLRELNSSLDKLQKTNEEQKNK 169
Query: 79 LHKLVFQENVLEDGAAVGYFVYEEVSPSKFEAHAGWASRRKPWPLIIEFLFPIVMNTAVS 138
+ KL + E+ + EAH W PW + F F T
Sbjct: 170 IGKLERAIKIAEEEMLRTKLEATTKAKELLEAHGSWL---PPWLAVHWFKFQTYTETHWE 226
Query: 139 ANKEPVESPPKEKWYCEYCRNKLQKDKNVE-HKENVVT--TQKIIESDPSEQIAKICTLS 195
A+ +P E + K Q +K E H ENV T I E+ TLS
Sbjct: 227 AHGKPA----VETVILKVTEAKAQAEKWAEPHVENVKTKYIPAIKETVTIHVEPHFRTLS 282
Query: 196 VKHKEVEHSSCALCSERHFNNGEF 219
+K KE HSS + S EF
Sbjct: 283 IKAKEAYHSSKSAVSPHIVTVQEF 306
>At5g24330 putative protein
Length = 349
Score = 37.0 bits (84), Expect = 0.027
Identities = 15/37 (40%), Positives = 23/37 (61%), Gaps = 4/37 (10%)
Query: 221 PWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFC 257
P +++CD+C+K +H+ CL+ L VPK WFC
Sbjct: 44 PAKLLLCDKCDKGFHLFCLR----PILVSVPKGSWFC 76
>At1g25250 Indeterminate 1 like Zn finger containing protein
Length = 385
Score = 37.0 bits (84), Expect = 0.027
Identities = 29/134 (21%), Positives = 59/134 (43%), Gaps = 13/134 (9%)
Query: 150 EKWYCEYCRNKLQKDKNVE-HKENVVTTQKIIESD-PSEQIAKICTLSVKHKEVEHSSCA 207
+++ CE C Q+D+N++ H+ K+++ D E++ K + + + H C
Sbjct: 60 DRYVCEICNQGFQRDQNLQMHRRRHKVPWKLLKRDKKDEEVRKRVYVCPEPTCLHHDPCH 119
Query: 208 LCSE-----RHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCY 262
+ +HF + S +C++C K Y V + A+LK C DC
Sbjct: 120 ALGDLVGIKKHFRR-KHSVHKQWVCERCSKGYAV---QSDYKAHLKTCGSRGHSC--DCG 173
Query: 263 DIHMKLKNFMARGD 276
+ ++++F+ D
Sbjct: 174 RVFSRVESFIEHQD 187
>At1g77250 unknown protein
Length = 522
Score = 36.2 bits (82), Expect = 0.046
Identities = 19/56 (33%), Positives = 25/56 (43%), Gaps = 11/56 (19%)
Query: 206 CALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDC 261
C LC E+ + CD CE YHV C + K +P H W+C +DC
Sbjct: 243 CKLCGEKA------EARDCLACDHCEDMYHVSCAQPGG----KGMPTHSWYC-LDC 287
Score = 31.6 bits (70), Expect = 1.1
Identities = 11/34 (32%), Positives = 19/34 (55%), Gaps = 4/34 (11%)
Query: 224 VMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFC 257
+++CD C+ YH+ C++ + VP WFC
Sbjct: 417 IVLCDGCDDAYHIYCMR----PPCESVPNGEWFC 446
>At3g28770 hypothetical protein
Length = 2081
Score = 35.8 bits (81), Expect = 0.060
Identities = 23/63 (36%), Positives = 33/63 (51%), Gaps = 5/63 (7%)
Query: 6 EELKDNIVVEAVAKDGDDVEKKMKNKKKKKNKNKKESNLR---SNESKNLQPSPKHTTQT 62
EE KD I + K D +KK K + K N KKE + + +NE K + + K TT++
Sbjct: 927 EENKDTINTSSKQKGKD--KKKKKKESKNSNMKKKEEDKKEYVNNELKKQEDNKKETTKS 984
Query: 63 SNS 65
NS
Sbjct: 985 ENS 987
Score = 29.6 bits (65), Expect = 4.3
Identities = 15/49 (30%), Positives = 27/49 (54%), Gaps = 1/49 (2%)
Query: 5 HEELKDNIVVEAVAKDGDDVEKKMKNKKKKKNKNKKESNLRSNESKNLQ 53
+EE + +E+ ++V+KK K K K + KKE ++ +E K L+
Sbjct: 1159 NEEKSETKEIESSKSQKNEVDKKEK-KSSKDQQKKKEKEMKESEEKKLK 1206
Score = 29.6 bits (65), Expect = 4.3
Identities = 16/43 (37%), Positives = 25/43 (57%), Gaps = 1/43 (2%)
Query: 18 AKDGDDVEKKMKNKKKKKNKNKKESNLR-SNESKNLQPSPKHT 59
+KD VE K K K+ K+NK K + R N+ +N+Q + K +
Sbjct: 740 SKDDKSVEAKGKKKESKENKKTKTNENRVRNKEENVQGNKKES 782
Score = 29.3 bits (64), Expect = 5.6
Identities = 22/64 (34%), Positives = 34/64 (52%), Gaps = 9/64 (14%)
Query: 7 ELKDNIVVEAVAKDGDDVEKKMKNKKKKKNKNK---KESNLRSN--ESKNLQPSPKHTTQ 61
E KD+ VEA K + E NKK K N+N+ KE N++ N ES+ ++ K ++
Sbjct: 739 ESKDDKSVEAKGKKKESKE----NKKTKTNENRVRNKEENVQGNKKESEKVEKGEKKESK 794
Query: 62 TSNS 65
+ S
Sbjct: 795 DAKS 798
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,450,017
Number of Sequences: 26719
Number of extensions: 556440
Number of successful extensions: 3489
Number of sequences better than 10.0: 112
Number of HSP's better than 10.0 without gapping: 52
Number of HSP's successfully gapped in prelim test: 62
Number of HSP's that attempted gapping in prelim test: 3064
Number of HSP's gapped (non-prelim): 310
length of query: 525
length of database: 11,318,596
effective HSP length: 104
effective length of query: 421
effective length of database: 8,539,820
effective search space: 3595264220
effective search space used: 3595264220
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC126007.11


