

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125480.1 - phase: 0 /pseudo
(250 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
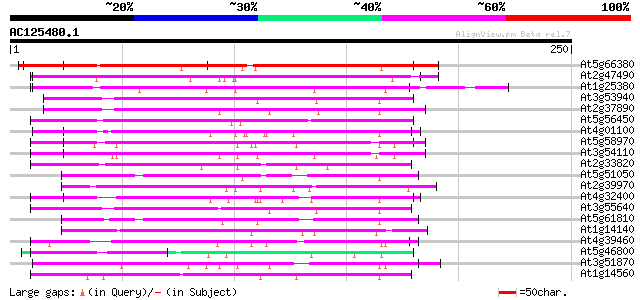
Score E
Sequences producing significant alignments: (bits) Value
At5g66380 unknown protein 207 4e-54
At2g47490 mitochondrial carrier like protein 89 2e-18
At1g25380 unknown protein 88 4e-18
At3g53940 unknown protein 78 4e-15
At2g37890 mitochondrial carrier like protein 77 7e-15
At5g56450 ADP/ATP translocase-like protein 71 6e-13
At4g01100 putative carrier protein 68 5e-12
At5g58970 uncoupling protein AtUCP2 67 1e-11
At3g54110 uncoupling protein (ucp/PUMP) 66 2e-11
At2g33820 mitochondrial basic amino acid carrier (BAC1) 63 2e-10
At5g51050 calcium-binding transporter-like protein 62 3e-10
At2g39970 36kDa-peroxisomal membrane protein (PMP36) 61 5e-10
At4g32400 adenylate translocator (brittle-1) - like protein 60 1e-09
At3g55640 Ca-dependent solute carrier - like protein 58 5e-09
At5g61810 peroxisomal Ca-dependent solute carrier - like protein 57 1e-08
At1g14140 mitochondrial uncoupling protein - like 57 1e-08
At4g39460 mitochondrial carrier like protein 55 4e-08
At5g46800 carnitine/acylcarnitine translocase-like protein 53 1e-07
At3g51870 carrier like protein 53 1e-07
At1g14560 mitochondrial carrier like protein 53 1e-07
>At5g66380 unknown protein
Length = 308
Score = 207 bits (527), Expect = 4e-54
Identities = 108/200 (54%), Positives = 134/200 (67%), Gaps = 16/200 (8%)
Query: 5 QWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLK 64
QWE A GFA + + LDVV+TR QV+DGR S P Y NT HA+FTIAR EGL+
Sbjct: 6 QWENATAGAVAGFATVAAMHSLDVVRTRFQVNDGR-GSSLPTYKNTAHAVFTIARLEGLR 64
Query: 65 GLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-------------SGALMTVCL 111
GLYAGF ++G+T+SW L F+YG K R+AR + ++ +GAL VCL
Sbjct: 65 GLYAGFFPAVIGSTVSWGLYFFFYGRAKQRYARGRDDEKLSPALHLASAAEAGAL--VCL 122
Query: 112 CANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQ 171
C NP ++VKTRLQLQTPLH +PYSGL DAFRTI +EEG A Y+GIVPG L+S AIQ
Sbjct: 123 CTNPIWLVKTRLQLQTPLHQTQPYSGLLDAFRTIVKEEGPRALYKGIVPGLVLVSHGAIQ 182
Query: 172 FIVYEQLRKTVVNLKTKGSK 191
F YE+LRK +V+LK + K
Sbjct: 183 FTAYEELRKIIVDLKERRRK 202
Score = 47.8 bits (112), Expect = 6e-06
Identities = 48/175 (27%), Positives = 70/175 (39%), Gaps = 21/175 (12%)
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLL----GATIS 80
P+ +V+TRLQ+ Q Y+ A TI ++EG + LY G GL+ GA
Sbjct: 126 PIWLVKTRLQLQTP--LHQTQPYSGLLDAFRTIVKEEGPRALYKGIVPGLVLVSHGAIQF 183
Query: 81 WSLLVFYYGIVKDRHARSKVEKSGALMT--------------VCLCANPAFVVKTRLQLQ 126
+ IV + R K E + L+ L P V++ RLQ +
Sbjct: 184 TAYEELRKIIVDLKERRRKSESTDNLLNSADYAALGGSSKVAAVLLTYPFQVIRARLQQR 243
Query: 127 TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFL-ISQAAIQFIVYEQLRK 180
+ Y R R EG FYRG+ + ++I FIVYE + K
Sbjct: 244 PSTNGIPRYIDSLHVIRETARYEGLRGFYRGLTANLLKNVPASSITFIVYENVLK 298
Score = 43.1 bits (100), Expect = 1e-04
Identities = 26/82 (31%), Positives = 38/82 (45%), Gaps = 2/82 (2%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGL 66
+Y ++ A + YP V++ RLQ + PRY ++ H I AR EGL+G
Sbjct: 214 DYAALGGSSKVAAVLLTYPFQVIRARLQQRPST--NGIPRYIDSLHVIRETARYEGLRGF 271
Query: 67 YAGFPAGLLGATISWSLLVFYY 88
Y G A LL + S+ Y
Sbjct: 272 YRGLTANLLKNVPASSITFIVY 293
>At2g47490 mitochondrial carrier like protein
Length = 312
Score = 89.0 bits (219), Expect = 2e-18
Identities = 64/191 (33%), Positives = 89/191 (46%), Gaps = 13/191 (6%)
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHD-GRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
A G TF PLDV++TR QVH +L + + ++ I ++EG++GLY G
Sbjct: 19 AGAAAGVVAATFVCPLDVIKTRFQVHGLPKLGDANIKGSLIVGSLEQIFKREGMRGLYRG 78
Query: 70 FPAGLLGATISWSLLVFYYGIVK-----DRHARSK----VEKSGALMTVCLCANPAFVVK 120
++ +W++ Y +K + H S + SGA + NP +VVK
Sbjct: 79 LSPTVMALLSNWAIYFTMYDQLKSFLCSNDHKLSVGANVLAASGAGAATTIATNPLWVVK 138
Query: 121 TRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIVYEQLRK 180
TRLQ Q PY + A R I EEG Y G+VP IS AIQF YE ++
Sbjct: 139 TRLQTQGMRVGIVPYKSTFSALRRIAYEEGIRGLYSGLVPALAGISHVAIQFPTYEMIK- 197
Query: 181 TVVNLKTKGSK 191
V L KG K
Sbjct: 198 --VYLAKKGDK 206
Score = 59.7 bits (143), Expect = 1e-09
Identities = 56/187 (29%), Positives = 86/187 (45%), Gaps = 16/187 (8%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A+ G A PL VV+TRLQ R+ Y +T A+ IA +EG++GLY+G
Sbjct: 118 LAASGAGAATTIATNPLWVVKTRLQTQGMRV--GIVPYKSTFSALRRIAYEEGIRGLYSG 175
Query: 70 FPAGLLGATI------SWSLLVFYYGIVKDR-----HARS-KVEKSGALMTVCLCANPAF 117
L G + ++ ++ Y D+ +AR V S A + P
Sbjct: 176 LVPALAGISHVAIQFPTYEMIKVYLAKKGDKSVDNLNARDVAVASSIAKIFASTLTYPHE 235
Query: 118 VVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA-IQFIVYE 176
VV+ RLQ Q H + YSG+ D + + ++GF FYRG + AA I F +E
Sbjct: 236 VVRARLQEQGH-HSEKRYSGVRDCIKKVFEKDGFPGFYRGCATNLLRTTPAAVITFTSFE 294
Query: 177 QLRKTVV 183
+ + +V
Sbjct: 295 MVHRFLV 301
>At1g25380 unknown protein
Length = 363
Score = 88.2 bits (217), Expect = 4e-18
Identities = 65/225 (28%), Positives = 98/225 (42%), Gaps = 22/225 (9%)
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFT----IARKEGLKGL 66
A T G TF PLDV++TRLQV + P I T I ++EG +G+
Sbjct: 23 AGATAGAIAATFVCPLDVIKTRLQVLG---LPEAPASGQRGGVIITSLKNIIKEEGYRGM 79
Query: 67 YAGFPAGLLGATISWSLLVFYYGIVKDRHARSK---------VEKSGALMTVCLCANPAF 117
Y G ++ +W++ YG +KD S + +GA + NP +
Sbjct: 80 YRGLSPTIIALLPNWAVYFSVYGKLKDVLQSSDGKLSIGSNMIAAAGAGAATSIATNPLW 139
Query: 118 VVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIVYEQ 177
VVKTRL Q PY + AF I EEG Y GI+P +S AIQF YE+
Sbjct: 140 VVKTRLMTQGIRPGVVPYKSVMSAFSRICHEEGVRGLYSGILPSLAGVSHVAIQFPAYEK 199
Query: 178 LRKTVVNLKTKGSKIQHQKPDQILVCYWTLDLFTSEISLVLDYLH 222
+++ + K + +++ P + + I+ +L Y H
Sbjct: 200 IKQYMA--KMDNTSVENLSPGNVAIA----SSIAKVIASILTYPH 238
Score = 48.9 bits (115), Expect = 3e-06
Identities = 50/185 (27%), Positives = 74/185 (39%), Gaps = 20/185 (10%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A+ G A PL VV+TRL R Y + A I +EG++GLY+G
Sbjct: 122 IAAAGAGAATSIATNPLWVVKTRLMTQGIR--PGVVPYKSVMSAFSRICHEEGVRGLYSG 179
Query: 70 FPAGLLGATISWSLLVF-YYGIVKDRHARSK-------------VEKSGALMTVCLCANP 115
L G +S + F Y +K A+ + S A + + P
Sbjct: 180 ILPSLAG--VSHVAIQFPAYEKIKQYMAKMDNTSVENLSPGNVAIASSIAKVIASILTYP 237
Query: 116 AFVVKTRLQLQTPLHHARP-YSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS-QAAIQFI 173
V++ +LQ Q + +A YSG+ D + R EG YRG + A I F
Sbjct: 238 HEVIRAKLQEQGQIRNAETKYSGVIDCITKVFRSEGIPGLYRGCATNLLRTTPSAVITFT 297
Query: 174 VYEQL 178
YE +
Sbjct: 298 TYEMM 302
>At3g53940 unknown protein
Length = 365
Score = 78.2 bits (191), Expect = 4e-15
Identities = 54/177 (30%), Positives = 85/177 (47%), Gaps = 17/177 (9%)
Query: 16 GFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLL 75
G + YPLD+V+TRL ++ Y HA TI R+EG+ GLY G A LL
Sbjct: 187 GLTAASATYPLDLVRTRLSAQRNSIY-----YQGVGHAFRTICREEGILGLYKGLGATLL 241
Query: 76 GATISWSLLVFYYGIVKDRHARSKVEKSGALMTV----------CLCANPAFVVKTRLQL 125
G S ++ Y K + S A++++ P +V+ R+QL
Sbjct: 242 GVGPSLAISFAAYETFKTFWLSHRPNDSNAVVSLGCGSLSGIVSSTATFPLDLVRRRMQL 301
Query: 126 QTPLHHARPY-SGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLRK 180
+ AR Y +GL+ F+ I + EG YRGI+P ++ ++ I F+ +E+L+K
Sbjct: 302 EGAGGRARVYTTGLFGTFKHIFKTEGMRGLYRGIIPEYYKVVPGVGIAFMTFEELKK 358
Score = 27.3 bits (59), Expect = 7.9
Identities = 15/42 (35%), Positives = 25/42 (58%), Gaps = 2/42 (4%)
Query: 139 YDAFRTIKREEGFSAFYRG-IVPGFFLISQAAIQFIVYEQLR 179
++A R +K EEGF AF++G +V + A+ F YE+ +
Sbjct: 116 HEASRIVK-EEGFRAFWKGNLVTVAHRLPYGAVNFYAYEEYK 156
>At2g37890 mitochondrial carrier like protein
Length = 337
Score = 77.4 bits (189), Expect = 7e-15
Identities = 57/182 (31%), Positives = 86/182 (46%), Gaps = 17/182 (9%)
Query: 16 GFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLL 75
G T YPLD+V+TRL ++ Y H TI R+EG+ GLY G A LL
Sbjct: 159 GITAATATYPLDLVRTRLAAQRNAIY-----YQGIEHTFRTICREEGILGLYKGLGATLL 213
Query: 76 GATISWSLLVFYYGIVK---------DRHARSKVEKSGALMTVCLCAN-PAFVVKTRLQL 125
G S ++ Y +K D + G V A P +V+ R+Q+
Sbjct: 214 GVGPSLAINFAAYESMKLFWHSHRPNDSDLVVSLVSGGLAGAVSSTATYPLDLVRRRMQV 273
Query: 126 QTPLHHARPYS-GLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLRKTVV 183
+ AR Y+ GL+ F+ I + EGF YRGI+P ++ ++ I F+ Y+ LR+ +
Sbjct: 274 EGAGGRARVYNTGLFGTFKHIFKSEGFKGIYRGILPEYYKVVPGVGIVFMTYDALRRLLT 333
Query: 184 NL 185
+L
Sbjct: 334 SL 335
>At5g56450 ADP/ATP translocase-like protein
Length = 330
Score = 70.9 bits (172), Expect = 6e-13
Identities = 47/181 (25%), Positives = 84/181 (45%), Gaps = 13/181 (7%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A G + YPLD+ TRL G+ + ++ H + TI +K+G++G+Y G
Sbjct: 146 MAGSAAGCTALIVVYPLDIAHTRLAADIGK--PEARQFRGIHHFLSTIHKKDGVRGIYRG 203
Query: 70 FPAGLLGATISWSLLVFYYGIVKDRHAR-SKVE---------KSGALMTVCLCANPAFVV 119
PA L G I L + VK+ + +K E + L + P V
Sbjct: 204 LPASLHGVIIHRGLYFGGFDTVKEIFSEDTKPELALWKRWGLAQAVTTSAGLASYPLDTV 263
Query: 120 KTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIVYEQLR 179
+ R+ +Q+ + H Y D ++ I R EG ++FYRG + F + +A + Y++++
Sbjct: 264 RRRIMMQSGMEHPM-YRSTLDCWKKIYRSEGLASFYRGALSNMFRSTGSAAILVFYDEVK 322
Query: 180 K 180
+
Sbjct: 323 R 323
>At4g01100 putative carrier protein
Length = 352
Score = 67.8 bits (164), Expect = 5e-12
Identities = 56/198 (28%), Positives = 89/198 (44%), Gaps = 32/198 (16%)
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGF 70
A T G ++ YP+D+V+ RL V S Y +Y HA+ T+ R+EG + LY G+
Sbjct: 147 AGATAGIIAMSATYPMDMVRGRLTVQTAN--SPY-QYRGIAHALATVLREEGPRALYRGW 203
Query: 71 PAGLLGATISWSLLVFYYGIVKDRHARSK----VEKSG-ALMTVCLC-----------AN 114
++G L Y +KD + VE + ++T C A
Sbjct: 204 LPSVIGVVPYVGLNFSVYESLKDWLVKENPYGLVENNELTVVTRLTCGAIAGTVGQTIAY 263
Query: 115 PAFVVKTRLQL------------QTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGF 162
P V++ R+Q+ + + Y+G+ DAFR R EGF A Y+G+VP
Sbjct: 264 PLDVIRRRMQMVGWKDASAIVTGEGRSTASLEYTGMVDAFRKTVRHEGFGALYKGLVPNS 323
Query: 163 F-LISQAAIQFIVYEQLR 179
++ AI F+ YE ++
Sbjct: 324 VKVVPSIAIAFVTYEMVK 341
Score = 47.8 bits (112), Expect = 6e-06
Identities = 50/177 (28%), Positives = 78/177 (43%), Gaps = 23/177 (12%)
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLL 84
PL+ ++ LQV + +Y+ T + I R EGL+GL+ G + ++
Sbjct: 58 PLERMKILLQVQNPHNI----KYSGTVQGLKHIWRTEGLRGLFKGNGTNCARIVPNSAVK 113
Query: 85 VFYY-----GIVKDRHARSKVEKS--------GALMTVCLCA----NPAFVVKTRLQLQT 127
F Y GI+ R+ E + GA T + A P +V+ RL +QT
Sbjct: 114 FFSYEQASNGILYMYRQRTGNENAQLTPLLRLGAGATAGIIAMSATYPMDMVRGRLTVQT 173
Query: 128 PLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLRKTVV 183
+ Y G+ A T+ REEG A YRG +P ++ + F VYE L+ +V
Sbjct: 174 -ANSPYQYRGIAHALATVLREEGPRALYRGWLPSVIGVVPYVGLNFSVYESLKDWLV 229
Score = 29.6 bits (65), Expect = 1.6
Identities = 23/84 (27%), Positives = 33/84 (38%), Gaps = 12/84 (14%)
Query: 21 TFQYPLDVVQTRLQV-----------HDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
T YPLDV++ R+Q+ +GR + Y A R EG LY G
Sbjct: 260 TIAYPLDVIRRRMQMVGWKDASAIVTGEGRSTASL-EYTGMVDAFRKTVRHEGFGALYKG 318
Query: 70 FPAGLLGATISWSLLVFYYGIVKD 93
+ S ++ Y +VKD
Sbjct: 319 LVPNSVKVVPSIAIAFVTYEMVKD 342
>At5g58970 uncoupling protein AtUCP2
Length = 305
Score = 66.6 bits (161), Expect = 1e-11
Identities = 59/184 (32%), Positives = 88/184 (47%), Gaps = 29/184 (15%)
Query: 25 PLDVVQTRLQVH------DGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGAT 78
PLD + RLQ+ DG P+Y + + TIAR+EG+ GL+ G AGL
Sbjct: 32 PLDTAKVRLQLQRKIPTGDGE---NLPKYRGSIGTLATIAREEGISGLWKGVIAGLHRQC 88
Query: 79 ISWSLLVFYYGIVKDRHARSKV--------EKSGALMT---VCLCANPAFVVKTRLQLQ- 126
I L + Y VK S + AL+T + ANP +VK RLQ +
Sbjct: 89 IYGGLRIGLYEPVKTLLVGSDFIGDIPLYQKILAALLTGAIAIIVANPTDLVKVRLQSEG 148
Query: 127 -TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAI----QFIVYEQLRKT 181
P R Y+G DA+ TI + EG SA + G+ P I++ AI + Y+Q+++T
Sbjct: 149 KLPAGVPRRYAGAVDAYFTIVKLEGVSALWTGLGPN---IARNAIVNAAELASYDQIKET 205
Query: 182 VVNL 185
++ +
Sbjct: 206 IMKI 209
Score = 59.7 bits (143), Expect = 1e-09
Identities = 53/184 (28%), Positives = 82/184 (43%), Gaps = 21/184 (11%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR-YNNTTHAIFTIARKEGLKGLYA 68
+A+L TG I P D+V+ RLQ +G+L + PR Y A FTI + EG+ L+
Sbjct: 121 LAALLTGAIAIIVANPTDLVKVRLQ-SEGKLPAGVPRRYAGAVDAYFTIVKLEGVSALWT 179
Query: 69 GFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGAL-----------MTVCLCANPAF 117
G + I + + Y +K+ + + L VC+ +P
Sbjct: 180 GLGPNIARNAIVNAAELASYDQIKETIMKIPFFRDSVLTHLLAGLAAGFFAVCI-GSPID 238
Query: 118 VVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYE 176
VVK+R+ + Y D F + EG AFY+G +P F L + AI F+ E
Sbjct: 239 VVKSRMMGDST------YRNTVDCFIKTMKTEGIMAFYKGFLPNFTRLGTWNAIMFLTLE 292
Query: 177 QLRK 180
Q++K
Sbjct: 293 QVKK 296
Score = 36.6 bits (83), Expect = 0.013
Identities = 20/56 (35%), Positives = 26/56 (45%), Gaps = 5/56 (8%)
Query: 111 LCANPAFVVKTRLQLQTPL-----HHARPYSGLYDAFRTIKREEGFSAFYRGIVPG 161
LC P K RLQLQ + + Y G TI REEG S ++G++ G
Sbjct: 28 LCTIPLDTAKVRLQLQRKIPTGDGENLPKYRGSIGTLATIAREEGISGLWKGVIAG 83
>At3g54110 uncoupling protein (ucp/PUMP)
Length = 306
Score = 65.9 bits (159), Expect = 2e-11
Identities = 54/180 (30%), Positives = 86/180 (47%), Gaps = 22/180 (12%)
Query: 25 PLDVVQTRLQVHDGRLFSQY--PRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWS 82
PLD + RLQ+ L P+Y + TIAR+EGL+ L+ G GL +
Sbjct: 31 PLDTAKVRLQLQKSALAGDVTLPKYRGLLGTVGTIAREEGLRSLWKGVVPGLHRQCLFGG 90
Query: 83 LLVFYYGIVKDRHARSKV-------EKSGALMTV----CLCANPAFVVKTRLQLQTPLHH 131
L + Y VK+ + +K A +T + ANP +VK RLQ + L
Sbjct: 91 LRIGMYEPVKNLYVGKDFVGDVPLSKKILAGLTTGALGIMVANPTDLVKVRLQAEGKLAA 150
Query: 132 ARP--YSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAI----QFIVYEQLRKTVVNL 185
P YSG +A+ TI R+EG A + G+ P +++ AI + Y+Q+++T++ +
Sbjct: 151 GAPRRYSGALNAYSTIVRQEGVRALWTGLGPN---VARNAIINAAELASYDQVKETILKI 207
Score = 63.5 bits (153), Expect = 1e-10
Identities = 57/188 (30%), Positives = 84/188 (44%), Gaps = 18/188 (9%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR-YNNTTHAIFTIARKEGLKGLYA 68
+A LTTG I P D+V+ RLQ +G+L + PR Y+ +A TI R+EG++ L+
Sbjct: 119 LAGLTTGALGIMVANPTDLVKVRLQA-EGKLAAGAPRRYSGALNAYSTIVRQEGVRALWT 177
Query: 69 GFPAGLLGATISWSLLVFYYGIVK----------DRHARSKVEKSGALMTVCLCANPAFV 118
G + I + + Y VK D + GA +P V
Sbjct: 178 GLGPNVARNAIINAAELASYDQVKETILKIPGFTDNVVTHILSGLGAGFFAVCIGSPVDV 237
Query: 119 VKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGF-FLISQAAIQFIVYEQ 177
VK+R+ + + Y G D F + +G AFY+G +P F L S I F+ EQ
Sbjct: 238 VKSRM-----MGDSGAYKGTIDCFVKTLKSDGPMAFYKGFIPNFGRLGSWNVIMFLTLEQ 292
Query: 178 LRKTVVNL 185
+K V L
Sbjct: 293 AKKYVREL 300
Score = 36.2 bits (82), Expect = 0.017
Identities = 21/55 (38%), Positives = 26/55 (47%), Gaps = 4/55 (7%)
Query: 111 LCANPAFVVKTRLQLQTPLHHAR----PYSGLYDAFRTIKREEGFSAFYRGIVPG 161
+C P K RLQLQ Y GL TI REEG + ++G+VPG
Sbjct: 27 VCTIPLDTAKVRLQLQKSALAGDVTLPKYRGLLGTVGTIAREEGLRSLWKGVVPG 81
>At2g33820 mitochondrial basic amino acid carrier (BAC1)
Length = 311
Score = 62.8 bits (151), Expect = 2e-10
Identities = 55/187 (29%), Positives = 86/187 (45%), Gaps = 21/187 (11%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
VA + G A + +P D V+ +LQ H+ + Q RY N H I + EG+KGLY G
Sbjct: 19 VAGMMAGLATVAVGHPFDTVKVKLQKHNTDV--QGLRYKNGLHCASRILQTEGVKGLYRG 76
Query: 70 FPAGLLGATISWSLL--------VFYYGIVKDRHARSKV-----EKSGALMTVCLCANPA 116
+ +G SL+ +F G + D R ++ GA+++ LC P
Sbjct: 77 ATSSFMGMAFESSLMFGIYSQAKLFLRGTLPDDGPRPEIIVPSAMFGGAIISFVLC--PT 134
Query: 117 FVVKTRLQLQ---TPLHHARPYSGLYD-AFRTIKREEGFSAFYRGIVPGFFLISQAAIQF 172
+VK R+Q+Q + + + R Y+ D A +T+K + F G + A+ F
Sbjct: 135 ELVKCRMQIQGTDSLVPNFRRYNSPLDCAVQTVKNDGVTGIFRGGSATLLRECTGNAVFF 194
Query: 173 IVYEQLR 179
VYE LR
Sbjct: 195 TVYEYLR 201
Score = 31.2 bits (69), Expect = 0.55
Identities = 25/86 (29%), Positives = 37/86 (42%), Gaps = 14/86 (16%)
Query: 16 GFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG------ 69
G A + P DV +T +Q S+ N + +I ++ GLKG YAG
Sbjct: 232 GIACWSAVLPFDVAKTIIQTS-----SEKATERNPFKVLSSIHKRAGLKGCYAGLGPTIV 286
Query: 70 --FPAGLLGATISWSLLVFYYGIVKD 93
FPA A ++W + GI +D
Sbjct: 287 RAFPAN-AAAIVAWEFSMKMLGIKRD 311
>At5g51050 calcium-binding transporter-like protein
Length = 487
Score = 62.0 bits (149), Expect = 3e-10
Identities = 50/174 (28%), Positives = 79/174 (44%), Gaps = 26/174 (14%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
YPLD+V+TRLQ + + PR T I EG + Y G LLG +
Sbjct: 322 YPLDLVKTRLQTYTSQAGVAVPRLGTLTKDILV---HEGPRAFYKGLFPSLLGIIPYAGI 378
Query: 84 LVFYYGIVKDRHARSKVEK--------------SGALMTVCLCANPAFVVKTRLQLQTPL 129
+ Y +KD ++ SGAL C+ P VV+TR+Q +
Sbjct: 379 DLAAYETLKDLSRTYILQDAEPGPLVQLGCGTISGALGATCVY--PLQVVRTRMQAE--- 433
Query: 130 HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLRKTV 182
R + + FR EEG+ A Y+G++P ++ A+I ++VYE ++K++
Sbjct: 434 ---RARTSMSGVFRRTISEEGYRALYKGLLPNLLKVVPAASITYMVYEAMKKSL 484
Score = 27.7 bits (60), Expect = 6.0
Identities = 20/72 (27%), Positives = 29/72 (39%), Gaps = 8/72 (11%)
Query: 21 TFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATIS 80
T YPL VV+TR+Q R + + +EG + LY G LL +
Sbjct: 418 TCVYPLQVVRTRMQAERAR--------TSMSGVFRRTISEEGYRALYKGLLPNLLKVVPA 469
Query: 81 WSLLVFYYGIVK 92
S+ Y +K
Sbjct: 470 ASITYMVYEAMK 481
>At2g39970 36kDa-peroxisomal membrane protein (PMP36)
Length = 331
Score = 61.2 bits (147), Expect = 5e-10
Identities = 54/207 (26%), Positives = 88/207 (42%), Gaps = 43/207 (20%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
YPL V TR Q R + R T + + ++EG + LY G L G S +
Sbjct: 23 YPLQTVNTRQQTE--RDLKREKRKLGTIEHMCQVVKQEGWERLYGGLAPSLAGTAASQGV 80
Query: 84 LVFYYGIVKDRH-----ARSK-------VEKSGALMTVC-------LCANPAFVVKTRLQ 124
++Y + ++R AR K V +L+ L NP +V+ TR+Q
Sbjct: 81 YYYFYQVFRNRAEATALARKKKGLGDGSVGMFASLLVAAFAGSVNVLMTNPIWVIVTRMQ 140
Query: 125 L------------QTPLHHA--------RPYSGLYDAFRTIKREEGFSAFYRGIVPGFFL 164
++P +A RPY G ++ R + E G + F++G++P +
Sbjct: 141 THRKMTKDQTAAPESPSSNAEALVAVEPRPY-GTFNTIREVYDEAGITGFWKGVIPTLIM 199
Query: 165 ISQAAIQFIVYE-QLRKTVVNLKTKGS 190
+S ++QF++YE L K KGS
Sbjct: 200 VSNPSMQFMLYETMLTKLKKKRALKGS 226
Score = 38.1 bits (87), Expect = 0.004
Identities = 47/193 (24%), Positives = 77/193 (39%), Gaps = 41/193 (21%)
Query: 25 PLDVVQTRLQVHDGR-----------------LFSQYPRYNNTTHAIFTIARKEGLKGLY 67
P+ V+ TR+Q H L + PR T + I + + G+ G +
Sbjct: 131 PIWVIVTRMQTHRKMTKDQTAAPESPSSNAEALVAVEPRPYGTFNTIREVYDEAGITGFW 190
Query: 68 AG-FPAGLLGATISWSLLVFYYGIVKDRHARS----------------KVEKSGALMTVC 110
G P ++ + S +++ + K + R+ V K GA +T
Sbjct: 191 KGVIPTLIMVSNPSMQFMLYETMLTKLKKKRALKGSNNVTALETFLLGAVAKLGATVTTY 250
Query: 111 LCANPAFVVKTRLQLQ--TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQ 167
P VVK+RLQ + T + Y G DA + R EG FY+G+ +
Sbjct: 251 ----PLLVVKSRLQAKQVTTGDKRQQYKGTLDAILKMIRYEGLYGFYKGMSTKIVQSVLA 306
Query: 168 AAIQFIVYEQLRK 180
AA+ F++ E+L K
Sbjct: 307 AAVLFMIKEELVK 319
Score = 37.0 bits (84), Expect = 0.010
Identities = 20/61 (32%), Positives = 32/61 (51%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
YPL VV++RLQ + +Y T AI + R EGL G Y G ++ + ++ ++
Sbjct: 250 YPLLVVKSRLQAKQVTTGDKRQQYKGTLDAILKMIRYEGLYGFYKGMSTKIVQSVLAAAV 309
Query: 84 L 84
L
Sbjct: 310 L 310
Score = 27.7 bits (60), Expect = 6.0
Identities = 18/65 (27%), Positives = 27/65 (40%), Gaps = 2/65 (3%)
Query: 111 LCANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAI 170
L P V TR Q + L + G + + ++EG+ Y G+ P L AA
Sbjct: 20 LLTYPLQTVNTRQQTERDLKREKRKLGTIEHMCQVVKQEGWERLYGGLAPS--LAGTAAS 77
Query: 171 QFIVY 175
Q + Y
Sbjct: 78 QGVYY 82
>At4g32400 adenylate translocator (brittle-1) - like protein
Length = 392
Score = 60.1 bits (144), Expect = 1e-09
Identities = 48/169 (28%), Positives = 78/169 (45%), Gaps = 26/169 (15%)
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLL 84
PL+ ++T L V G N++T I + EG GL+ G ++ + ++
Sbjct: 130 PLETIRTHLMVGSGG--------NSSTEVFSDIMKHEGWTGLFRGNLVNVIRVAPARAVE 181
Query: 85 VFYYGIVKDRHA-----RSKVEKSGALMT-VC------LCANPAFVVKTRLQLQTPLHHA 132
+F + V + + SK+ +L+ C L P +VKTRL +Q +
Sbjct: 182 LFVFETVNKKLSPPHGQESKIPIPASLLAGACAGVSQTLLTYPLELVKTRLTIQRGV--- 238
Query: 133 RPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLRK 180
Y G++DAF I REEG + YRG+ P ++ AA + Y+ LRK
Sbjct: 239 --YKGIFDAFLKIIREEGPTELYRGLAPSLIGVVPYAATNYFAYDSLRK 285
Score = 53.9 bits (128), Expect = 8e-08
Identities = 51/185 (27%), Positives = 77/185 (41%), Gaps = 18/185 (9%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A G + YPL++V+TRL + G Y A I R+EG LY G
Sbjct: 209 LAGACAGVSQTLLTYPLELVKTRLTIQRG-------VYKGIFDAFLKIIREEGPTELYRG 261
Query: 70 FPAGLLGATISWSLLVFYY-GIVKDRHARSKVEKSGALMTV-------CLCANPAFVVK- 120
L+G + F Y + K + SK EK G + T+ L + F ++
Sbjct: 262 LAPSLIGVVPYAATNYFAYDSLRKAYRSFSKQEKIGNIETLLIGSLAGALSSTATFPLEV 321
Query: 121 TRLQLQTPLHHAR-PYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQL 178
R +Q R Y + A TI EG +Y+G+ P L+ A I F+ YE
Sbjct: 322 ARKHMQVGAVSGRVVYKNMLHALVTILEHEGILGWYKGLGPSCLKLVPAAGISFMCYEAC 381
Query: 179 RKTVV 183
+K ++
Sbjct: 382 KKILI 386
>At3g55640 Ca-dependent solute carrier - like protein
Length = 332
Score = 57.8 bits (138), Expect = 5e-09
Identities = 50/183 (27%), Positives = 80/183 (43%), Gaps = 19/183 (10%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
VA G + YPLD+V+TRL ++ Y+ H + +I EG+ GLY G
Sbjct: 146 VAGGLAGITAASATYPLDLVRTRLAAQTKVIY-----YSGIWHTLRSITTDEGILGLYKG 200
Query: 70 FPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCAN-----------PAFV 118
L+G S ++ Y ++ + RS +M C + P +
Sbjct: 201 LGTTLVGVGPSIAISFSVYESLRS-YWRSTRPHDSPIMVSLACGSLSGIASSTATFPLDL 259
Query: 119 VKTRLQLQTPLHHARPY-SGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYE 176
V+ R QL+ A Y +GL + I + EG YRGI+P ++ ++ I F+ YE
Sbjct: 260 VRRRKQLEGIGGRAVVYKTGLLGTLKRIVQTEGARGLYRGILPEYYKVVPGVGICFMTYE 319
Query: 177 QLR 179
L+
Sbjct: 320 TLK 322
>At5g61810 peroxisomal Ca-dependent solute carrier - like protein
Length = 478
Score = 56.6 bits (135), Expect = 1e-08
Identities = 48/173 (27%), Positives = 77/173 (43%), Gaps = 25/173 (14%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
YP+D+V+TRLQ + + P+ T I+ +EG + Y G L+G +
Sbjct: 314 YPMDLVKTRLQTFVSEVGT--PKLWKLTKDIWI---QEGPRAFYRGLCPSLIGIIPYAGI 368
Query: 84 LVFYYGIVKD---RHARSKVEKSGALMTV----------CLCANPAFVVKTRLQLQTPLH 130
+ Y +KD H + G L+ + C P V++TR+Q +
Sbjct: 369 DLAAYETLKDLSRAHFLHDTAEPGPLIQLGCGMTSGALGASCVYPLQVIRTRMQADSSK- 427
Query: 131 HARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLRKTV 182
+ + F R EG FYRGI P FF +I A+I ++VYE ++K +
Sbjct: 428 -----TSMGQEFLKTLRGEGLKGFYRGIFPNFFKVIPSASISYLVYEAMKKNL 475
Score = 30.0 bits (66), Expect = 1.2
Identities = 21/80 (26%), Positives = 31/80 (38%), Gaps = 8/80 (10%)
Query: 13 LTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPA 72
+T+G + YPL V++TR+Q + + R EGLKG Y G
Sbjct: 401 MTSGALGASCVYPLQVIRTRMQADSSK--------TSMGQEFLKTLRGEGLKGFYRGIFP 452
Query: 73 GLLGATISWSLLVFYYGIVK 92
S S+ Y +K
Sbjct: 453 NFFKVIPSASISYLVYEAMK 472
>At1g14140 mitochondrial uncoupling protein - like
Length = 305
Score = 56.6 bits (135), Expect = 1e-08
Identities = 48/182 (26%), Positives = 82/182 (44%), Gaps = 22/182 (12%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
+P+D+ +TR+Q+H S R + IARKEG+ GLY G ++ +
Sbjct: 31 FPIDLTKTRMQLHGSGSASGAHRIG-AFGVVSEIARKEGVIGLYKGLSPAIIRHLFYTPI 89
Query: 84 LVFYYGIVKDRHARSKVEKSGAL-------------MTVCLCANPAFVVKTRLQLQTPL- 129
+ Y +K RS+ S +L + + A+PA +VK R+Q L
Sbjct: 90 RIIGYENLKGLIVRSETNNSESLPLATKALVGGFSGVIAQVVASPADLVKVRMQADGRLV 149
Query: 130 -HHARP-YSGLYDAFRTIKREEGFSAFYRGIVPGF---FLISQAAIQFIVYEQLRKTVVN 184
+P YSG +AF I + EG ++G++P FL++ + Y+ + V++
Sbjct: 150 SQGLKPRYSGPIEAFTKILQSEGVKGLWKGVLPNIQRAFLVNMG--ELACYDHAKHFVID 207
Query: 185 LK 186
K
Sbjct: 208 KK 209
Score = 31.2 bits (69), Expect = 0.55
Identities = 17/63 (26%), Positives = 31/63 (48%), Gaps = 5/63 (7%)
Query: 8 YPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLY 67
+ +AS+ +G A + P DVV+TR+ + Y N+ + + EG++ L+
Sbjct: 218 HTLASIMSGLASTSLSCPADVVKTRMMNQ-----GENAVYRNSYDCLVKTVKFEGIRALW 272
Query: 68 AGF 70
GF
Sbjct: 273 KGF 275
>At4g39460 mitochondrial carrier like protein
Length = 325
Score = 55.1 bits (131), Expect = 4e-08
Identities = 57/188 (30%), Positives = 83/188 (43%), Gaps = 27/188 (14%)
Query: 10 VASLTTG----FAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
VA LT G A + P +VV+ R+Q ++ + A+ IA KEG +G
Sbjct: 136 VAHLTAGAIGGLAASLIRVPTEVVKQRMQTG---------QFTSAPSAVRMIASKEGFRG 186
Query: 66 LYAGFPAGLLGATISWSLLVFYYGIV----KDRHARSKVEKSGALMTVCLCA------NP 115
LYAG+ + LL ++ Y + K R + AL+ A P
Sbjct: 187 LYAGYRSFLLRDLPFDAIQFCIYEQLCLGYKKAARRELSDPENALIGAFAGALTGAVTTP 246
Query: 116 AFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS-QAAIQFIV 174
V+KTRL +Q A+ Y G+ D +TI REEG A +GI P I +I F V
Sbjct: 247 LDVIKTRLMVQGS---AKQYQGIVDCVQTIVREEGAPALLKGIGPRVLWIGIGGSIFFGV 303
Query: 175 YEQLRKTV 182
E ++T+
Sbjct: 304 LESTKRTL 311
Score = 48.1 bits (113), Expect = 4e-06
Identities = 50/181 (27%), Positives = 70/181 (38%), Gaps = 39/181 (21%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A T G T YP+D ++TRLQ G K LKGLY+G
Sbjct: 59 IAGGTAGVVVETALYPIDTIKTRLQAARG-------------------GGKIVLKGLYSG 99
Query: 70 FPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGAL----------MTVCLCANPAFVV 119
+ G + +L V Y K + ++ + A+ + L P VV
Sbjct: 100 LAGNIAGVLPASALFVGVYEPTKQKLLKTFPDHLSAVAHLTAGAIGGLAASLIRVPTEVV 159
Query: 120 KTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFL--ISQAAIQFIVYEQ 177
K R+Q ++ A R I +EGF Y G F L + AIQF +YEQ
Sbjct: 160 KQRMQ-------TGQFTSAPSAVRMIASKEGFRGLYAG-YRSFLLRDLPFDAIQFCIYEQ 211
Query: 178 L 178
L
Sbjct: 212 L 212
>At5g46800 carnitine/acylcarnitine translocase-like protein
Length = 300
Score = 53.1 bits (126), Expect = 1e-07
Identities = 52/198 (26%), Positives = 79/198 (39%), Gaps = 25/198 (12%)
Query: 6 WEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
W+ + G A + +P D ++ +LQ Q PRY A+ EG KG
Sbjct: 5 WKDLASGTVGGAAQLVVGHPFDTIKVKLQSQPTPAPGQLPRYTGAIDAVKQTVASEGTKG 64
Query: 66 LYAGFPAGLLGATISWSLLVFY------YGIVKDRH------ARSKVEKSGALMTVCLCA 113
LY G A L AT++ V + G+++ ++ V +GA V A
Sbjct: 65 LYKGMGAPL--ATVAAFNAVLFTVRGQMEGLLRSEAGVPLTISQQFVAGAGAGFAVSFLA 122
Query: 114 NPAFVVKTRLQLQTPLHHAR---------PYSGLYDAFRTIKREEGFS-AFYRGIVPGFF 163
P ++K RLQ Q L A Y G D R + R EG + ++G+ P F
Sbjct: 123 CPTELIKCRLQAQGALAGASTTSSVVAAVKYGGPMDVARHVLRSEGGARGLFKGLFPTFA 182
Query: 164 L-ISQAAIQFIVYEQLRK 180
+ A F YE ++
Sbjct: 183 REVPGNATMFAAYEAFKR 200
Score = 42.7 bits (99), Expect = 2e-04
Identities = 25/61 (40%), Positives = 33/61 (53%), Gaps = 4/61 (6%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A G +F YP DVV++ LQV D + PRY + A I + EG+KGLY G
Sbjct: 218 MAGGVAGASFWGIVYPTDVVKSVLQVDDYK----NPRYTGSMDAFRKILKSEGVKGLYKG 273
Query: 70 F 70
F
Sbjct: 274 F 274
Score = 36.6 bits (83), Expect = 0.013
Identities = 44/194 (22%), Positives = 71/194 (35%), Gaps = 24/194 (12%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYP-------RYNNTTHAIFTIARKEG 62
VA GFA P ++++ RLQ + +Y + R EG
Sbjct: 109 VAGAGAGFAVSFLACPTELIKCRLQAQGALAGASTTSSVVAAVKYGGPMDVARHVLRSEG 168
Query: 63 -----LKGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVCLCAN--- 114
KGL+ F + G ++ + + S + + +M +
Sbjct: 169 GARGLFKGLFPTFAREVPGNATMFAAYEAFKRFLAGGSDTSSLGQGSLIMAGGVAGASFW 228
Query: 115 ----PAFVVKTRLQLQTPLHHARP-YSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQA 168
P VVK+ LQ+ + P Y+G DAFR I + EG Y+G P +
Sbjct: 229 GIVYPTDVVKSVLQVDD---YKNPRYTGSMDAFRKILKSEGVKGLYKGFGPAMARSVPAN 285
Query: 169 AIQFIVYEQLRKTV 182
A F+ YE R ++
Sbjct: 286 AACFLAYEMTRSSL 299
>At3g51870 carrier like protein
Length = 381
Score = 53.1 bits (126), Expect = 1e-07
Identities = 45/193 (23%), Positives = 79/193 (40%), Gaps = 26/193 (13%)
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGF 70
A G YPLDV++ RL V PRY + ++ R EG+ Y G
Sbjct: 193 AGACAGMTSTLLTYPLDVLRLRLAVE--------PRYRTMSQVALSMLRDEGIASFYYGL 244
Query: 71 PAGLLGATISWSLLVFYYGIVK---DRHARSKVEKSGALMTVCLCAN-------PAFVVK 120
L+G ++ + +VK R K + S L+T L A P V+
Sbjct: 245 GPSLVGIAPYIAVNFCIFDLVKKSLPEEYRKKAQSS--LLTAVLSAGIATLTCYPLDTVR 302
Query: 121 TRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLR 179
++Q++ PY + +AF I +G YRG +P + ++I+ ++ ++
Sbjct: 303 RQMQMR-----GTPYKSIPEAFAGIIDRDGLIGLYRGFLPNALKTLPNSSIRLTTFDMVK 357
Query: 180 KTVVNLKTKGSKI 192
+ + + + KI
Sbjct: 358 RLIATSEKQLQKI 370
Score = 41.2 bits (95), Expect = 5e-04
Identities = 46/183 (25%), Positives = 82/183 (44%), Gaps = 17/183 (9%)
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYN-NTTHAIFTIARKEGLKGLYAG 69
A G A T PLD ++ +Q H RL Q + AI IA++EG+KG + G
Sbjct: 93 AGALAGAAAKTVTAPLDRIKLLMQTHGIRLGQQSAKKAIGFIEAITLIAKEEGVKGYWKG 152
Query: 70 FPAGLLGAT--ISWSLLVF--YYGIVKDRHARSKV-----EKSGALMTVCLCANPAFVVK 120
++ + LL + Y + K + + V + A MT L P V++
Sbjct: 153 NLPQVIRVLPYSAVQLLAYESYKNLFKGKDDQLSVIGRLAAGACAGMTSTLLTYPLDVLR 212
Query: 121 TRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLIS-QAAIQFIVYEQLR 179
RL ++ Y + ++ R+EG ++FY G+ P I+ A+ F +++ ++
Sbjct: 213 LRLAVEP------RYRTMSQVALSMLRDEGIASFYYGLGPSLVGIAPYIAVNFCIFDLVK 266
Query: 180 KTV 182
K++
Sbjct: 267 KSL 269
>At1g14560 mitochondrial carrier like protein
Length = 331
Score = 53.1 bits (126), Expect = 1e-07
Identities = 48/195 (24%), Positives = 82/195 (41%), Gaps = 26/195 (13%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRL--QVHDGRL--------FSQYPRYNNTTHAIFTIAR 59
VA G + YPLD+ +T+L QV D R F + P Y+ + +
Sbjct: 124 VAGSAAGGTAVLCTYPLDLARTKLAYQVSDTRQSLRGGANGFYRQPTYSGIKEVLAMAYK 183
Query: 60 KEGLKGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTVC--------- 110
+ G +GLY G L+G + ++ L FY RH + + S + C
Sbjct: 184 EGGPRGLYRGIGPTLIG-ILPYAGLKFYIYEELKRHVPEEHQNSVRMHLPCGALAGLFGQ 242
Query: 111 LCANPAFVVKTRLQLQ-----TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-L 164
P VV+ ++Q++ T + + Y +D TI R +G+ + G+ + +
Sbjct: 243 TITYPLDVVRRQMQVENLQPMTSEGNNKRYKNTFDGLNTIVRTQGWKQLFAGLSINYIKI 302
Query: 165 ISQAAIQFIVYEQLR 179
+ AI F VYE ++
Sbjct: 303 VPSVAIGFTVYESMK 317
Score = 38.5 bits (88), Expect = 0.003
Identities = 25/71 (35%), Positives = 37/71 (51%), Gaps = 4/71 (5%)
Query: 2 NELQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQY---PRYNNTTHAIFTIA 58
N ++ P +L G T YPLDVV+ ++QV + + + RY NT + TI
Sbjct: 225 NSVRMHLPCGALA-GLFGQTITYPLDVVRRQMQVENLQPMTSEGNNKRYKNTFDGLNTIV 283
Query: 59 RKEGLKGLYAG 69
R +G K L+AG
Sbjct: 284 RTQGWKQLFAG 294
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.143 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,402,007
Number of Sequences: 26719
Number of extensions: 214139
Number of successful extensions: 868
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 596
Number of HSP's gapped (non-prelim): 154
length of query: 250
length of database: 11,318,596
effective HSP length: 97
effective length of query: 153
effective length of database: 8,726,853
effective search space: 1335208509
effective search space used: 1335208509
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC125480.1


