

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126006.10 - phase: 0 /pseudo
(352 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
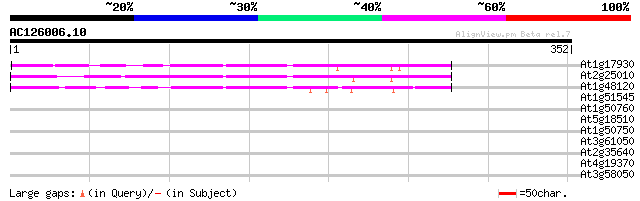
Score E
Sequences producing significant alignments: (bits) Value
At1g17930 unknown protein 104 8e-23
At2g25010 unknown protein 84 2e-16
At1g48120 serine/threonine phosphatase PP7, putative 67 2e-11
At1g51545 unknown protein 38 0.007
At1g50760 hypothetical protein 35 0.047
At5g18510 putative protein 32 0.68
At1g50750 hypothetical protein 31 0.89
At3g61050 CaLB protein 29 3.4
At2g35640 unknown protein 28 5.8
At4g19370 hypothetical protein 28 9.9
At3g58050 putative protein 28 9.9
>At1g17930 unknown protein
Length = 478
Score = 104 bits (259), Expect = 8e-23
Identities = 77/283 (27%), Positives = 133/283 (46%), Gaps = 32/283 (11%)
Query: 2 PMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYG 61
P GEMT+TLD+V+ + L ++G+ + G K + + + +R LG + G+ G
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVV-GVKEKDEDPSQVCLRLLGKLPK------GELSG 136
Query: 62 GYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRSN 121
++ L++ + A ++E+ Y R YL+Y++G +F
Sbjct: 137 NRVTAKWLKESF----------AECPKGATMKEIE----YHTRAYLIYIVGSTIFATTDP 182
Query: 122 KRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDKALGGSVTLLTIRKLIF 181
+I + YL ED + Y+WG LA+LY ++ +A + +GG +TLL +
Sbjct: 183 SKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLL---QCWS 238
Query: 182 YFFVNVDPNTDYMENYPVAARWK---LQKGHEEGIMYRSLLDRIQLDDVCWRPYEEHREI 238
YF +N+D +P+A WK + + YR LD + +V W P+E +I
Sbjct: 239 YFHLNIDRPKRTTRQFPLALLWKGRQQSRSKNDLFKYRKALDDLDPSNVSWCPFEGDLDI 298
Query: 239 --QDFED--VFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIP 277
Q F+D + S + G + V H P+R ++Q+G Q IP
Sbjct: 299 VPQSFKDNLLLGRSRTKLIGPKVVEWHFPDRCMKQFGLCQVIP 341
>At2g25010 unknown protein
Length = 509
Score = 83.6 bits (205), Expect = 2e-16
Identities = 75/283 (26%), Positives = 115/283 (40%), Gaps = 28/283 (9%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEY 60
+P+GEMT+TLD+VA + L I+G + K V + A +CG+
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIVGSK----------------VGDEVAMDMCGRLL 135
Query: 61 GGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRS 120
G S + N L T + V R YLLYLIG +F
Sbjct: 136 GKLPSAANKE--VNCSRVKLNWLKRTFSECPEDASFDVVKCHTRAYLLYLIGSTIFATTD 193
Query: 121 NKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDKALGGSVTLLTIRKLI 180
++ + YL ED + Y+WG LA LY L +A + G +TLL +
Sbjct: 194 GDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLL---QCW 249
Query: 181 FYFFVNVDPNTDYMENYPVAARWKLQKGHEEGIM--YRSLLDRIQLDDVCWRPYEEHREI 238
YF +++ +P+A WK + + + YR LD + + W PYE +
Sbjct: 250 SYFHLDIGRPEKSEACFPLALLWKGKGSRSKTDLSEYRRELDDLDPSKITWCPYERFENL 309
Query: 239 ----QDFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIP 277
+ + S + ++ H P+R LRQ+G Q IP
Sbjct: 310 IPPHIKAKLILGRSKTTLVCFEKIELHFPDRCLRQFGKRQPIP 352
>At1g48120 serine/threonine phosphatase PP7, putative
Length = 1338
Score = 66.6 bits (161), Expect = 2e-11
Identities = 82/302 (27%), Positives = 129/302 (42%), Gaps = 55/302 (18%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEY 60
+P GE+TVTL DV L L ++G ++ K + A L LG + +
Sbjct: 101 LPAGEITVTLQDVNILLGLRVDGPAVTGSTK---YNWADLCEDLLGHRPGPKDL-----H 152
Query: 61 GGYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYC-VRCYLLYLIGCLLFGDR 119
G ++S LR+ + L + D V L+ C R ++L L+ L+GD+
Sbjct: 153 GSHVSLAWLRENFRN-------LPADPDEVTLK--------CHTRAFVLALMSGFLYGDK 197
Query: 120 SNKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDKALGGSVTLLTIRKL 179
S + L +L + D + + SWG TLA LY EL A + + G + LL +L
Sbjct: 198 SKHDVALTFLPLLRD-FDEVAKLSWGSATLALLYRELCRASKRTVSTICGPLVLL---QL 253
Query: 180 IFYFFVNV-------DPNTDYMENY------PVAARWKLQKGHEEGI-----MYRSLLDR 221
+ ++V D YM+ P+ R L H+E YR D+
Sbjct: 254 WAWERLHVGRPGRLKDVGASYMDGIDGPLPDPLGCRASLS--HKENPRGGLDFYRDQFDQ 311
Query: 222 IQLDDVCWRPYEEHREIQ------DFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQT 275
+ + V W+PY + E+++ ++C V H P+RVLRQ+G QT
Sbjct: 312 QKDEQVIWQPYTPDLLAKIPLICVSGENIWRTVAPLIC-FDVVEWHRPDRVLRQFGLHQT 370
Query: 276 IP 277
IP
Sbjct: 371 IP 372
>At1g51545 unknown protein
Length = 629
Score = 38.1 bits (87), Expect = 0.007
Identities = 70/308 (22%), Positives = 113/308 (35%), Gaps = 66/308 (21%)
Query: 2 PMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYG 61
P GE T+TL+DV L ++G + A L + S ++ EK E
Sbjct: 49 PWGEATITLEDVLVLLGFSVQGSPVF----------APLESSEMRDSVEKLEK-ARLENR 97
Query: 62 GYISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRSN 121
G R + +++LGR D +E E +L + + +F D
Sbjct: 98 GQDGLVRQNLWVSSFLGRG-------DQMEHE-----------AFLAFWLSQFVFPDMCR 139
Query: 122 KRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDKALGGSVTLLTIRKLIF 181
+ I L M A ++ LA LY +L + +VTL ++ KL+
Sbjct: 140 RSISTKVL-PMAVRLARGERIAFAPAVLARLYRDLGQIQASAREKSTPNVTLKSLFKLVQ 198
Query: 182 YF----FVNVDPNTDYM-ENYPVAARWKLQKGHEEGIMYRSLLDRIQLDDVCWRPYEEHR 236
+ F + P + + P +RW Q R+ L D WRPY +
Sbjct: 199 LWAWERFKSTSPKARVIPKGEPRISRWHSQTSKNV---------RLNLVDFDWRPYTKPL 249
Query: 237 EIQD-----FEDVFWYS-------GWI----------MCGVRRVYRHLPERVLRQYGYMQ 274
+I + E+ W + G++ + GV V + P RV Q+G Q
Sbjct: 250 QIWNPPRFYPEEAMWMTVDDNLDDGFVSFARCMRVSQLVGVGIVEDYYPNRVAMQFGLAQ 309
Query: 275 TIPRHPTD 282
+P TD
Sbjct: 310 DLPGLVTD 317
>At1g50760 hypothetical protein
Length = 649
Score = 35.4 bits (80), Expect = 0.047
Identities = 26/109 (23%), Positives = 41/109 (36%), Gaps = 28/109 (25%)
Query: 198 PVAARWKLQKGHEEGIMYRSLLDRIQLDDVCWRPYEEHREIQDFEDVFWYSGWIMC---- 253
P A W K + R +LD ++D WRPY R + +++ +Y MC
Sbjct: 280 PRVAIWNKLKQQTSNV--RQILDNSKIDSFEWRPYT--RSVTNWKFPQFYPEKAMCVHVS 335
Query: 254 --------------------GVRRVYRHLPERVLRQYGYMQTIPRHPTD 282
G+ +V + P RV Q+G +Q +P D
Sbjct: 336 PSLDEELISFARCIKDSELVGIEKVEHYFPNRVASQFGMLQDVPSPHVD 384
>At5g18510 putative protein
Length = 702
Score = 31.6 bits (70), Expect = 0.68
Identities = 36/158 (22%), Positives = 63/158 (39%), Gaps = 38/158 (24%)
Query: 149 LAYLYGELADACRPGDKALGGSVTLLTIRKLIFYF----FVNVDPNT-DYMENYPVAARW 203
LA+LY +L C G V L ++ KL+ + F N+ P + + P A+W
Sbjct: 232 LAFLYKDLDRICDFSRGKCAGKVNLKSLFKLVQVWTWERFSNIRPKAKEIPKGEPRIAQW 291
Query: 204 KLQKGHEEGIMYRSLLDRIQLDDVCWRPYEEHREIQDFEDVFWY---SGWI--------- 251
+G+ S ++ D WRPY + ++++ + +Y + W+
Sbjct: 292 -------DGLQQISKNVKLSFDVFEWRPYT--KPLKNWNPLRFYVDEAMWLTVDDSVDDA 342
Query: 252 ------------MCGVRRVYRHLPERVLRQYGYMQTIP 277
+ G V + P RV RQ+G Q +P
Sbjct: 343 FASFARCVKVSYLAGNGFVEDYFPYRVARQFGLSQDLP 380
>At1g50750 hypothetical protein
Length = 816
Score = 31.2 bits (69), Expect = 0.89
Identities = 16/62 (25%), Positives = 30/62 (47%), Gaps = 1/62 (1%)
Query: 216 RSLLDRIQLDDVCWRPYEEHREIQDFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQT 275
R +LD +++D V W P + +++ + G+ V + P RV Q+G +Q
Sbjct: 258 RQILDSLKIDTV-WIPASPNLDVEFVSFARCIKVSQLVGIDNVEHYFPNRVASQFGMLQD 316
Query: 276 IP 277
+P
Sbjct: 317 VP 318
>At3g61050 CaLB protein
Length = 510
Score = 29.3 bits (64), Expect = 3.4
Identities = 16/45 (35%), Positives = 28/45 (61%), Gaps = 3/45 (6%)
Query: 24 RMLSHGKKMPKHEGAALLMRHLG-VSQQEAEKICGQEYGGYISYP 67
RM++H + K A+ M+ LG +S+ + +KICG + +IS+P
Sbjct: 23 RMMTH--RSSKRVAKAVDMKLLGSLSRDDLKKICGDNFPQWISFP 65
>At2g35640 unknown protein
Length = 340
Score = 28.5 bits (62), Expect = 5.8
Identities = 39/177 (22%), Positives = 64/177 (36%), Gaps = 26/177 (14%)
Query: 160 CRPGDKALGGSVTLLTIRKLIFYFFVNVDPNTDYMENYPVAARWKL------QKG-HEEG 212
CR G+ + ++ L+ +K+ V N P RWK ++G +
Sbjct: 17 CRKGNWTVSETLVLIEAKKMDDQRRVRRSEKQPEGRNKPAELRWKWIEEYCWRRGCYRNQ 76
Query: 213 IMYRSLLDRIQLDDVCWRPYEEHREIQDFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGY 272
D + D R YE R F V S W M R ++LP +L Q
Sbjct: 77 NQCNDKWDNLMRDYKKIREYERSRVESSFNTVTSSSYWKMDKTERKEKNLPSNMLPQ--- 133
Query: 273 MQTIPRHPTDVVELPLPYIVQAFVDFRTHTLKAAIGVSRQESRHGGWLMAICYGTLG 329
+ + D LP + + AA+G + +GG ++ +C +LG
Sbjct: 134 IYDVLSELVDRKTLP-----------SSSSAAAAVG-----NGNGGQILRVCQQSLG 174
>At4g19370 hypothetical protein
Length = 217
Score = 27.7 bits (60), Expect = 9.9
Identities = 24/84 (28%), Positives = 35/84 (41%), Gaps = 12/84 (14%)
Query: 258 VYRHLPERVLRQYGYMQTIPRHPTDVVELPLPYIVQAFVDFRTHTLKAAIGVSRQESRHG 317
V+R+ R R+ GY T PT ++ L V + L AI +SR ++
Sbjct: 71 VFRNHRTRTKREDGYKITDLTLPTVLLLLSWSNFVVVVL-----ILSTAISMSRAQAYGE 125
Query: 318 GWLMAICY-------GTLGCLTLI 334
GWL CY GCL ++
Sbjct: 126 GWLDEDCYLVKDGVFAASGCLAIL 149
>At3g58050 putative protein
Length = 1209
Score = 27.7 bits (60), Expect = 9.9
Identities = 18/51 (35%), Positives = 24/51 (46%)
Query: 64 ISYPRLRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCL 114
+SY +L+ F++ +A D L E AR YC RC L L G L
Sbjct: 24 VSYNQLQKFWSELSPKARQELLKIDKQTLFEQARKNMYCSRCNGLLLEGFL 74
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.143 0.453
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,082,859
Number of Sequences: 26719
Number of extensions: 343883
Number of successful extensions: 853
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 829
Number of HSP's gapped (non-prelim): 15
length of query: 352
length of database: 11,318,596
effective HSP length: 100
effective length of query: 252
effective length of database: 8,646,696
effective search space: 2178967392
effective search space used: 2178967392
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC126006.10


