

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124972.13 - phase: 0
(935 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
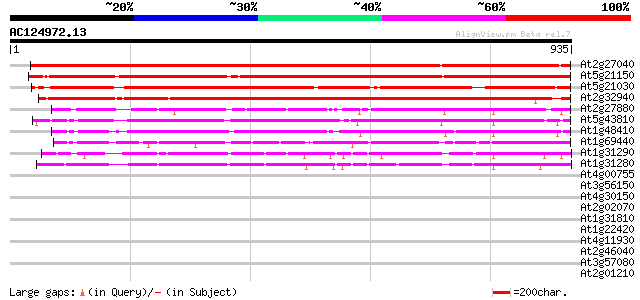
Score E
Sequences producing significant alignments: (bits) Value
At2g27040 Argonaute (AGO1)-like protein 1350 0.0
At5g21150 zwille/pinhead-like protein 1230 0.0
At5g21030 zwille/pinhead-like protein 1017 0.0
At2g32940 Argonaute (AGO1)-like protein 940 0.0
At2g27880 Argonaute (AGO1)-like protein 485 e-137
At5g43810 PINHEAD (gb|AAD40098.1); translation initiation factor 476 e-134
At1g48410 Argonaute protein (AGO1) 458 e-129
At1g69440 hypothetical protein 363 e-100
At1g31290 hypothetical protein 325 7e-89
At1g31280 unknown protein 325 1e-88
At4g00755 putative F-box protein 33 0.98
At3g56150 PROBABLE EUKARYOTIC TRANSLATION INITIATION FACTOR 3 SU... 32 1.7
At4g30150 hypothetical protein 32 2.2
At2g02070 putative C2H2-type zinc finger protein 31 2.8
At1g31810 hypothetical protein 31 2.8
At1g22420 31 2.8
At4g11930 hypothetical protein 30 4.8
At2g46040 unknown protein 30 4.8
At3g57080 unknown protein 30 6.3
At2g01210 putative receptor-like protein kinase 30 6.3
>At2g27040 Argonaute (AGO1)-like protein
Length = 924
Score = 1350 bits (3495), Expect = 0.0
Identities = 644/902 (71%), Positives = 777/902 (85%), Gaps = 4/902 (0%)
Query: 35 ECLPPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNT 94
E LPPPPP++P ++EP++++ ++ +KK P +VPMAR+G G++G K+PLLTNHFKV+V N
Sbjct: 24 EALPPPPPVIPPNVEPVRVKTELAEKKGPVRVPMARKGFGTRGQKIPLLTNHFKVDVANL 83
Query: 95 DGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFTIGSLA 154
G+FF YSVALFY+DGRPVE KG GRKILD+V +TY S+L+GK+ AYDGEKTLFT G+L
Sbjct: 84 QGHFFHYSVALFYDDGRPVEQKGVGRKILDKVHQTYHSDLDGKEFAYDGEKTLFTYGALP 143
Query: 155 QNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQ 214
NK++F+VVLE+V++ R NGN SP+G+ SP+D DRKRL++ +RSK ++VEIS+A+KIPLQ
Sbjct: 144 SNKMDFSVVLEEVSATRANGNGSPNGNESPSDGDRKRLRRPNRSKNFRVEISYAAKIPLQ 203
Query: 215 AIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCR 274
A+ANA++G E+EN QEAIRVLDIILRQHAA+QGCLLVRQ+FFHNDP N VGG +LGCR
Sbjct: 204 ALANAMRGQESENSQEAIRVLDIILRQHAARQGCLLVRQSFFHNDPTNCEPVGGNILGCR 263
Query: 275 GLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNKAKRTLKNLRI 334
G HSSFRTTQ G+SLN+DV+TTMI+ PGPVVDFLIANQN RDP+S+DW+KAKRTLKNLR+
Sbjct: 264 GFHSSFRTTQGGMSLNMDVTTTMIIKPGPVVDFLIANQNARDPYSIDWSKAKRTLKNLRV 323
Query: 335 TTSPTNQEYKITGLSEMPCKDQLFTLKKRGA-VPGEDDTEEITVYDYFVNRRKISLQYSA 393
SP+ QE+KITGLS+ PC++Q F LKKR GE +T E+TV DYF + R I LQYSA
Sbjct: 324 KVSPSGQEFKITGLSDKPCREQTFELKKRNPNENGEFETTEVTVADYFRDTRHIDLQYSA 383
Query: 394 DLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDAL 453
DLPCINVGKPKRPT++P+ELC+LV LQRYTKAL+T QRS+LVEKSRQKPQERM VL+ AL
Sbjct: 384 DLPCINVGKPKRPTYIPLELCALVPLQRYTKALTTFQRSALVEKSRQKPQERMTVLSKAL 443
Query: 454 KTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKKIVQP 513
K S+Y +EP+LR+CGISI+S FTQV+GRVL AP+LK G G + PRNGRWNFNNK+ V+P
Sbjct: 444 KVSNYDAEPLLRSCGISISSNFTQVEGRVLPAPKLKMGCGSETFPRNGRWNFNNKEFVEP 503
Query: 514 VKIEKWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKM 573
KI++W VVNFSARC+VR +V DLIK GG KGI + PF FEE QFRRAPP++RVE M
Sbjct: 504 TKIQRWVVVNFSARCNVRQVVDDLIKIGGSKGIEIASPFQVFEEGNQFRRAPPMIRVENM 563
Query: 574 FEHVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTR-VNDQYLTNV 632
F+ +QSKLPG P+F+LC+L ++KNSDLYGPWKKKNL EFGIVTQC+APTR NDQYLTN+
Sbjct: 564 FKDIQSKLPGVPQFILCVLPDKKNSDLYGPWKKKNLTEFGIVTQCMAPTRQPNDQYLTNL 623
Query: 633 LLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQW 692
LLKINAKLGG+NS+L VE +P+ ++SK PT+ILGMDVSHGSPGQ+++PSIAAVVSSR+W
Sbjct: 624 LLKINAKLGGLNSMLSVERTPAFTVISKVPTIILGMDVSHGSPGQSDVPSIAAVVSSREW 683
Query: 693 PLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRD 752
PLISKYRA VRTQ +K EMI++L K + TED+GII+ELL+DFY SS RKP++IIIFRD
Sbjct: 684 PLISKYRASVRTQPSKAEMIESLVKK-NGTEDDGIIKELLVDFYTSSNKRKPEHIIIFRD 742
Query: 753 GVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPDNVPPGTV 812
GVSESQFNQVLNIEL QIIEACK LD WNPKFL++VAQKNHHTKFFQP SP+NVPPGT+
Sbjct: 743 GVSESQFNQVLNIELDQIIEACKLLDANWNPKFLLLVAQKNHHTKFFQPTSPENVPPGTI 802
Query: 813 VDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTT 872
+DNKICHP+N DFY+CAHAGMIGT+RPTHYHVL DEIGFS D+LQELVHSLSYVYQRST+
Sbjct: 803 IDNKICHPKNNDFYLCAHAGMIGTTRPTHYHVLYDEIGFSADELQELVHSLSYVYQRSTS 862
Query: 873 AISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIPQLPKLMDSVCNSM 932
AISVVAPICYAHLAA+Q+G FMKFED+SETSSSHGG S + QLP+L D+V NSM
Sbjct: 863 AISVVAPICYAHLAAAQLGTFMKFEDQSETSSSHGGITAPGPIS-VAQLPRLKDNVANSM 921
Query: 933 FF 934
FF
Sbjct: 922 FF 923
>At5g21150 zwille/pinhead-like protein
Length = 892
Score = 1230 bits (3182), Expect = 0.0
Identities = 609/906 (67%), Positives = 733/906 (80%), Gaps = 21/906 (2%)
Query: 31 QENEECLPPPPPIVPADIEPIKIEPQIVKKKLPTKVPMAR-RGLGSKGAKLPLLTNHFKV 89
+ N LPPPPP VPA++ P ++EP VKK + +PMAR RG GSKG K+PLLTNHF V
Sbjct: 5 EPNGSGLPPPPPFVPANLVP-EVEP--VKKNI--LLPMARPRGSGSKGQKIPLLTNHFGV 59
Query: 90 NVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFT 149
GYFF YSVA+ YEDGRPVE KG GRKILD+VQETY S+L K AYDGEKTLFT
Sbjct: 60 KFNKPSGYFFHYSVAINYEDGRPVEAKGIGRKILDKVQETYQSDLGAKYFAYDGEKTLFT 119
Query: 150 IGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFAS 209
+G+L NKL+F+VVLE++ S+RN+ ND DRKR ++ +++K + VEIS+A+
Sbjct: 120 VGALPSNKLDFSVVLEEIPSSRNHAG------NDTNDADRKRSRRPNQTKKFMVEISYAA 173
Query: 210 KIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGG 269
KIP+QAIA+AL+G ETEN Q+A+RVLDIILRQ AA+QGCLLVRQ+FFHND KNF +GGG
Sbjct: 174 KIPMQAIASALQGKETENLQDALRVLDIILRQSAARQGCLLVRQSFFHNDVKNFVPIGGG 233
Query: 270 VLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNKAKRTL 329
V GCRG HSSFRTTQ GLSLNID STTMIV PGPVVDFL+ANQN +DP+ +DWNKA+R L
Sbjct: 234 VSGCRGFHSSFRTTQGGLSLNIDTSTTMIVQPGPVVDFLLANQNKKDPYGMDWNKARRVL 293
Query: 330 KNLRITTSPTNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISL 389
KNLR+ + +N+EYKI+GLSE CKDQL K GE + EITV +Y+ R I +
Sbjct: 294 KNLRVQITLSNREYKISGLSEHSCKDQLKPNDK-----GEFEEVEITVLNYY-KERNIEV 347
Query: 390 QYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVL 449
+YS D PCINVGKPKRPT+ P+E C+LVSLQRYTK+L+ QR++LVEKSRQKP ERM L
Sbjct: 348 RYSGDFPCINVGKPKRPTYFPIEFCNLVSLQRYTKSLTNFQRAALVEKSRQKPPERMASL 407
Query: 450 TDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKK 509
T LK S+Y ++P+L++ G+SI + FTQV+GR+L P LK G GE+ +P G+WNF K
Sbjct: 408 TKGLKDSNYNADPVLQDSGVSIITNFTQVEGRILPTPMLKVGKGENLSPIKGKWNFMRKT 467
Query: 510 IVQPVKIEKWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPF-DCFEENGQFRRAPPLV 568
+ +P + +WAVVNFSARCD L+RDLIKCG KGI+VE PF D EN QFR AP V
Sbjct: 468 LAEPTTVTRWAVVNFSARCDTNTLIRDLIKCGREKGINVEPPFKDVINENPQFRNAPATV 527
Query: 569 RVEKMFEHVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVNDQY 628
RVE MFE ++SKLP P FLLC+L+ERKNSD+YGPWKKKNL + GIVTQCIAPTR+NDQY
Sbjct: 528 RVENMFEQIKSKLPKPPLFLLCILAERKNSDVYGPWKKKNLVDLGIVTQCIAPTRLNDQY 587
Query: 629 LTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVS 688
LTNVLLKINAKLGG+NSLL +E SP++P V++ PT+I+GMDVSHGSPGQ++IPSIAAVVS
Sbjct: 588 LTNVLLKINAKLGGLNSLLAMERSPAMPKVTQVPTIIVGMDVSHGSPGQSDIPSIAAVVS 647
Query: 689 SRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNII 748
SRQWPLISKY+ACVRTQ K+EMIDNLFKPV+ +DEG+ RELL+DFY SS NRKP++II
Sbjct: 648 SRQWPLISKYKACVRTQSRKMEMIDNLFKPVNG-KDEGMFRELLLDFYYSSENRKPEHII 706
Query: 749 IFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPDNVP 808
IFRDGVSESQFNQVLNIEL Q+++ACKFLD+ W+PKF VIVAQKNHHTKFFQ PDNVP
Sbjct: 707 IFRDGVSESQFNQVLNIELDQMMQACKFLDDTWHPKFTVIVAQKNHHTKFFQSRGPDNVP 766
Query: 809 PGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQ 868
PGT++D++ICHPRN+DFY+CAHAGMIGT+RPTHYHVL DEIGF+ DDLQELVHSLSYVYQ
Sbjct: 767 PGTIIDSQICHPRNFDFYLCAHAGMIGTTRPTHYHVLYDEIGFATDDLQELVHSLSYVYQ 826
Query: 869 RSTTAISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIPQLPKLMDSV 928
RSTTAISVVAP+CYAHLAA+Q+G MK+E+ SETSSSHGG A P+P +P+L ++V
Sbjct: 827 RSTTAISVVAPVCYAHLAAAQMGTVMKYEELSETSSSHGGITTP-GAVPVPPMPQLHNNV 885
Query: 929 CNSMFF 934
SMFF
Sbjct: 886 STSMFF 891
>At5g21030 zwille/pinhead-like protein
Length = 850
Score = 1017 bits (2629), Expect = 0.0
Identities = 542/904 (59%), Positives = 668/904 (72%), Gaps = 65/904 (7%)
Query: 37 LPPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDG 96
LPPP + + EP+K + ++ PM RRG GSKG K+ LLTNHF+VN +
Sbjct: 5 LPPPQHM---EREPLKSKSSLL--------PMTRRGNGSKGQKILLLTNHFRVNFRKPNS 53
Query: 97 Y-FFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFTIGSLAQ 155
+ FF YSV + YEDG P+ KG GRKIL++VQ+T ++L K AYDG+K L+T+G L +
Sbjct: 54 HNFFHYSVTITYEDGSPLLAKGFGRKILEKVQQTCQADLGCKHFAYDGDKNLYTVGPLPR 113
Query: 156 NKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFAS-KIPLQ 214
+ L+F+VVLE S RN KRLK H+SK + V I FA +IP++
Sbjct: 114 SSLDFSVVLETAPSRRNAD---------------KRLKLPHQSKKFNVAILFAPPEIPME 158
Query: 215 AIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCR 274
AIANAL+G +T++ +AIRV+D IL Q+AA+QGCLLVRQ+FFHND K F ++G GV C+
Sbjct: 159 AIANALQGKKTKHLLDAIRVMDCILSQNAARQGCLLVRQSFFHNDAKYFANIGEGVDCCK 218
Query: 275 GLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNKAKRTLKNLRI 334
G HSSFRTTQ GLSLNIDVST MIV PGPVVDFLIANQ V DPFS++W KAK TLKNLR+
Sbjct: 219 GFHSSFRTTQGGLSLNIDVSTAMIVKPGPVVDFLIANQGVNDPFSINWKKAKNTLKNLRV 278
Query: 335 TTSPTNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSAD 394
P+NQEYKITGLS + CKDQ FT KKR E + EITV DYF R+I L+YS
Sbjct: 279 KVLPSNQEYKITGLSGLHCKDQTFTWKKRNQ-NREFEEVEITVSDYFTRIREIELRYSGG 337
Query: 395 LPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALK 454
LPCINVGKP RPT+ P+ELC LVSLQRYTKAL+ QRS+L+++SRQ PQ+R+ VLT ALK
Sbjct: 338 LPCINVGKPNRPTYFPIELCELVSLQRYTKALTKFQRSNLIKESRQNPQQRIGVLTRALK 397
Query: 455 TSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKKIVQPV 514
TS+Y +PML+ CG+ I S FTQV+GRVL P+LK G +D P NG WNF NK P
Sbjct: 398 TSNYNDDPMLQECGVRIGSDFTQVEGRVLPTPKLKAGKEQDIYPINGSWNFKNK----PA 453
Query: 515 KIEKWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPFD-CFEENGQFRRAPPLVRVEKM 573
+ +WAVVNFSARCD + ++ DL +CG MKGI+V+ P+ FEEN QF+ A VRV+KM
Sbjct: 454 TVTRWAVVNFSARCDPQKIIDDLTRCGKMKGINVDSPYHVVFEENPQFKDATGSVRVDKM 513
Query: 574 FEHVQSKLPGA-PKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTR-VNDQYLTN 631
F+H+QS L PKFLLC+L E+KNSD+Y +K+ + + +CI P + +NDQYLTN
Sbjct: 514 FQHLQSILGEVPPKFLLCIL-EKKNSDVY----EKSCSMWN--CECIVPPQNLNDQYLTN 566
Query: 632 VLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTE-IPSIAAVVSSR 690
+LLKINAKLGG+NS+L +E S ++P+V + PT+I+GMDVSHGSPGQ++ IPSIAAVVSSR
Sbjct: 567 LLLKINAKLGGLNSVLDMELSGTMPLVMRVPTIIIGMDVSHGSPGQSDHIPSIAAVVSSR 626
Query: 691 QWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIF 750
+WPLISKYRACVRTQ KVEMID+LFKPVSD +D+GI+RELL+DF++SSG +KP++IIIF
Sbjct: 627 EWPLISKYRACVRTQSPKVEMIDSLFKPVSDKDDQGIMRELLLDFHSSSG-KKPNHIIIF 685
Query: 751 RDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPDNVPPG 810
RDGVSESQFNQVLNIEL Q++ Q NHHTKFFQ SP+NV PG
Sbjct: 686 RDGVSESQFNQVLNIELDQMM-------------------QINHHTKFFQTESPNNVLPG 726
Query: 811 TVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRS 870
T++D+ ICH N DFY+CAHAG IGT+RPTHYHVL DEIGF D LQELVHSLSYVYQRS
Sbjct: 727 TIIDSNICHQHNNDFYLCAHAGKIGTTRPTHYHVLYDEIGFDTDQLQELVHSLSYVYQRS 786
Query: 871 TTAISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIPQLPKLMDSVCN 930
TTAIS+VAPICYAHLAA+Q+ MKFED SETSSSHGG A P+P +PKL +V +
Sbjct: 787 TTAISLVAPICYAHLAAAQMATAMKFEDMSETSSSHGGI-TTAGAVPVPPMPKLNTNVAS 845
Query: 931 SMFF 934
SMFF
Sbjct: 846 SMFF 849
>At2g32940 Argonaute (AGO1)-like protein
Length = 887
Score = 940 bits (2430), Expect = 0.0
Identities = 496/900 (55%), Positives = 632/900 (70%), Gaps = 35/900 (3%)
Query: 48 IEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFY 107
+ PI IEP+ + RRG+G+ G + L TNHF V+V D F+QY+V++
Sbjct: 9 LSPISIEPEQPSHR--DYDITTRRGVGTTGNPIELCTNHFNVSVRQPDVVFYQYTVSITT 66
Query: 108 EDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDV 167
E+G V+G G RK++D++ +TY S+L+GK LAYDGEKTL+T+G L QN+ +F V++E
Sbjct: 67 ENGDAVDGTGISRKLMDQLFKTYSSDLDGKRLAYDGEKTLYTVGPLPQNEFDFLVIVEGS 126
Query: 168 TSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHET-- 225
S R+ G + DG GS + T KR K+S ++YKV+I +A++IPL+ + +G T
Sbjct: 127 FSKRDCGVS--DG-GSSSGTC-KRSKRSFLPRSYKVQIHYAAEIPLKTVLGTQRGAYTPD 182
Query: 226 ENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQS 285
++ Q+A+RVLDI+LRQ AA++GCLLVRQ FFH+D VGGGV+G RGLHSSFR T
Sbjct: 183 KSAQDALRVLDIVLRQQAAERGCLLVRQAFFHSDGHPMK-VGGGVIGIRGLHSSFRPTHG 241
Query: 286 GLSLNIDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNK-AKRTLKNLRITTSPTNQEYK 344
GLSLNIDVSTTMI+ PGPV++FL ANQ+V P +DW K A + LK++R+ + N E+K
Sbjct: 242 GLSLNIDVSTTMILEPGPVIEFLKANQSVETPRQIDWIKVAAKMLKHMRVKATHRNMEFK 301
Query: 345 ITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPK 404
I GLS PC QLF++K + E EITVYDYF + SA PC++VGKP
Sbjct: 302 IIGLSSKPCNQQLFSMKIKDG-EREVPIREITVYDYFKQTYTEPIS-SAYFPCLDVGKPD 359
Query: 405 RPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPML 464
RP ++P+E C+LVSLQRYTK LS QR LVE SRQKP ER++ L DA+ T Y +P L
Sbjct: 360 RPNYLPLEFCNLVSLQRYTKPLSGRQRVLLVESSRQKPLERIKTLNDAMHTYCYDKDPFL 419
Query: 465 RNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNF 524
CGISI TQV+GRVL+ P LKFG EDF P NGRWNFNNK +++P I+ WA+VNF
Sbjct: 420 AGCGISIEKEMTQVEGRVLKPPMLKFGKNEDFQPCNGRWNFNNKMLLEPRAIKSWAIVNF 479
Query: 525 SARCDVRGLVRDLIKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPGA 584
S CD + R+LI CG KGI +++PF EE+ Q+++A P+ RVEKM ++ K P
Sbjct: 480 SFPCDSSHISRELISCGMRKGIEIDRPFALVEEDPQYKKAGPVERVEKMIATMKLKFPDP 539
Query: 585 PKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVNDQYLTNVLLKINAKLGGMN 644
P F+LC+L ERK SD+YGPWKK L E GI TQCI P +++DQYLTNVLLKIN+KLGG+N
Sbjct: 540 PHFILCILPERKTSDIYGPWKKICLTEEGIHTQCICPIKISDQYLTNVLLKINSKLGGIN 599
Query: 645 SLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRT 704
SLLG+E+S +IP+++K PTLILGMDVSHG PG+ ++PS+AAVV S+ WPLIS+YRA VRT
Sbjct: 600 SLLGIEYSYNIPLINKIPTLILGMDVSHGPPGRADVPSVAAVVGSKCWPLISRYRAAVRT 659
Query: 705 QGAKVEMIDNLFKPVSDTE--DEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQV 762
Q ++EMID+LF+P+ +TE D GI+ EL ++FY +S RKP IIIFRDGVSESQF QV
Sbjct: 660 QSPRLEMIDSLFQPIENTEKGDNGIMNELFVEFYRTSRARKPKQIIIFRDGVSESQFEQV 719
Query: 763 LNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPDNVPPGTVVDNKICHPRN 822
L IE+ QII+A + L E PKF VIVAQKNHHTK FQ P+NVP GTVVD KI HP N
Sbjct: 720 LKIEVDQIIKAYQRLGESDVPKFTVIVAQKNHHTKLFQAKGPENVPAGTVVDTKIVHPTN 779
Query: 823 YDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAIS------- 875
YDFYMCAHAG IGTSRP HYHVLLDEIGFSPDDLQ L+HSLSY S +S
Sbjct: 780 YDFYMCAHAGKIGTSRPAHYHVLLDEIGFSPDDLQNLIHSLSYKLLNSIFNVSSLLCVFV 839
Query: 876 -VVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIPQLPKLMDSVCNSMFF 934
VAP+ YAHLAA+QV QF KFE SE +P+LP+L ++V +MFF
Sbjct: 840 LSVAPVRYAHLAAAQVAQFTKFEGISEDGK-------------VPELPRLHENVEGNMFF 886
>At2g27880 Argonaute (AGO1)-like protein
Length = 997
Score = 485 bits (1249), Expect = e-137
Identities = 325/907 (35%), Positives = 475/907 (51%), Gaps = 102/907 (11%)
Query: 70 RRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQET 129
R G G+ G K+ + NHF V V + D Y + S+ V K R ++ + +
Sbjct: 150 RPGRGTLGKKVMVRANHFLVQVADRDLYHYDVSI------NPEVISKTVNRNVMKLLVKN 203
Query: 130 Y-GSELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTD 188
Y S L GK AYDG K+L+T G L + EF V L + ++ ++G P
Sbjct: 204 YKDSHLGGKSPAYDGRKSLYTAGPLPFDSKEFVVNLAEKRADGSSGKDRP---------- 253
Query: 189 RKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGC 248
+KV + + L + L + E + I+VLD++LR +
Sbjct: 254 ------------FKVAVKNVTSTDLYQLQQFLDRKQREAPYDTIQVLDVVLRDKPSND-Y 300
Query: 249 LLVRQNFFHNDPKNFTDVGGGVLG-----CRGLHSSFRTTQSGLSLNIDVSTTMIVHPGP 303
+ V ++FFH G G LG RG S R TQ GLSLNIDVS P
Sbjct: 301 VSVGRSFFHTSLGKDARDGRGELGDGIEYWRGYFQSLRLTQMGLSLNIDVSARSFYEPIV 360
Query: 304 VVDFLIANQNVRD---PF-SLDWNKAKRTLKNLRITTSPTN--QEYKITGLSEMPCKDQL 357
V DF+ N+RD P D K K+ L+ L++ N + KI+G+S +P ++
Sbjct: 361 VTDFISKFLNIRDLNRPLRDSDRLKVKKVLRTLKVKLLHWNCTKSAKISGISSLPIRELR 420
Query: 358 FTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLV 417
FTL +D E TV YF + ++Y A LP I G RP ++P+ELC +
Sbjct: 421 FTL---------EDKSEKTVVQYFAEKYNYRVKYQA-LPAIQTGSDTRPVYLPMELCQID 470
Query: 418 SLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQ 477
QRYTK L+ Q ++L++ + Q+P +R + + + ++Y + + + G+S+T+
Sbjct: 471 EGQRYTKRLNEKQVTALLKATCQRPPDRENSIKNLVVKNNYNDD-LSKEFGMSVTTQLAS 529
Query: 478 VDGRVLQAPRLKF---GNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCDVRGLV 534
++ RVL P LK+ G + NPR G+WN +KK+V K+ W V+FS R D RGL
Sbjct: 530 IEARVLPPPMLKYHDSGKEKMVNPRLGQWNMIDKKMVNGAKVTSWTCVSFSTRID-RGLP 588
Query: 535 RDLIKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHVQSKL-------PGAPKF 587
++ C + G+ C + +F+ P + + EH++ L PG +
Sbjct: 589 QEF--CKQLIGM-------CVSKGMEFKPQPAIPFISCPPEHIEEALLDIHKRAPGL-QL 638
Query: 588 LLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVND---QYLTNVLLKINAKLGGMN 644
L+ +L + S YG K+ E GIV+QC P +VN QY+ NV LKIN K GG N
Sbjct: 639 LIVILPDVTGS--YGKIKRICETELGIVSQCCQPRQVNKLNKQYMENVALKINVKTGGRN 696
Query: 645 SLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRT 704
++L +IP+++ PT+I+G DV+H PG+ PSIAAVV+S WP I+KYR V
Sbjct: 697 TVLNDAIRRNIPLITDRPTIIMGADVTHPQPGEDSSPSIAAVVASMDWPEINKYRGLVSA 756
Query: 705 QGAKVEMIDNLFKPVSDTE----DEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFN 760
Q + E+I +L+K V D + G+IRE I F ++G + P II +RDGVSE QF+
Sbjct: 757 QAHREEIIQDLYKLVQDPQRGLVHSGLIREHFIAFRRATG-QIPQRIIFYRDGVSEGQFS 815
Query: 761 QVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF--QPGSPD------NVPPGTV 812
QVL E++ I +AC L E + P+ ++ QK HHT+ F Q G+ D N+ PGTV
Sbjct: 816 QVLLHEMTAIRKACNSLQENYVPRVTFVIVQKRHHTRLFPEQHGNRDMTDKSGNIQPGTV 875
Query: 813 VDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTT 872
VD KICHP +DFY+ +HAG+ GTSRP HYHVLLDE GF+ D LQ L ++L Y Y R T
Sbjct: 876 VDTKICHPNEFDFYLNSHAGIQGTSRPAHYHVLLDENGFTADQLQMLTNNLCYTYARCTK 935
Query: 873 AISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASP-----IPQLPKLMDS 927
++S+V P YAHLAA + +M E+ S GGS R +++ I QLP + D+
Sbjct: 936 SVSIVPPAYYAHLAAFRARYYM------ESEMSDGGSSRSRSSTTGVGQVISQLPAIKDN 989
Query: 928 VCNSMFF 934
V MF+
Sbjct: 990 VKEVMFY 996
>At5g43810 PINHEAD (gb|AAD40098.1); translation initiation factor
Length = 988
Score = 476 bits (1224), Expect = e-134
Identities = 321/948 (33%), Positives = 485/948 (50%), Gaps = 104/948 (10%)
Query: 39 PPPP----------IVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFK 88
PPPP +I + + Q+ +K P R G G+ G K + NHF
Sbjct: 92 PPPPSQTTSSAVSVATAGEIVAVNHQMQMGVRKNSNFAP--RPGFGTLGTKCIVKANHFL 149
Query: 89 VNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDL-AYDGEKTL 147
++ D QY V + E V K R I+ + Y G+ L AYDG K+L
Sbjct: 150 ADLPTKD--LNQYDVTITPE----VSSKSVNRAIIAELVRLYKESDLGRRLPAYDGRKSL 203
Query: 148 FTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISF 207
+T G L EF+V + D NG R ++YKV I F
Sbjct: 204 YTAGELPFTWKEFSVKIVDEDDGIING--------------------PKRERSYKVAIKF 243
Query: 208 ASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVG 267
++ + + L G + QEA+++LDI+LR+ + K+ C + R +FF D K +G
Sbjct: 244 VARANMHHLGEFLAGKRADCPQEAVQILDIVLRELSVKRFCPVGR-SFFSPDIKTPQRLG 302
Query: 268 GGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDF---LIANQNVRDPFS-LDWN 323
G+ G + S R TQ GLSLNID+++ + P PV++F L+ + P S D
Sbjct: 303 EGLESWCGFYQSIRPTQMGLSLNIDMASAAFIEPLPVIEFVAQLLGKDVLSKPLSDSDRV 362
Query: 324 KAKRTLKNLRITTSP---TNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDY 380
K K+ L+ +++ + ++Y++ GL+ P ++ +F P +++ +V +Y
Sbjct: 363 KIKKGLRGVKVEVTHRANVRRKYRVAGLTTQPTRELMF--------PVDENCTMKSVIEY 414
Query: 381 FVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQ 440
F ++Q++ LPC+ VG K+ +++P+E C +V QRYTK L+ Q ++L++ + Q
Sbjct: 415 FQEMYGFTIQHT-HLPCLQVGNQKKASYLPMEACKIVEGQRYTKRLNEKQITALLKVTCQ 473
Query: 441 KPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDFN 497
+P++R + ++ + Y +P + G++I+ V+ R+L AP LK+ G +D
Sbjct: 474 RPRDRENDILRTVQHNAYDQDPYAKEFGMNISEKLASVEARILPAPWLKYHENGKEKDCL 533
Query: 498 PRNGRWNFNNKKIVQPVKIEKWAVVNFSARCD---VRGLVRDLIKCGGMKGIHVEQPFDC 554
P+ G+WN NKK++ + + +WA VNFS RG +L + + G+
Sbjct: 534 PQVGQWNMMNKKMINGMTVSRWACVNFSRSVQENVARGFCNELGQMCEVSGMEFNP---- 589
Query: 555 FEENGQFRRAPPLVRVEKMFEHV----QSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLA 610
E A P +VEK +HV +K G LL + N LYG K+
Sbjct: 590 -EPVIPIYSARP-DQVEKALKHVYHTSMNKTKGKELELLLAILPDNNGSLYGDLKRICET 647
Query: 611 EFGIVTQCIAPT---RVNDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILG 667
E G+++QC +++ QYL NV LKIN K+GG N++L S IP+VS PT+I G
Sbjct: 648 ELGLISQCCLTKHVFKISKQYLANVSLKINVKMGGRNTVLVDAISCRIPLVSDIPTIIFG 707
Query: 668 MDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFK----PVSDTE 723
DV+H G+ PSIAAVV+S+ WP ++KY V Q + E+I +L+K PV T
Sbjct: 708 ADVTHPENGEESSPSIAAVVASQDWPEVTKYAGLVCAQAHRQELIQDLYKTWQDPVRGTV 767
Query: 724 DEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNP 783
G+IR+LLI F ++G +KP II +RDGVSE QF QVL EL I +AC L+ + P
Sbjct: 768 SGGMIRDLLISFRKATG-QKPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLEPNYQP 826
Query: 784 KFLVIVAQKNHHTKFFQPGSPD--------NVPPGTVVDNKICHPRNYDFYMCAHAGMIG 835
IV QK HHT+ F D N+ PGTVVD KICHP +DFY+C+HAG+ G
Sbjct: 827 PVTFIVVQKRHHTRLFANNHRDKNSTDRSGNILPGTVVDTKICHPTEFDFYLCSHAGIQG 886
Query: 836 TSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFMK 895
TSRP HYHVL DE F+ D +Q L ++L Y Y R T ++S+V P YAHLAA + +
Sbjct: 887 TSRPAHYHVLWDENNFTADGIQSLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA----R 942
Query: 896 FEDKSETSSSHGGSGR---------DINASPIPQLPKLMDSVCNSMFF 934
F + E +G G+ D+ P LP L ++V MF+
Sbjct: 943 FYLEPEIMQDNGSPGKKNTKTTTVGDVGVKP---LPALKENVKRVMFY 987
>At1g48410 Argonaute protein (AGO1)
Length = 870
Score = 458 bits (1179), Expect = e-129
Identities = 310/916 (33%), Positives = 467/916 (50%), Gaps = 99/916 (10%)
Query: 70 RRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQET 129
R G G G + + NHF + + D Y V + E V +G R ++ ++ +
Sbjct: 2 RPGKGQSGKRCIVKANHFFAELPDKD--LHHYDVTITPE----VTSRGVNRAVMKQLVDN 55
Query: 130 Y-GSELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTD 188
Y S L + AYDG K+L+T G L N EF + L D G G
Sbjct: 56 YRDSHLGSRLPAYDGRKSLYTAGPLPFNSKEFRINLLD----------EEVGAGG----- 100
Query: 189 RKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQ-HAAKQG 247
R + +KV I ++ L + L+G +++ QEA++VLDI+LR+ ++
Sbjct: 101 ------QRREREFKVVIKLVARADLHHLGMFLEGKQSDAPQEALQVLDIVLRELPTSRIR 154
Query: 248 CLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDF 307
+ V ++F+ D +G G+ RG + S R TQ GLSLNID+S+T + PV+ F
Sbjct: 155 YIPVGRSFYSPDIGKKQSLGDGLESWRGFYQSIRPTQMGLSLNIDMSSTAFIEANPVIQF 214
Query: 308 L--IANQNVRD-PFS-LDWNKAKRTLKNLRITTSPTN---QEYKITGLSEMPCKDQLFTL 360
+ + N+++ P S D K K+ L+ +++ + ++Y+I+GL+ + ++ F +
Sbjct: 215 VCDLLNRDISSRPLSDADRVKIKKALRGVKVEVTHRGNMRRKYRISGLTAVATRELTFPV 274
Query: 361 KKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQ 420
+R + +V +YF +Q++ LPC+ VG RP ++P+E+C +V Q
Sbjct: 275 DERNT--------QKSVVEYFHETYGFRIQHT-QLPCLQVGNSNRPNYLPMEVCKIVEGQ 325
Query: 421 RYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDG 480
RY+K L+ Q ++L++ + Q+P +R + + ++ +DY + + GI I++ V+
Sbjct: 326 RYSKRLNERQITALLKVTCQRPIDREKDILQTVQLNDYAKDNYAQEFGIKISTSLASVEA 385
Query: 481 RVLQAPRLKF---GNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCDVRGLVRDL 537
R+L P LK+ G P+ G+WN NKK++ + W +NFS + L R
Sbjct: 386 RILPPPWLKYHESGREGTCLPQVGQWNMMNKKMINGGTVNNWICINFSRQVQ-DNLARTF 444
Query: 538 IKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLP------------GAP 585
+ E C+ F P L V E V+ L G
Sbjct: 445 CQ---------ELAQMCYVSGMAFNPEPVLPPVSARPEQVEKVLKTRYHDATSKLSQGKE 495
Query: 586 KFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRV---NDQYLTNVLLKINAKLGG 642
LL ++ N LYG K+ E GIV+QC V + QY+ NV LKIN K+GG
Sbjct: 496 IDLLIVILPDNNGSLYGDLKRICETELGIVSQCCLTKHVFKMSKQYMANVALKINVKVGG 555
Query: 643 MNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACV 702
N++L S IP+VS PT+I G DV+H PG+ PSIAAVV+S+ WP I+KY V
Sbjct: 556 RNTVLVDALSRRIPLVSDRPTIIFGADVTHPHPGEDSSPSIAAVVASQDWPEITKYAGLV 615
Query: 703 RTQGAKVEMIDNLFKPVSDTEDE----GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQ 758
Q + E+I +LFK D + G+I+ELLI F S+G+ KP II +RDGVSE Q
Sbjct: 616 CAQAHRQELIQDLFKEWKDPQKGVVTGGMIKELLIAFRRSTGH-KPLRIIFYRDGVSEGQ 674
Query: 759 FNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPD--------NVPPG 810
F QVL EL I +AC L+ + P +V QK HHT+ F D N+ PG
Sbjct: 675 FYQVLLYELDAIRKACASLEAGYQPPVTFVVVQKRHHTRLFAQNHNDRHSVDRSGNILPG 734
Query: 811 TVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRS 870
TVVD+KICHP +DFY+C+HAG+ GTSRP HYHVL DE F+ D LQ L ++L Y Y R
Sbjct: 735 TVVDSKICHPTEFDFYLCSHAGIQGTSRPAHYHVLWDENNFTADGLQSLTNNLCYTYARC 794
Query: 871 TTAISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGR------------DINASPI 918
T ++S+V P YAHLAA + +M+ E S + G R ++NA+
Sbjct: 795 TRSVSIVPPAYYAHLAAFRARFYMEPETSDSGSMASGSMARGGGMAGRSTRGPNVNAAVR 854
Query: 919 PQLPKLMDSVCNSMFF 934
P LP L ++V MF+
Sbjct: 855 P-LPALKENVKRVMFY 869
>At1g69440 hypothetical protein
Length = 990
Score = 363 bits (932), Expect = e-100
Identities = 269/891 (30%), Positives = 443/891 (49%), Gaps = 89/891 (9%)
Query: 74 GSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSE 133
G G+ + LL NHF V ++ + Y+V + P K R I ++ ET +
Sbjct: 158 GQDGSVIYLLANHFLVKFDSSQR-IYHYNVEI-----SPQPSKEIARMIKQKLVETDRNS 211
Query: 134 LNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTD-RKRL 192
+G A+DG + +++ ++LEF V L P N D R++
Sbjct: 212 FSGVVPAFDGRQNIYSPVEFQGDRLEFFVNLP-----------IPSCKAVMNYGDLREKQ 260
Query: 193 KKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQ----EAIRVLDIILRQHAAKQGC 248
+ K ++V + SK + + E E++ E I LD+ILR++ ++ C
Sbjct: 261 PQKKIEKLFRVNMKLVSKFDGKE-----QRKEGEDWAPLPPEYIHALDVILRENPMEK-C 314
Query: 249 LLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFL 308
+ ++F+ + ++GGG +G RG S R TQ GL+LN+D+S T V+ +L
Sbjct: 315 TSIGRSFYSSSMGGSKEIGGGAVGLRGFFQSLRHTQQGLALNMDLSITAFHESIGVIAYL 374
Query: 309 ---------IANQNVRDPFSLDWNKAKRTLKNLRITTS--PTNQEYKITGLSEMPCKDQL 357
+ R+ + + ++ LKN+R+ T Q Y++ GL+E ++
Sbjct: 375 QKRLEFLTDLPRNKGRELSLEEKREVEKALKNIRVFVCHRETVQRYRVYGLTEEITENIW 434
Query: 358 FTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLV 417
F + + + + + YF + +Q+ +LPC+ + + RP ++P+ELC +
Sbjct: 435 FP---------DREGKYLRLMSYFKDHYGYEIQFK-NLPCLQISRA-RPCYLPMELCMIC 483
Query: 418 SLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYG--SEPMLRNCGISITSGF 475
Q++ LS Q + +++ QKP ER ++ D + T G S R + ++
Sbjct: 484 EGQKFLGKLSDDQAAKIMKMGCQKPNERKAII-DKVMTGSVGPSSGNQTREFNLEVSREM 542
Query: 476 TQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCDVRGLVR 535
T + GR+LQ P+LK PRN K+ + +IE+WA+++ D + +
Sbjct: 543 TLLKGRILQPPKLKLDR-----PRN----LKESKVFKGTRIERWALMSIGGSSDQKSTIP 593
Query: 536 DLIKCGGMKGIHVEQPFDCFEENGQFRRAPPLVR----VEKMFEHVQSKLPGAPKFLLCL 591
I K H+ + F ++ +E + +Q + ++C+
Sbjct: 594 KFINELTQKCEHLGVFLSKNTLSSTFFEPSHILNNISLLESKLKEIQRAASNNLQLIICV 653
Query: 592 LSERKNSDLYGPWKKKNLAEFGIVTQC-IAP--TRVNDQYLTNVLLKINAKLGGMNSLLG 648
+ ++ YG K+ + G+VTQC + P T+++ Q+++N+ LKINAK+GG + L
Sbjct: 654 MEKKHKG--YGDLKRISETRIGVVTQCCLYPNITKLSSQFVSNLALKINAKIGGSMTELY 711
Query: 649 VEHSPSIPIVSKA--PTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQG 706
IP + + P + +G DV+H P PS+AAVV S WP ++Y + +R+Q
Sbjct: 712 NSIPSHIPRLLRPDEPVIFMGADVTHPHPFDDCSPSVAAVVGSINWPEANRYVSRMRSQT 771
Query: 707 AKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIE 766
+ E+I +L + +++ELL DFY + + P+ II FRDGVSE+QF +VL E
Sbjct: 772 HRQEIIQDL---------DLMVKELLDDFYKAV-KKLPNRIIFFRDGVSETQFKKVLQEE 821
Query: 767 LSQIIEAC-KFLDEKWNPKFLVIVAQKNHHTKFFQPGSPD--NVPPGTVVDNKICHPRNY 823
L I AC KF D +NP V QK HHT+ F+ PD N+PPGTVVD I HP+ +
Sbjct: 822 LQSIKTACSKFQD--YNPSITFAVVQKRHHTRLFRC-DPDHENIPPGTVVDTVITHPKEF 878
Query: 824 DFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYA 883
DFY+C+H G+ GTSRPTHYH+L DE F+ D+LQ LV++L Y + R T IS+V P YA
Sbjct: 879 DFYLCSHLGVKGTSRPTHYHILWDENEFTSDELQRLVYNLCYTFVRCTKPISIVPPAYYA 938
Query: 884 HLAASQVGQFMKFEDKSETSSSHGGSGRDINASPIPQLPKLMDSVCNSMFF 934
HLAA + +++ +S S + S + LPKL D+V N MF+
Sbjct: 939 HLAAYRGRLYIERSSESNGGSMNPSSVSRVGPPKTIPLPKLSDNVKNLMFY 989
>At1g31290 hypothetical protein
Length = 1194
Score = 325 bits (833), Expect = 7e-89
Identities = 276/937 (29%), Positives = 442/937 (46%), Gaps = 121/937 (12%)
Query: 54 EPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPV 113
+P KK P K P + KG + L NHF+V+ + T+ Y V +
Sbjct: 324 KPSSSDKKEPVKRPDKGGNIKVKGV-INLSVNHFRVSFS-TESVIRHYDV--------DI 373
Query: 114 EGKGAGRKI----LDRVQETYGSELN---GKDLAYDGEKTLFTIGSLAQNKLEFTVVLED 166
+G+ + +KI L V+E + N AYDG+K +F+ L +
Sbjct: 374 KGENSSKKISRFELAMVKEKLFKDNNDFPNAMTAYDGQKNIFSAVELPTGSFK------- 426
Query: 167 VTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETE 226
D ++ R ++Y I ++ L + + G T
Sbjct: 427 --------------------VDFSETEEIMRGRSYTFIIKQVKELKLLDLQAYIDGRSTF 466
Query: 227 NYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSG 286
++ ++ +D+++++H +K+ + V + FF + D G GV +G H + + T G
Sbjct: 467 IPRDVLQGMDVVMKEHPSKR-MITVGKRFFSTRLE--IDFGYGVGAAKGFHHTLKPTVQG 523
Query: 287 LSLNIDVSTTMIVHPGPVVDFL---IANQNVRDPFSLDWNKAKRTLKNLRITTS--PTNQ 341
LSL ++ S V+++L +N+R + + + L L++T T Q
Sbjct: 524 LSLCLNSSLLAFRKAISVIEYLKLYFGWRNIRQFKNCRPDDVVQELIGLKVTVDHRKTKQ 583
Query: 342 EYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVG 401
++ I GLS+ KD F G +I++ +YF + + + D+PC+N+G
Sbjct: 584 KFIIMGLSKDDTKDIKFDFIDHA---GNQPPRKISIVEYFKEKYGRDIDHK-DIPCLNLG 639
Query: 402 KPKRPTFVPVELCSLVSLQRYTKALSTLQRSS---LVEKSRQKPQERMRVLTDALKTSD- 457
K R FVP+E C+LV Q + K L R S L E S PQ+R+ + +K+SD
Sbjct: 640 KKGRENFVPMEFCNLVEGQIFPK--EKLYRDSAAWLKELSLVTPQQRLENINKMIKSSDG 697
Query: 458 -YGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF----GNG--EDFNPRNGRWNFNNKKI 510
G + ++ N G+ + T V+GRVL+AP LK GN E + +WN K +
Sbjct: 698 PRGGD-IIGNFGLRVDPNMTTVEGRVLEAPTLKLTDRRGNPIHEKLMSESNQWNLTTKGV 756
Query: 511 VQPVKIEKWAVVNFSARCDVRG-----LVRDLI-KCGGMKGIHVEQPFDC----FEENGQ 560
+ I+ WAV++F+A ++ V LI +C G+ G+ +E P C E
Sbjct: 757 TKGSIIKHWAVLDFTASESLKKKMPGYFVNKLIERCKGL-GMQMEAPIVCKTSSMETLYD 815
Query: 561 FRRAPPLVRVEKMFEHVQSKLPGA-PKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQC- 618
L+R + + GA P +LC ++ + D Y K + G+VTQC
Sbjct: 816 GNALEELLR--SVIDEASHNHGGACPTLVLCAMTGKH--DGYKTLKWIAETKLGLVTQCF 871
Query: 619 -----IAPTRVNDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHG 673
I V+DQYL N+ LKINAK+GG N L V++ S + + +G DV+H
Sbjct: 872 LTISAIKGETVSDQYLANLALKINAKVGGTNVEL-VDNIFSF-FKKEDKVMFIGADVNHP 929
Query: 674 SPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLI 733
+ PSI AVV + WP ++Y A V+ Q + E I + + LI
Sbjct: 930 AAHDNMSPSIVAVVGTLNWPEANRYAARVKAQSHRKEEIQGFGETCWE----------LI 979
Query: 734 DFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKN 793
+ ++ + ++P+ I+IFRDGVS+ QF+ VLN+EL + + F +NP+ VIVAQK
Sbjct: 980 EAHSQAPEKRPNKIVIFRDGVSDGQFDMVLNVELQNVKDV--FAKVGYNPQITVIVAQKR 1037
Query: 794 HHTKFFQPGSPD------NVPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLD 847
H T+FF + NVP GTVVD I HP YDFY+C+ G IGTS+PTHY+VL D
Sbjct: 1038 HQTRFFPATTSKDGRAKGNVPSGTVVDTTIIHPFEYDFYLCSQHGAIGTSKPTHYYVLSD 1097
Query: 848 EIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQ-----VGQFMKFEDKSET 902
EIGF+ + +Q+L+ L + + R T +++V P+ YA AAS+ MK K
Sbjct: 1098 EIGFNSNQIQKLIFDLCFTFTRCTKPVALVPPVSYADKAASRGRVYYEASLMKKNSKQSR 1157
Query: 903 SSSHGGSGRDINASPI----PQLPKLMDSVCNSMFFV 935
+S + ++S + ++ K+ + N MFFV
Sbjct: 1158 GASSSSASVASSSSSVTMEDKEIFKVHAGIENFMFFV 1194
>At1g31280 unknown protein
Length = 1013
Score = 325 bits (832), Expect = 1e-88
Identities = 270/937 (28%), Positives = 429/937 (44%), Gaps = 107/937 (11%)
Query: 45 PADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGA--KLPLLTNHFKVNVTNTDGYFFQYS 102
P E + ++P + K PM R G A ++ L NH+KVN N + Y
Sbjct: 138 PVRAEVMNLKPSVQVATSDRKEPMKRPDRGGVVAVRRVNLYVNHYKVNF-NPESVIRHYD 196
Query: 103 VALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFTIGSLAQN--KLEF 160
V + E + + D+V E AYDG+K +F+ L K+E+
Sbjct: 197 VEIKGEIPTKKVSRFELAMVRDKVFTDNPDEFPLAMTAYDGQKNIFSAVELPTGSYKVEY 256
Query: 161 TVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANAL 220
E R ++Y I + + L + +
Sbjct: 257 PKTEE------------------------------MRGRSYTFTIKQVNVLKLGDLKEYM 286
Query: 221 KGHETENYQEAIRVLDIILRQHAAKQGCLL-VRQNFFHNDPKNFTDVGGGVLGCRGLHSS 279
G + N ++ ++ +D+++++H +K C++ V ++FF + + D GV+ +G +
Sbjct: 287 TGRSSFNPRDVLQGMDVVMKEHPSK--CMITVGKSFFTRETEPDEDFRFGVIAAKGYRHT 344
Query: 280 FRTTQSGLSLNIDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNKAKRTLKNLRITTSPT 339
+ T GLSL +D S V+++L N D + L L++T +
Sbjct: 345 LKPTAQGLSLCLDYSVLAFRKAMSVIEYLKLYFNWSDMRQFRRRDVEEELIGLKVTVNHR 404
Query: 340 NQEYKIT--GLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPC 397
+ K+T GLS KD F L + G + + ++ +YF + + + D+PC
Sbjct: 405 KNKQKLTIVGLSMQNTKDIKFDLIDQ---EGNEPPRKTSIVEYFRIKYGRHIVHK-DIPC 460
Query: 398 INVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSS---LVEKSRQKPQERMRVLTDALK 454
+++GK R FVP+E C LV Q Y K L + S L + S PQ+R R + +K
Sbjct: 461 LDLGKNGRQNFVPMEFCDLVEGQIYPK--DNLDKDSALWLKKLSLVNPQQRQRNIDKMIK 518
Query: 455 TSDYGSE-PMLRNCGISITSGFTQVDGRVLQAPRLKFGNG-----EDFNPR-NGRWNFNN 507
+ S ++ N G+ + + T V+GRVL+AP LK E+ NPR N +WN
Sbjct: 519 ARNGPSGGEIIGNFGLKVDTNMTPVEGRVLKAPSLKLAERGRVVREEPNPRQNNQWNLMK 578
Query: 508 KKIVQPVKIEKWAVVNFSARCDVRGLVRDLI-----KCGGMKGIHVEQPF----DCFEEN 558
K + + ++ WAV++F+A + D + +C + G+ +E P E
Sbjct: 579 KGVTRGSIVKHWAVLDFTASERFNKMPNDFVDNLIDRCWRL-GMQMEAPIVYKTSRMETL 637
Query: 559 GQFRRAPPLVRVEKMFEHVQSKLPGA-PKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQ 617
L+R + + K GA P +LC +S + D Y K + G+VTQ
Sbjct: 638 SNGNAIEELLR--SVIDEASRKHGGARPTLVLCAMSRK--DDGYKTLKWIAETKLGLVTQ 693
Query: 618 CIAP---TRVNDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAP-TLILGMDVSHG 673
C T+ DQY N+ LK+NAK+GG N VE + K + +G DV+H
Sbjct: 694 CFLTGPATKGGDQYRANLALKMNAKVGGSN----VELMDTFSFFKKEDEVMFIGADVNHP 749
Query: 674 SPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLI 733
+ PSI AVV + WP ++Y A V Q + E I + L+
Sbjct: 750 AARDKMSPSIVAVVGTLNWPEANRYAARVIAQPHRKEEIQGFGDACLE----------LV 799
Query: 734 DFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKN 793
+ + ++P+ I+IFRDGVS++QF+ VLN+EL + F +NPK VIVAQK
Sbjct: 800 KAHVQATGKRPNKIVIFRDGVSDAQFDMVLNVELLDV--KLTFEKNGYNPKITVIVAQKR 857
Query: 794 HHTKFFQPGSPD-----NVPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDE 848
H T+FF + D NVP GTVVD K+ HP YDFY+C+H G IGTS+PTHY+ L DE
Sbjct: 858 HQTRFFPATNNDGSDKGNVPSGTVVDTKVIHPYEYDFYLCSHHGGIGTSKPTHYYTLWDE 917
Query: 849 IGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYA----------HLAASQVGQFMKFED 898
+GF+ D +Q+L+ + + + R T +S+V P+ YA H A+S+ F +
Sbjct: 918 LGFTSDQVQKLIFEMCFTFTRCTKPVSLVPPVYYADMVAFRGRMYHEASSREKNFKQPRG 977
Query: 899 KSETSSSHGGSGRDINASPIPQLPKLMDSVCNSMFFV 935
S +++S S + + KL + N MFFV
Sbjct: 978 ASTSAASLASSLSSLTIED-KAIFKLHAELENVMFFV 1013
>At4g00755 putative F-box protein
Length = 377
Score = 32.7 bits (73), Expect = 0.98
Identities = 22/61 (36%), Positives = 31/61 (50%), Gaps = 3/61 (4%)
Query: 168 TSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETEN 227
TSN NG +S G S DT + L+K HR+ +S I IA+A++ T+N
Sbjct: 65 TSNDRNGESSEAGSSSLMDT--RLLEKEHRAFALLARGCMSSPIE-SCIADAIRASSTDN 121
Query: 228 Y 228
Y
Sbjct: 122 Y 122
>At3g56150 PROBABLE EUKARYOTIC TRANSLATION INITIATION FACTOR 3
SUBUNIT 8
Length = 900
Score = 32.0 bits (71), Expect = 1.7
Identities = 21/95 (22%), Positives = 51/95 (53%), Gaps = 9/95 (9%)
Query: 150 IGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFAS 209
+GS ++++ ++ V + +V ++ N G +DTD KR+ K + K ++ E+++
Sbjct: 9 VGSESEDESDYEVEVNEVQNDDVNNRYLQSGSEDDDDTDTKRVVKPAKDKRFE-EMTYT- 66
Query: 210 KIPLQAIANALKGHE----TENYQEAIRVLDIILR 240
+ + NA+K ++ EN+ + + L+ ++R
Sbjct: 67 ---VDQMKNAMKINDWVSLQENFDKVNKQLEKVMR 98
>At4g30150 hypothetical protein
Length = 1966
Score = 31.6 bits (70), Expect = 2.2
Identities = 21/65 (32%), Positives = 31/65 (47%), Gaps = 5/65 (7%)
Query: 99 FQYSVALFYEDGRPV-----EGKGAGRKILDRVQETYGSELNGKDLAYDGEKTLFTIGSL 153
+Q+S AL + D + + EG G I++ + E + LN + EKT F SL
Sbjct: 1323 YQFSKALPFSDEKLISSEISEGTGQANLIIENLTEQAETLLNALRATFRDEKTAFKCESL 1382
Query: 154 AQNKL 158
NKL
Sbjct: 1383 ILNKL 1387
>At2g02070 putative C2H2-type zinc finger protein
Length = 602
Score = 31.2 bits (69), Expect = 2.8
Identities = 18/56 (32%), Positives = 27/56 (48%), Gaps = 1/56 (1%)
Query: 38 PPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTN 93
PPPPP A + P++ PQ K+ P + P + + + K + TN F V N
Sbjct: 33 PPPPPHHQAPLPPLEAPPQKKKRNQP-RTPNSDAEVIALSPKTLMATNRFICEVCN 87
>At1g31810 hypothetical protein
Length = 1201
Score = 31.2 bits (69), Expect = 2.8
Identities = 18/51 (35%), Positives = 22/51 (42%), Gaps = 7/51 (13%)
Query: 38 PPPPPIVPADIEPIKIEPQIVKKKLPTKVPMARRG-------LGSKGAKLP 81
PPPPP P P P + K +P P RG LG+KG+ P
Sbjct: 714 PPPPPPPPLSKTPAPPPPPLSKTPVPPPPPGLGRGTSSGPPPLGAKGSNAP 764
>At1g22420
Length = 480
Score = 31.2 bits (69), Expect = 2.8
Identities = 14/36 (38%), Positives = 20/36 (54%), Gaps = 2/36 (5%)
Query: 32 ENEECLPPPPPIVPADIEPIKIEPQIVKKKLPTKVP 67
+ +E +PPPP VPA+I P + I +PT P
Sbjct: 158 QQQESIPPPP--VPANITPPPVPANITPPPVPTSAP 191
>At4g11930 hypothetical protein
Length = 272
Score = 30.4 bits (67), Expect = 4.8
Identities = 23/67 (34%), Positives = 35/67 (51%), Gaps = 8/67 (11%)
Query: 349 SEMPCKDQLFTLKKRGAVPGEDDTEEITVYD----YFVNRRKISLQYSADLPCINVGKPK 404
+E+P KD F +KR PG DD E + + D +V R IS+ +A + ++ K K
Sbjct: 205 NEVPSKDTSFVPEKRNVGPGPDD-EVVVISDDEDEEYVGRDIISMARTAKVRIVDGRKVK 263
Query: 405 RPTFVPV 411
FVP+
Sbjct: 264 ---FVPL 267
>At2g46040 unknown protein
Length = 562
Score = 30.4 bits (67), Expect = 4.8
Identities = 21/79 (26%), Positives = 34/79 (42%), Gaps = 1/79 (1%)
Query: 91 VTNTDGYFFQYSVALFYEDGRPVEGKGAGRK-ILDRVQETYGSELNGKDLAYDGEKTLFT 149
V +G+ + GRP + GA K + + + Y GK+ A + F
Sbjct: 158 VARLNGFLSEVKKKYELRKGRPAKELGAELKWFISKTKRRYDKHHVGKESASNDAVKEFQ 217
Query: 150 IGSLAQNKLEFTVVLEDVT 168
LA+ +LE ++LE VT
Sbjct: 218 GSKLAERRLEQIMILESVT 236
>At3g57080 unknown protein
Length = 222
Score = 30.0 bits (66), Expect = 6.3
Identities = 26/89 (29%), Positives = 40/89 (44%), Gaps = 6/89 (6%)
Query: 181 HGSPNDTDRKRLKKSHRS-KTYKVEISF--ASKIPLQAIANALKGHETENYQEAIRVLDI 237
+G D DR R+ HRS T KV+I F S + + AI + + + QE I L +
Sbjct: 62 YGERPDVDRLRISALHRSDSTKKVKIVFFGTSMVKVNAIRSVVADILS---QETITGLIL 118
Query: 238 ILRQHAAKQGCLLVRQNFFHNDPKNFTDV 266
+L+ H Q + F + TD+
Sbjct: 119 VLQNHVTNQALKAIELFSFKVEIFQITDL 147
>At2g01210 putative receptor-like protein kinase
Length = 716
Score = 30.0 bits (66), Expect = 6.3
Identities = 16/64 (25%), Positives = 33/64 (51%), Gaps = 1/64 (1%)
Query: 682 SIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGI-IRELLIDFYNSSG 740
S A V + + L+ + C+ + +++D P ++TEDE + + ++ I NSS
Sbjct: 636 SPAVEVGTSEMDLVRWVQVCIEEKKPLCDVLDPCLAPEAETEDEIVAVLKIAISCVNSSP 695
Query: 741 NRKP 744
++P
Sbjct: 696 EKRP 699
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.137 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,136,793
Number of Sequences: 26719
Number of extensions: 1018582
Number of successful extensions: 2858
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 2744
Number of HSP's gapped (non-prelim): 31
length of query: 935
length of database: 11,318,596
effective HSP length: 109
effective length of query: 826
effective length of database: 8,406,225
effective search space: 6943541850
effective search space used: 6943541850
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC124972.13


