

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125480.3 + phase: 0
(476 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
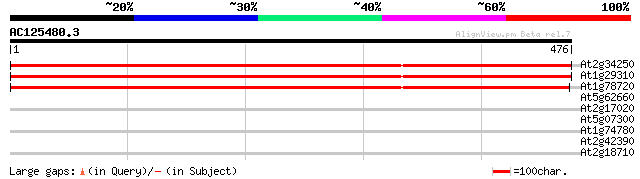
Score E
Sequences producing significant alignments: (bits) Value
At2g34250 putative protein transport protein SEC61 alpha subunit 880 0.0
At1g29310 transport protein sec61 alpha subunit, putative 873 0.0
At1g78720 hypothetical protein 861 0.0
At5g62660 putative protein 33 0.26
At2g17020 unknown protein 30 2.2
At5g07300 copine-like protein 30 3.8
At1g74780 unknown protein (At1g74780) 29 6.5
At2g42390 unknown protein 28 8.5
At2g18710 putative preprotein translocase SECY protein 28 8.5
>At2g34250 putative protein transport protein SEC61 alpha subunit
Length = 475
Score = 880 bits (2273), Expect = 0.0
Identities = 445/476 (93%), Positives = 467/476 (97%), Gaps = 1/476 (0%)
Query: 1 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 60
MGGGFRVLHLVRPFL+FLPEVQ+ADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG
Sbjct: 1 MGGGFRVLHLVRPFLAFLPEVQSADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 60
Query: 61 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 120
ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK
Sbjct: 61 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 120
Query: 121 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGS 180
LLGILIA+GEAVAYVLSGMYG VGQLGVGNAILII+QLFFAGIIVICLDELLQKGYGLGS
Sbjct: 121 LLGILIAIGEAVAYVLSGMYGPVGQLGVGNAILIILQLFFAGIIVICLDELLQKGYGLGS 180
Query: 181 GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVRALREAFYRQ 240
GISLFIATNICE+IIWKAFSPTTIN+GRGAEFEGAVIALFH+LIT+++KV ALR+AFYRQ
Sbjct: 181 GISLFIATNICESIIWKAFSPTTINTGRGAEFEGAVIALFHMLITKSNKVAALRQAFYRQ 240
Query: 241 NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL 300
NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL
Sbjct: 241 NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL 300
Query: 301 VSNLYFISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAPSSLADMAANP 360
VSNLYFISQLL+RK+SGNF VNLLG+WKESEY G SIPV G+AY ITAP+S +DMAA+P
Sbjct: 301 VSNLYFISQLLYRKFSGNFFVNLLGQWKESEY-SGQSIPVSGLAYLITAPASFSDMAAHP 359
Query: 361 FHALFYLVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP 420
FHALFY+VFML+ACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP
Sbjct: 360 FHALFYIVFMLTACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP 419
Query: 421 TAAAFGGICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF 476
TAAAFGG+CIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKE+ASELGFFGF
Sbjct: 420 TAAAFGGVCIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKEKASELGFFGF 475
>At1g29310 transport protein sec61 alpha subunit, putative
Length = 475
Score = 873 bits (2256), Expect = 0.0
Identities = 442/476 (92%), Positives = 465/476 (96%), Gaps = 1/476 (0%)
Query: 1 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 60
MGGGFRVLHLVRPFL+FLPEVQ+ADRK+PFREKVIYTVISLFIFLVCSQLPLYGIHSTTG
Sbjct: 1 MGGGFRVLHLVRPFLAFLPEVQSADRKIPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 60
Query: 61 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 120
ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK
Sbjct: 61 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 120
Query: 121 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGS 180
LL IL A+GEAVAYVLSGMYG VGQLGVGNAILII+QLFFAGIIVICLDELLQKGYGLGS
Sbjct: 121 LLWILSAIGEAVAYVLSGMYGPVGQLGVGNAILIILQLFFAGIIVICLDELLQKGYGLGS 180
Query: 181 GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVRALREAFYRQ 240
GISLFIATNICE+IIWKAFSPTTIN+GRGAEFEGAVIALFH+LIT+++KV ALR+AFYRQ
Sbjct: 181 GISLFIATNICESIIWKAFSPTTINTGRGAEFEGAVIALFHMLITKSNKVAALRQAFYRQ 240
Query: 241 NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL 300
NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSK+ARGQQGSYPIKLFYTSNMPIILQSAL
Sbjct: 241 NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKSARGQQGSYPIKLFYTSNMPIILQSAL 300
Query: 301 VSNLYFISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAPSSLADMAANP 360
VSNLYFISQLL+RK+SGNF VNLLG+WKESEY G SIPV G+AY ITAP+S ADMAA+P
Sbjct: 301 VSNLYFISQLLYRKFSGNFFVNLLGQWKESEY-SGQSIPVSGLAYLITAPASFADMAAHP 359
Query: 361 FHALFYLVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP 420
FHALFY+VFML+ACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP
Sbjct: 360 FHALFYIVFMLTACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP 419
Query: 421 TAAAFGGICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF 476
TAAAFGG+CIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKE+ASELGFFGF
Sbjct: 420 TAAAFGGVCIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKEKASELGFFGF 475
>At1g78720 hypothetical protein
Length = 475
Score = 861 bits (2224), Expect = 0.0
Identities = 433/475 (91%), Positives = 463/475 (97%), Gaps = 1/475 (0%)
Query: 1 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 60
M GGFRV+HLVRPFL+FLPEVQ+ +RK+PFREKVIYTVISLFIFLVCSQLPLYGIHSTTG
Sbjct: 1 MVGGFRVIHLVRPFLAFLPEVQSPERKIPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 60
Query: 61 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 120
ADPFYWMRVILAS+RGTVMELGITPIVTSG+VMQLLAGSKIIE+DNNVREDRALLNGAQK
Sbjct: 61 ADPFYWMRVILASSRGTVMELGITPIVTSGMVMQLLAGSKIIEIDNNVREDRALLNGAQK 120
Query: 121 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGS 180
LLGILIAVG+AVAYVLSGMYGSVG+LGVGNAILIIVQL FA IIV+CLDELLQKGYGLGS
Sbjct: 121 LLGILIAVGQAVAYVLSGMYGSVGELGVGNAILIIVQLCFAAIIVLCLDELLQKGYGLGS 180
Query: 181 GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVRALREAFYRQ 240
GISLFIATNICE+IIWKAFSPTTINSGRGA+FEGAVIALFHLLITRTDKVRALREAF+RQ
Sbjct: 181 GISLFIATNICESIIWKAFSPTTINSGRGAQFEGAVIALFHLLITRTDKVRALREAFFRQ 240
Query: 241 NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL 300
NLPNVTNL ATVLIFLIVIYFQGFRVVLPVRSKNARGQ+GSYPIKLFYTSNMPIILQSAL
Sbjct: 241 NLPNVTNLHATVLIFLIVIYFQGFRVVLPVRSKNARGQRGSYPIKLFYTSNMPIILQSAL 300
Query: 301 VSNLYFISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAPSSLADMAANP 360
VSN+YFISQ+L+RK+ GNF+VNL+G WKESEY G SIPVGGIAYYITAPSSLA+MA +P
Sbjct: 301 VSNIYFISQILYRKFGGNFLVNLIGTWKESEY-SGQSIPVGGIAYYITAPSSLAEMATHP 359
Query: 361 FHALFYLVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP 420
FHALFYLVFML+ACALFSKTWIEVSGSSA+DVA+QL+EQQMVMPGHR+SNLQKELNRYIP
Sbjct: 360 FHALFYLVFMLAACALFSKTWIEVSGSSAKDVARQLREQQMVMPGHRDSNLQKELNRYIP 419
Query: 421 TAAAFGGICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFG 475
TAAAFGG+CIGALTVLAD MGAIGSGTGILLAVTIIYQYFETFEKE+ASELGFFG
Sbjct: 420 TAAAFGGLCIGALTVLADLMGAIGSGTGILLAVTIIYQYFETFEKEKASELGFFG 474
>At5g62660 putative protein
Length = 379
Score = 33.5 bits (75), Expect = 0.26
Identities = 24/101 (23%), Positives = 43/101 (41%), Gaps = 7/101 (6%)
Query: 266 VVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSALVSNLYFISQLLHRKYSGN--FIVNL 323
V LP N Q+ + L++ P++ Q +V + S+ L R S + F++
Sbjct: 167 VTLPAVKSNILAQKSHWNSLLYFFGYDPVLDQYKVVCTVALFSKRLKRITSEHWVFVLEP 226
Query: 324 LGKWKESEYGGGH-----SIPVGGIAYYITAPSSLADMAAN 359
G WK E+ H + V G+ YY+ + D+ +
Sbjct: 227 GGSWKRIEFDQPHLPTRLGLCVNGVIYYLASTWKRRDIVVS 267
>At2g17020 unknown protein
Length = 656
Score = 30.4 bits (67), Expect = 2.2
Identities = 25/83 (30%), Positives = 42/83 (50%), Gaps = 3/83 (3%)
Query: 71 LASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQKLLGILIAVGE 130
L S+R T +ELG +TS ++ QLL ++I D+N +L Q+L + + +
Sbjct: 192 LVSDRLTHLELGH---ITSRMMTQLLTSTEISGQDSNRVTTSTVLQNVQRLRLSVDCITD 248
Query: 131 AVAYVLSGMYGSVGQLGVGNAIL 153
AV +S S+ L + +A L
Sbjct: 249 AVVKAISKSLPSLIDLDIRDAPL 271
>At5g07300 copine-like protein
Length = 586
Score = 29.6 bits (65), Expect = 3.8
Identities = 23/77 (29%), Positives = 31/77 (39%), Gaps = 9/77 (11%)
Query: 327 WKESEYGGGHSIPVGGIAYYITAPSSLADMAANPFHALFYLVFMLSACALFSKTWIEVSG 386
W + Y GG + VGG A AA P A+ Y + LFS+ + S
Sbjct: 5 WSDGSYAGGGMVGVGGGA---------NSSAATPNDAVDYYLKSRGYNGLFSQIELSFSA 55
Query: 387 SSARDVAKQLKEQQMVM 403
S+ RD K MV+
Sbjct: 56 SNLRDRDVISKSDAMVV 72
>At1g74780 unknown protein (At1g74780)
Length = 533
Score = 28.9 bits (63), Expect = 6.5
Identities = 22/81 (27%), Positives = 40/81 (49%), Gaps = 6/81 (7%)
Query: 78 VMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQKLLGILIAVGEAVAYVLS 137
++ L +TP V S LVM L+ +I E +V +D+ LNG + ++IA + +L
Sbjct: 185 ILLLAVTPTVLSLLVMPLV---RIYE--TSVADDKKHLNGL-SAVSLIIAAYLMIIIILK 238
Query: 138 GMYGSVGQLGVGNAILIIVQL 158
+G + + ++V L
Sbjct: 239 NTFGLSSWANIVTLVCLLVML 259
>At2g42390 unknown protein
Length = 212
Score = 28.5 bits (62), Expect = 8.5
Identities = 19/55 (34%), Positives = 29/55 (52%), Gaps = 5/55 (9%)
Query: 245 VTNLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSA 299
V NL L+F + + F GF V+L + AR ++ Y +K N+P I +SA
Sbjct: 162 VKNLQGMKLVFALQMVFIGFLVILWMLRSRARSKRRRYLLK-----NVPPINRSA 211
>At2g18710 putative preprotein translocase SECY protein
Length = 551
Score = 28.5 bits (62), Expect = 8.5
Identities = 36/154 (23%), Positives = 65/154 (41%), Gaps = 12/154 (7%)
Query: 78 VMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQKLLGI--LIAVGEAVAYV 135
+ LGI P + + +V QLLA D +E A G +K+L +VG A+
Sbjct: 194 ICSLGIVPFINAQIVFQLLAQVYPKLQDLQKKEGEA---GRKKILQYTRYASVGFAIVQA 250
Query: 136 LSGMY---GSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGSGISLFIATNICE 192
+ ++ V + + L ++ + E + LG+G SL I T+I
Sbjct: 251 IGQVFYLRPYVNDFSTEWVVSSVTLLTLGSVLTTYIGERI-SDLKLGNGTSLLIFTSII- 308
Query: 193 NIIWKAFSPTTINSGRGAEFE--GAVIALFHLLI 224
+ + +F TT + + + G ++ F LL+
Sbjct: 309 SYLPASFGRTTAEALQEGNYTGLGTIVVSFLLLV 342
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.143 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,915,361
Number of Sequences: 26719
Number of extensions: 410202
Number of successful extensions: 1201
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1194
Number of HSP's gapped (non-prelim): 9
length of query: 476
length of database: 11,318,596
effective HSP length: 103
effective length of query: 373
effective length of database: 8,566,539
effective search space: 3195319047
effective search space used: 3195319047
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC125480.3


