

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124971.1 + phase: 0
(397 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
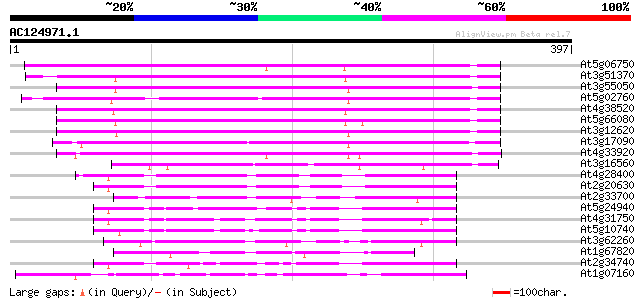
Score E
Sequences producing significant alignments: (bits) Value
At5g06750 protein phosphatase 2C-like 220 1e-57
At3g51370 protein phosphatase 2C like protein 211 7e-55
At3g55050 protein phosphatase 2C - like protein 207 1e-53
At5g02760 protein phosphatase - like protein 203 1e-52
At4g38520 putative protein phosphatase-2c 202 3e-52
At5g66080 protein phosphatase 2C-like protein 200 1e-51
At3g12620 unknown protein 195 4e-50
At3g17090 protein phosphatase-2c, putative 192 4e-49
At4g33920 unknown protein 183 2e-46
At3g16560 unknown protein 104 7e-23
At4g28400 protein phosphatase 2C like protein 87 1e-17
At2g20630 putative protein phosphatase 2C 87 2e-17
At2g33700 putative protein phosphatase 2C 82 4e-16
At5g24940 Protein phosphatase 2C-like protein 82 7e-16
At4g31750 unknown protein 80 2e-15
At5g10740 protein phosphatase 2C -like protein 80 2e-15
At3g62260 unknown protein 79 6e-15
At1g67820 protein phosphatase-like protein 78 1e-14
At2g34740 putative protein phosphatase 2C 77 1e-14
At1g07160 unknown protein 77 2e-14
>At5g06750 protein phosphatase 2C-like
Length = 393
Score = 220 bits (560), Expect = 1e-57
Identities = 130/347 (37%), Positives = 199/347 (56%), Gaps = 14/347 (4%)
Query: 11 RSGKGKDEEEEKKEKEKAEEEVHDNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFG 69
R G+ ++++ + + D+L+W + + + + SIA Q+N V+ED QVE G
Sbjct: 18 RYGRMNRDDDDDDDHDGDSSSSGDSLLWSRELERHSFGDFSIAVVQANEVIEDHSQVETG 77
Query: 70 KNSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVT 129
++FVGVYDGH G +A+R+I LF L R+ E +SE+ + A E+GF V
Sbjct: 78 NGAVFVGVYDGHGGPEASRYISDHLFSHLMRVSRERSCISEEALRAAFSATEEGFLTLVR 137
Query: 130 NNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVV---QLTVDH 186
+ +VGSCCL G+IW TL +ANVGDSRA+LGS R + + QLT DH
Sbjct: 138 RTCGLKPLIAAVGSCCLVGVIWKGTLLIANVGDSRAVLGSMGSNNNRSNKIVAEQLTSDH 197
Query: 187 HVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWS----DP-HPSF 241
+ + R+E+R+ +D ++ G R+K +I+++RSIGDAYLK DP P F
Sbjct: 198 NAALEEVRQELRSLHPDDSHIVVLKHGVWRIKGIIQVSRSIGDAYLKRPEFSLDPSFPRF 257
Query: 242 ETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKR 301
+ V+S +P R + SDKF+IFAS G W+ MTN +A +IV + + GI++R
Sbjct: 258 HLAEELQRPVLSAEPCVYTRVLQTSDKFVIFASDGLWEQMTNQQAVEIVNKHPRPGIARR 317
Query: 302 LVRAALEKAIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
LVR A+ A + Y +L+ ++ G RR ++DD+TV+VIF++
Sbjct: 318 LVRRAITIAAKKREMNYDDLKKVERG----VRRFFHDDITVVVIFID 360
>At3g51370 protein phosphatase 2C like protein
Length = 379
Score = 211 bits (536), Expect = 7e-55
Identities = 125/351 (35%), Positives = 199/351 (56%), Gaps = 28/351 (7%)
Query: 12 SGKGKDEEEEKKEKEKAEEEVHDNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGK 70
S GK + K+ D L+W +D ++ S+A Q+N ++ED QVE G
Sbjct: 17 SSSGKSSDSTGKQ---------DGLLWYKDFGQHLVGEFSMAVVQANNLLEDQSQVESGP 67
Query: 71 NSL--------FVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEK 122
S F+G+YDGH G + +RF+ LF L R E +S D++++A + E+
Sbjct: 68 LSTLDSGPYGTFIGIYDGHGGPETSRFVNDHLFQHLKRFAAEQASMSVDVIKKAYEATEE 127
Query: 123 GFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQL 182
GF VT ++ +VGSCCL G+I G L++ANVGDSRA+LG + +QL
Sbjct: 128 GFLGVVTKQWPTKPQIAAVGSCCLVGVICGGMLYIANVGDSRAVLGRAMKATGEVIALQL 187
Query: 183 TVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD-----P 237
+ +H+VS + R+E+ + +D ++ RVK LI+I+RSIGD YLK ++
Sbjct: 188 SAEHNVSIESVRQEMHSLHPDDSHIVMLKHNVWRVKGLIQISRSIGDVYLKKAEFNKEPL 247
Query: 238 HPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDG 297
+ + ++ ++S +P +I DKFLIFAS G W+ M+N EA DIV N+ ++G
Sbjct: 248 YTKYRIREPFKRPILSGEPTITEHEIQPQDKFLIFASDGLWEQMSNQEAVDIVQNHPRNG 307
Query: 298 ISKRLVRAALEKAIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
I++RLV+ AL++A + Y +L+ ++ G RRH++DD+TV++IFL+
Sbjct: 308 IARRLVKMALQEAAKKREMRYSDLKKIERG----VRRHFHDDITVVIIFLD 354
>At3g55050 protein phosphatase 2C - like protein
Length = 409
Score = 207 bits (526), Expect = 1e-53
Identities = 123/329 (37%), Positives = 190/329 (57%), Gaps = 19/329 (5%)
Query: 34 DNLVWCEDRKEN-DYHCSIATSQSNTVMEDFYQVEFGKNSL--------FVGVYDGHKGL 84
D L+W +D + S+A Q+N ++ED Q+E G SL FVGVYDGH G
Sbjct: 60 DGLLWYKDSGNHITGEFSMAVVQANNLLEDHSQLESGPISLHESGPEATFVGVYDGHGGP 119
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+AARF+ LF + R +E + +S D++ + E+ F V ++ SVG+C
Sbjct: 120 EAARFVNDRLFYNIKRYTSEQRGMSPDVITRGFVATEEEFLGLVQEQWKTKPQIASVGAC 179
Query: 145 CLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITND 204
CL GI+ L+VAN GDSR +LG FK VQL+ +H+ S + REE+R +D
Sbjct: 180 CLVGIVCNGLLYVANAGDSRVVLGKVANPFKELKAVQLSTEHNASIESVREELRLLHPDD 239
Query: 205 PFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH-----PSFETFSRYEANVISEKPFTD 259
P ++ RVK +I+++RSIGDAYLK ++ + P F R+E ++ +P
Sbjct: 240 PNIVVLKHKVWRVKGIIQVSRSIGDAYLKRAEFNQEPLLPKFRVPERFEKPIMRAEPTIT 299
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYC 318
I D+FLIFAS G W+ ++N EA DIV + ++G++++LV+AAL++A + Y
Sbjct: 300 VHKIHPEDQFLIFASDGLWEHLSNQEAVDIVNSCPRNGVARKLVKAALQEAAKKREMRYS 359
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ ++ G RRH++DD+TVIV+FL+
Sbjct: 360 DLEKIERGI----RRHFHDDITVIVVFLH 384
>At5g02760 protein phosphatase - like protein
Length = 361
Score = 203 bits (517), Expect = 1e-52
Identities = 130/354 (36%), Positives = 197/354 (54%), Gaps = 37/354 (10%)
Query: 9 CGRSGKGKDEEEEKKEKEKAEEEVHDNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVE 67
C R G G E+ K D L W +D + + S+A Q+N+VMED Q+E
Sbjct: 5 CWRIGAGM-------ERSKINPTKVDGLTWYKDLGLHTFGEFSMAMIQANSVMEDQCQIE 57
Query: 68 FGK--------NSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDF 119
G FVGVYDGH G +A+RFI +FP K +SE ++ +A
Sbjct: 58 SGPLTFNNPTVQGTFVGVYDGHGGPEASRFIADNIFP---------KEISEQVISKAFAE 108
Query: 120 IEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHV 179
+K F + VT + ++ SVGSCCL G+I +++AN GDSRA+LG S+ R
Sbjct: 109 TDKDFLKTVTKQWPTNPQMASVGSCCLAGVICNGLVYIANTGDSRAVLGRSERGGVR--A 166
Query: 180 VQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH- 238
VQL+V+H+ + +AR+E+ + NDP +L RVK +I++TRSIGDAYLK ++ +
Sbjct: 167 VQLSVEHNANLESARQELWSLHPNDPTILVMKHRLWRVKGVIQVTRSIGDAYLKRAEFNR 226
Query: 239 ----PSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNS 294
P F + ++S P + D+F+I AS G W+ ++N EA DIV+N+
Sbjct: 227 EPLLPKFRLPEHFTKPILSADPSVTITRLSPQDEFIILASDGLWEHLSNQEAVDIVHNSP 286
Query: 295 QDGISKRLVRAALEKAIND-IITYCNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+ GI++RL++AAL++A + Y +L + G RRH++DD+TVIV++LN
Sbjct: 287 RQGIARRLLKAALKEAAKKREMRYSDLTEIHPG----VRRHFHDDITVIVVYLN 336
>At4g38520 putative protein phosphatase-2c
Length = 400
Score = 202 bits (513), Expect = 3e-52
Identities = 117/329 (35%), Positives = 186/329 (55%), Gaps = 19/329 (5%)
Query: 34 DNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGKNS--------LFVGVYDGHKGL 84
+ L+W D ++ + S+A Q+N+++ED Q+E G S FVGVYDGH G
Sbjct: 32 EGLLWFRDSGQHVFGDFSMAVVQANSLLEDQSQLESGSLSSHDSGPFGTFVGVYDGHGGP 91
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+ +RFI +F L R E + +S +++++A E+GF VTN ++ +VGSC
Sbjct: 92 ETSRFINDHMFHHLKRFTAEQQCMSSEVIKKAFQATEEGFLSIVTNQFQTRPQIATVGSC 151
Query: 145 CLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITND 204
CL +I L+VAN GDSRA+LG H QL+ +H+ S + R E++ +
Sbjct: 152 CLVSVICDGKLYVANAGDSRAVLGQVMRVTGEAHATQLSAEHNASIESVRRELQALHPDH 211
Query: 205 PFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD-----PHPSFETFSRYEANVISEKPFTD 259
P ++ RVK +I+++RSIGD YLK S+ + F S + ++S +P
Sbjct: 212 PDIVVLKHNVWRVKGIIQVSRSIGDVYLKRSEFNREPLYAKFRLRSPFSKPLLSAEPAIT 271
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYC 318
++ D+F+I AS G W+ M+N EA DIV N+ ++GI+KRLV+ AL++A + Y
Sbjct: 272 VHTLEPHDQFIICASDGLWEHMSNQEAVDIVQNHPRNGIAKRLVKVALQEAAKKREMRYS 331
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ + G RRH++DD+TVIV+F +
Sbjct: 332 DLKKIDRG----VRRHFHDDITVIVVFFD 356
>At5g66080 protein phosphatase 2C-like protein
Length = 385
Score = 200 bits (508), Expect = 1e-51
Identities = 119/330 (36%), Positives = 189/330 (57%), Gaps = 20/330 (6%)
Query: 34 DNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGKNSL---------FVGVYDGHKG 83
D L+W +D + + S+A Q+N ++ED QVE G + FVGVYDGH G
Sbjct: 32 DGLLWYKDSAHHLFGDFSMAVVQANNLLEDQSQVESGPLTTLSSSGPYGTFVGVYDGHGG 91
Query: 84 LDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGS 143
+ +RF+ LF L R E +S D++ +A + E+GF V + +VGS
Sbjct: 92 PETSRFVNDHLFHHLKRFAAEQDSMSVDVIRKAYEATEEGFLGVVAKQWAVKPHIAAVGS 151
Query: 144 CCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITN 203
CCL G++ L+VANVGDSRA+LG + +QL+ +H+VS + R+E+ + +
Sbjct: 152 CCLIGVVCDGKLYVANVGDSRAVLGKVIKATGEVNALQLSAEHNVSIESVRQEMHSLHPD 211
Query: 204 DPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSD--PHPSFETFSRYE---ANVISEKPFT 258
D ++ RVK +I+++RSIGD YLK S+ P + + E ++S +P
Sbjct: 212 DSHIVVLKHNVWRVKGIIQVSRSIGDVYLKKSEFNKEPLYTKYRLREPMKRPILSWEPSI 271
Query: 259 DRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITY 317
D+ D+FLIFAS G W+ ++N EA +IV N+ ++GI++RLV+AAL++A + Y
Sbjct: 272 TVHDLQPDDQFLIFASDGLWEQLSNQEAVEIVQNHPRNGIARRLVKAALQEAAKKREMRY 331
Query: 318 CNLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L ++ G RRH++DD+TV+V+FL+
Sbjct: 332 SDLNKIERG----VRRHFHDDITVVVLFLD 357
>At3g12620 unknown protein
Length = 385
Score = 195 bits (495), Expect = 4e-50
Identities = 118/329 (35%), Positives = 186/329 (55%), Gaps = 19/329 (5%)
Query: 34 DNLVWCEDRKENDY-HCSIATSQSNTVMEDFYQVEFGKNSLF--------VGVYDGHKGL 84
D L+W +D + S++ Q+N ++ED ++E G S+F VGVYDGH G
Sbjct: 34 DGLLWYKDSGNHVAGEFSMSVIQANNLLEDHSKLESGPVSMFDSGPQATFVGVYDGHGGP 93
Query: 85 DAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSC 144
+AARF+ LF + + +EN +S +++ +A E+ F V ++ SVG+C
Sbjct: 94 EAARFVNKHLFDNIRKFTSENHGMSANVITKAFLATEEDFLSLVRRQWQIKPQIASVGAC 153
Query: 145 CLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITND 204
CL GII L++AN GDSR +LG + FK VQL+ +H+ S + REE+R+ ND
Sbjct: 154 CLVGIICSGLLYIANAGDSRVVLGRLEKAFKIVKAVQLSSEHNASLESVREELRSLHPND 213
Query: 205 PFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH-----PSFETFSRYEANVISEKPFTD 259
P ++ RVK +I+++RSIGDAYLK ++ + F + ++ +P
Sbjct: 214 PQIVVLKHKVWRVKGIIQVSRSIGDAYLKKAEFNREPLLAKFRVPEVFHKPILRAEPAIT 273
Query: 260 RRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAAL-EKAINDIITYC 318
I D+FLIFAS G W+ ++N EA DIV ++GI+++L++ AL E A + Y
Sbjct: 274 VHKIHPEDQFLIFASDGLWEHLSNQEAVDIVNTCPRNGIARKLIKTALREAAKKREMRYS 333
Query: 319 NLQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
+L+ + G RRH++DD+TVIV+FL+
Sbjct: 334 DLKKIDRG----VRRHFHDDITVIVVFLD 358
>At3g17090 protein phosphatase-2c, putative
Length = 384
Score = 192 bits (487), Expect = 4e-49
Identities = 120/328 (36%), Positives = 188/328 (56%), Gaps = 19/328 (5%)
Query: 31 EVHDNLVWCEDRKENDYHC----SIATSQSNTVMEDFYQVEFGKNSLFVGVYDGHKGLDA 86
E D L+W D + +C S+A Q+N V+ED QVE G FVGVYDGH G +A
Sbjct: 40 EGKDGLLWFRDLGK---YCGGDFSMAVIQANQVLEDQSQVESGNFGTFVGVYDGHGGPEA 96
Query: 87 ARFIRVCLFPELSRLVTENK-VVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCC 145
AR++ LF + E + VV+ + +E+A E+GF V+ + + +VG+CC
Sbjct: 97 ARYVCDHLFNHFREISAETQGVVTRETIERAFHATEEGFASIVSELWQEIPNLATVGTCC 156
Query: 146 LFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITNDP 205
L G+I+ TLFVA++GDSR +LG KG +QL+ +H+ ++ R E+++ +DP
Sbjct: 157 LVGVIYQNTLFVASLGDSRVVLG-KKGNCGGLSAIQLSTEHNANNEDIRWELKDLHPDDP 215
Query: 206 FVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPH-----PSFETFSRYEANVISEKPFTDR 260
++ G RVK +I+++RSIGD Y+K + + F + ++S P
Sbjct: 216 QIVVFRHGVWRVKGIIQVSRSIGDMYMKRPEFNKEPISQKFRIAEPMKRPLMSATPTILS 275
Query: 261 RDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAAL-EKAINDIITYCN 319
+ +D FLIFAS G W+ +TN +A +IV+N+ + G +KRL++AAL E A + Y +
Sbjct: 276 HPLHPNDSFLIFASDGLWEHLTNEKAVEIVHNHPRAGSAKRLIKAALHEAARKREMRYSD 335
Query: 320 LQNLKAGNGLLGRRHYYDDVTVIVIFLN 347
L+ + RRH++DD+TVIV+FLN
Sbjct: 336 LRKIDK----KVRRHFHDDITVIVVFLN 359
>At4g33920 unknown protein
Length = 380
Score = 183 bits (464), Expect = 2e-46
Identities = 116/327 (35%), Positives = 191/327 (57%), Gaps = 18/327 (5%)
Query: 34 DNLVWCEDRKEN---DYHCSIATSQSNTVMEDFYQVEFGKNSLFVGVYDGHKGLDAARFI 90
D L+W + + + DY SIA Q+N+ +ED QV ++ +VGVYDGH G +A+RF+
Sbjct: 20 DGLLWQSELRPHAGGDY--SIAVVQANSRLEDQSQVFTSSSATYVGVYDGHGGPEASRFV 77
Query: 91 RVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGII 150
LFP + + E+ +S D++++A E+ F V ++ ++ +VGSCCL G I
Sbjct: 78 NRHLFPYMHKFAREHGGLSVDVIKKAFKETEEEFCGMVKRSLPMKPQMATVGSCCLVGAI 137
Query: 151 WGRTLFVANVGDSRAILGS-SKGFFKRPHVV--QLTVDHHVSHAAAREEIRNHITNDPFV 207
TL+VAN+GDSRA+LGS G V +L+ DH+V+ R+E++ +D +
Sbjct: 138 SNDTLYVANLGDSRAVLGSVVSGVDSNKGAVAERLSTDHNVAVEEVRKEVKALNPDDSQI 197
Query: 208 LCKNRGSLRVKSLIEITRSIGDAYLKWSDPH--PSFETFSR---YEANVISEKPFTDRRD 262
+ RG R+K +I+++RSIGD YLK + + P F+ ++ +P R
Sbjct: 198 VLYTRGVWRIKGIIQVSRSIGDVYLKKPEYYRDPIFQRHGNPIPLRRPAMTAEPSIIVRK 257
Query: 263 IDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISKRLVRAALEKAIND-IITYCNLQ 321
+ D FLIFAS G W+ +++ A +IV + + GI++RLVRAALE+A + Y +++
Sbjct: 258 LKPQDLFLIFASDGLWEHLSDETAVEIVLKHPRTGIARRLVRAALEEAAKKREMRYGDIK 317
Query: 322 NLKAGNGLLGRRHYYDDVTVIVIFLNK 348
+ G RRH++DD++VIV++L++
Sbjct: 318 KIAKGI----RRHFHDDISVIVVYLDQ 340
>At3g16560 unknown protein
Length = 493
Score = 104 bits (260), Expect = 7e-23
Identities = 90/316 (28%), Positives = 138/316 (43%), Gaps = 51/316 (16%)
Query: 73 LFVGVYDGHKGLDAARFIRVCLFPE-------LSRLVTENKVVSE--------------- 110
LF +YDG G DAA F+ L+ L R + + K +
Sbjct: 174 LFCAIYDGFNGRDAADFLACTLYESIVFHLQLLDRQMKQTKSDDDGEKLELLSNISNVDY 233
Query: 111 -----------DIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVAN 159
D + +A+ E F V +++ + SVGSC L ++ G+ L+V N
Sbjct: 234 SSTDLFRQGVLDCLNRALFQAETDFLRMVEQEMEERPDLVSVGSCVLVTLLVGKDLYVLN 293
Query: 160 VGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKS 219
+GDSRA+L + G K+ VQLT DH V + + + +DP ++ ++K
Sbjct: 294 LGDSRAVLATYNG-NKKLQAVQLTEDHTVDNEVEEARLLSEHLDDPKIVIGG----KIKG 348
Query: 220 LIEITRSIGDAYLKWSDPHPSFETFSR----YEANVISEKPFTDRRDIDESDKFLIFASH 275
+++TR++G YLK + + R +S +P I ESD F+I AS
Sbjct: 349 KLKVTRALGVGYLKKEKLNDALMGILRVRNLLSPPYVSVEPSMRVHKITESDHFVIVASD 408
Query: 276 GFWKLMTNSEAADIVY-----NNSQDGISKRLVRAALEKAINDIITYCNLQNLKAGNGLL 330
G + +N EA +V+ N S D L R + A T L N+ AG
Sbjct: 409 GLFDFFSNEEAIGLVHSFVSSNPSGDPAKFLLERLVAKAAARAGFTLEELTNVPAGR--- 465
Query: 331 GRRHYYDDVTVIVIFL 346
RR Y+DDVT++VI L
Sbjct: 466 -RRRYHDDVTIMVITL 480
>At4g28400 protein phosphatase 2C like protein
Length = 283
Score = 87.4 bits (215), Expect = 1e-17
Identities = 76/278 (27%), Positives = 123/278 (43%), Gaps = 47/278 (16%)
Query: 47 YHCSIATSQSNTVMEDFYQVEF----GKNSLFVGVYDGHKGLDAARFIRVCLFPELSRLV 102
+HC +S+ MED+ EF G ++DGH G D A++++ LF
Sbjct: 38 FHC--VKGKSSHPMEDYVVSEFKKLEGHELGLFAIFDGHLGHDVAKYLQTNLF------- 88
Query: 103 TENKVVSEDIMEQAVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGI-IWGRTLFVANVG 161
+N + +D + I ++ + ++G GS + GI I G+ L VANVG
Sbjct: 89 -DNILKEKDFWTDTENAIRNAYRSTDAVILQQSLKLGKGGSTAVTGILIDGKKLVVANVG 147
Query: 162 DSRAILGSSKGFFKRPHVVQLTVDHHVSHAAAREEIRN-HITNDPFVLCKNRGSL-RVKS 219
DSRA++ K QL+VDH S E R ++N P G + RV
Sbjct: 148 DSRAVMS------KNGVAHQLSVDHEPSKEKKEIESRGGFVSNIP-------GDVPRVDG 194
Query: 220 LIEITRSIGDAYLKWSDPHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWK 279
+ + R+ GD LK +S +P + ID+ +F++FAS G WK
Sbjct: 195 QLAVARAFGDKSLKLH----------------LSSEPDITHQTIDDHTEFILFASDGIWK 238
Query: 280 LMTNSEAADIVYN-NSQDGISKRLVRAALEKAINDIIT 316
+++N EA D + + +K L+ A+ + D I+
Sbjct: 239 VLSNQEAVDAIKSIKDPHAAAKHLIEEAISRKSKDDIS 276
>At2g20630 putative protein phosphatase 2C
Length = 279
Score = 86.7 bits (213), Expect = 2e-17
Identities = 75/265 (28%), Positives = 118/265 (44%), Gaps = 45/265 (16%)
Query: 60 MEDFYQVEF----GKNSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQ 115
MED+ EF G + ++DGH G D A++++ LF +N + +D
Sbjct: 45 MEDYVVSEFKKVDGHDLGLFAIFDGHLGHDVAKYLQTNLF--------DNILKEKDFWTD 96
Query: 116 AVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGI-IWGRTLFVANVGDSRAILGSSKGFF 174
+ I + ++ ++G GS + GI I G+TL +ANVGDSRA++
Sbjct: 97 TKNAIRNAYISTDAVILEQSLKLGKGGSTAVTGILIDGKTLVIANVGDSRAVMS------ 150
Query: 175 KRPHVVQLTVDHHVSHAAAREEIRN-HITNDPFVLCKNRGSL-RVKSLIEITRSIGDAYL 232
K QL+VDH S E R ++N P G + RV + + R+ GD L
Sbjct: 151 KNGVASQLSVDHEPSKEQKEIESRGGFVSNIP-------GDVPRVDGQLAVARAFGDKSL 203
Query: 233 KWSDPHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYN 292
K +S P +ID +F++FAS G WK+M+N EA D++ +
Sbjct: 204 KIH----------------LSSDPDIRDENIDHETEFILFASDGVWKVMSNQEAVDLIKS 247
Query: 293 -NSQDGISKRLVRAALEKAINDIIT 316
+K L+ A+ K D I+
Sbjct: 248 IKDPQAAAKELIEEAVSKQSTDDIS 272
>At2g33700 putative protein phosphatase 2C
Length = 380
Score = 82.4 bits (202), Expect = 4e-16
Identities = 67/250 (26%), Positives = 110/250 (43%), Gaps = 41/250 (16%)
Query: 74 FVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIEKGFKEYVTNNID 133
F GV+DGH G DAA F+R + R + E+ + + K E+ D
Sbjct: 123 FYGVFDGHGGTDAAHFVR----KNILRFIVEDSSFPLCVKKAIKSAFLKADYEFA----D 174
Query: 134 DDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLTVDHHVSHAAA 193
D S G+ L I+GR L +AN GD RA+LG +R ++L+ DH + A
Sbjct: 175 DSSLDISSGTTALTAFIFGRRLIIANAGDCRAVLG------RRGRAIELSKDHKPNCTAE 228
Query: 194 REEIR--NHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPHPSFETFSRYEANV 251
+ I + D + + + + R+IGD ++K + A
Sbjct: 229 KVRIEKLGGVVYDGY----------LNGQLSVARAIGDWHMKG----------PKGSACP 268
Query: 252 ISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAA-----DIVYNNSQDGISKRLVRAA 306
+S +P D+ E D+FLI G W +M++ A +++ +N + S+ LVR A
Sbjct: 269 LSPEPELQETDLSEDDEFLIMGCDGLWDVMSSQCAVTIARKELMIHNDPERCSRELVREA 328
Query: 307 LEKAINDIIT 316
L++ D +T
Sbjct: 329 LKRNTCDNLT 338
>At5g24940 Protein phosphatase 2C-like protein
Length = 447
Score = 81.6 bits (200), Expect = 7e-16
Identities = 76/262 (29%), Positives = 114/262 (43%), Gaps = 39/262 (14%)
Query: 60 MEDFYQVEF----GKNSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQ 115
MEDF++ G+ GV+DGH G AA +++ LF S L+T K +S D
Sbjct: 46 MEDFFETRIDGIDGEIVGLFGVFDGHGGSRAAEYVKRHLF---SNLITHPKFIS-DTKSA 101
Query: 116 AVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFK 175
D E + + ++ GS I+ G L VANVGDSRA++ F
Sbjct: 102 IADAYTHTDSELLKS---ENSHTRDAGSTASTAILVGDRLLVANVGDSRAVICRGGNAFA 158
Query: 176 RPHVVQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWS 235
++ DH + RE I N FV+ G+ RV ++ ++R+ GD LK
Sbjct: 159 ------VSRDHKPDQSDERERIENA---GGFVMWA--GTWRVGGVLAVSRAFGDRLLK-- 205
Query: 236 DPHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYN-NS 294
+ P ID+S +FLI AS G W + +N EA +V
Sbjct: 206 --------------QYVVADPEIQEEKIDDSLEFLILASDGLWDVFSNEEAVAVVKEVED 251
Query: 295 QDGISKRLVRAALEKAINDIIT 316
+ +K+LV A+++ D IT
Sbjct: 252 PEESTKKLVGEAIKRGSADNIT 273
>At4g31750 unknown protein
Length = 311
Score = 80.5 bits (197), Expect = 2e-15
Identities = 73/263 (27%), Positives = 119/263 (44%), Gaps = 41/263 (15%)
Query: 60 MEDFYQVEF----GKNSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQ 115
MEDFY+ G+ GV+DGH G AA +++ LF S L+ K +S D
Sbjct: 46 MEDFYETRIDGVEGEIVGLFGVFDGHGGARAAEYVKQNLF---SNLIRHPKFIS-DTTAA 101
Query: 116 AVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFK 175
D + E++ + + GS S I+ G L VANVGDSRA++ +
Sbjct: 102 IADAYNQTDSEFLKSENSQNRDAGSTASTA---ILVGDRLLVANVGDSRAVI------CR 152
Query: 176 RPHVVQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWS 235
+ + ++ DH + R+ I + FV+ G+ RV ++ ++R+ GD LK
Sbjct: 153 GGNAIAVSRDHKPDQSDERQRIEDA---GGFVMWA--GTWRVGGVLAVSRAFGDRLLK-- 205
Query: 236 DPHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIV--YNN 293
+ P +D S +FLI AS G W +++N EA ++ +
Sbjct: 206 --------------QYVVADPEIQEEKVDSSLEFLILASDGLWDVVSNEEAVGMIKAIED 251
Query: 294 SQDGISKRLVRAALEKAINDIIT 316
++G +KRL+ A ++ D IT
Sbjct: 252 PEEG-AKRLMMEAYQRGSADNIT 273
>At5g10740 protein phosphatase 2C -like protein
Length = 354
Score = 80.1 bits (196), Expect = 2e-15
Identities = 78/262 (29%), Positives = 117/262 (43%), Gaps = 39/262 (14%)
Query: 60 MEDFYQVEF-GKNSLFVG---VYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQ 115
MEDF++ G N VG V+DGH G AA +++ LF S L+T K +S D
Sbjct: 46 MEDFFETRIDGINGEIVGLFGVFDGHGGARAAEYVKRHLF---SNLITHPKFIS-DTKSA 101
Query: 116 AVDFIEKGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFK 175
D E + + + GS S I+ G L VANVGDSRA++ S+G
Sbjct: 102 ITDAYNHTDSELLKSENSHNRDAGSTASTA---ILVGDRLVVANVGDSRAVI--SRG--- 153
Query: 176 RPHVVQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWS 235
+ ++ DH + RE I N FV+ G+ RV ++ ++R+ GD LK
Sbjct: 154 -GKAIAVSRDHKPDQSDERERIENA---GGFVMWA--GTWRVGGVLAVSRAFGDRLLK-- 205
Query: 236 DPHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYN-NS 294
+ P ID++ +FLI AS G W + +N A +V
Sbjct: 206 --------------QYVVADPEIQEEKIDDTLEFLILASDGLWDVFSNEAAVAMVKEVED 251
Query: 295 QDGISKRLVRAALEKAINDIIT 316
+ +K+LV A+++ D IT
Sbjct: 252 PEDSAKKLVGEAIKRGSADNIT 273
>At3g62260 unknown protein
Length = 383
Score = 78.6 bits (192), Expect = 6e-15
Identities = 74/261 (28%), Positives = 117/261 (44%), Gaps = 38/261 (14%)
Query: 67 EFGKNSLFVGVYDGHKGLDAARFIR---VCLFPELSRLVTENKVVSEDIMEQAVDFIEKG 123
E K S F V+DGH G +AA ++R + F E + + VS +E+ +
Sbjct: 109 ELPKPSAFYAVFDGHGGPEAAAYVRENAIRFFFEDEQF-PQTSEVSSVYVEEVETSLRNA 167
Query: 124 FKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQLT 183
F + +D S G+ L +I GR L VAN GD RA+L ++ + ++
Sbjct: 168 FLQADLALAEDCSISDSCGTTALTALICGRLLMVANAGDCRAVL------CRKGRAIDMS 221
Query: 184 VDHHVSHAAAR---EEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLKWSDPHPS 240
DH + R EE ITND + + ++ +TR++GD LK PH S
Sbjct: 222 EDHKPINLLERRRVEESGGFITNDGY----------LNEVLAVTRALGDWDLKL--PHGS 269
Query: 241 FETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIV-----YNNSQ 295
++ +ISE P + + E D+FL+ G W ++T+ EA IV +N
Sbjct: 270 -------QSPLISE-PEIKQITLTEDDEFLVIGCDGIWDVLTSQEAVSIVRRGLNRHNDP 321
Query: 296 DGISKRLVRAALEKAINDIIT 316
++ LV AL + D +T
Sbjct: 322 TRCARELVMEALGRNSFDNLT 342
>At1g67820 protein phosphatase-like protein
Length = 447
Score = 77.8 bits (190), Expect = 1e-14
Identities = 66/230 (28%), Positives = 108/230 (46%), Gaps = 49/230 (21%)
Query: 74 FVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDI-------MEQAVDFIEKGFKE 126
F GVYDGH G AA F+ L + ++ K E + + DF+EK KE
Sbjct: 134 FFGVYDGHGGAKAAEFVAENLHKYVVEMMENCKGKEEKVEAFKAAFLRTDRDFLEKVIKE 193
Query: 127 YVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQ-LTVD 185
+ G+CC+ +I + + V+N+GD RA+L R V + LT D
Sbjct: 194 QSLKGVVS-------GACCVTAVIQDQEMIVSNLGDCRAVL-------CRAGVAEALTDD 239
Query: 186 HHVSHAAAREEIRNHITNDPFV--------LCKNRGSLRVKSLIEITRSIGDAYL-KWSD 236
H +E I + + PF+ + ++G+ RV+ ++ ++RSIGDA+L KW
Sbjct: 240 HKPGRDDEKERIESQ-SLIPFMTFGLQGGYVDNHQGAWRVQGILAVSRSIGDAHLKKW-- 296
Query: 237 PHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEA 286
V++E P T ++++ +FL+ AS G W +++N EA
Sbjct: 297 --------------VVAE-PETRVLELEQDMEFLVLASDGLWDVVSNQEA 331
>At2g34740 putative protein phosphatase 2C
Length = 239
Score = 77.4 bits (189), Expect = 1e-14
Identities = 78/265 (29%), Positives = 115/265 (42%), Gaps = 45/265 (16%)
Query: 60 MEDFYQVEF----GKNSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQ 115
MEDF + G N ++DGH G D A +++ LF +N + D
Sbjct: 1 MEDFIVADTKTVKGHNLGLYAIFDGHSGSDVADYLQNHLF--------DNILSQPDFWRN 52
Query: 116 AVDFIEKGFK---EYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKG 172
I++ +K +Y+ N+ R GS + +I G+ + VANVGDSRAIL
Sbjct: 53 PKKAIKRAYKSTDDYILQNVVGP-RGGSTAVTAI--VIDGKKIVVANVGDSRAILCRESD 109
Query: 173 FFKRPHVVQLTVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYL 232
K Q+TVDH R+ +++ FV K RV + +TR+ GD L
Sbjct: 110 VVK-----QITVDHEPDKE--RDLVKS---KGGFVSQKPGNVPRVDGQLAMTRAFGDGGL 159
Query: 233 KWSDPHPSFETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAAD-IVY 291
K IS P + +I + KFLI AS G WK+M+N E D I
Sbjct: 160 K----------------EHISVIPNIEIAEIHDDTKFLILASDGLWKVMSNDEVWDQIKK 203
Query: 292 NNSQDGISKRLVRAALEKAINDIIT 316
+ + +K L+ AL + D I+
Sbjct: 204 RGNAEEAAKMLIDKALARGSKDDIS 228
>At1g07160 unknown protein
Length = 380
Score = 77.0 bits (188), Expect = 2e-14
Identities = 86/323 (26%), Positives = 138/323 (42%), Gaps = 50/323 (15%)
Query: 5 PFLTCGRSGKGKDEEEEKKEKEKAEEEVHDNLVWCEDRKEN---DYHCSIATSQSNTVME 61
P G + + + ++E E E V+C+ K D +I Q +
Sbjct: 93 PVAPVGIAAPISNADTPREESRAVEREGDGYSVYCKRGKREAMEDRFSAITNLQGDP--- 149
Query: 62 DFYQVEFGKNSLFVGVYDGHKGLDAARFIRVCLFPELSRLVTENKVVSEDIMEQAVDFIE 121
K ++F GVYDGH G AA F L + + + +E +E+AV +
Sbjct: 150 --------KQAIF-GVYDGHGGPTAAEFAAKNLCSNILGEIVGGR--NESKIEEAV---K 195
Query: 122 KGFKEYVTNNIDDDGRVGSVGSCCLFGIIWGRTLFVANVGDSRAILGSSKGFFKRPHVVQ 181
+G+ + + + G GSCC+ +I L VAN GD RA+L S GF +
Sbjct: 196 RGYLATDSEFLKEKNVKG--GSCCVTALISDGNLVVANAGDCRAVL-SVGGFAE-----A 247
Query: 182 LTVDHHVSHAAAREEIRNHITNDPFVLCKNRGSLRVKSLIEITRSIGDAYLK-WSDPHPS 240
LT DH S R++ RN I + + R++ + ++R IGDA+LK W P
Sbjct: 248 LTSDHRPS----RDDERNRIESSGGYVDTFNSVWRIQGSLAVSRGIGDAHLKQWIISEP- 302
Query: 241 FETFSRYEANVISEKPFTDRRDIDESDKFLIFASHGFWKLMTNSEAADIVYNNSQDGISK 300
E N++ I+ +FLI AS G W ++N EA DI + K
Sbjct: 303 -------EINILR---------INPQHEFLILASDGLWDKVSNQEAVDIARPFCKGTDQK 346
Query: 301 RLVRAALEKAINDIITYCNLQNL 323
R A +K ++ ++ +L ++
Sbjct: 347 RKPLLACKKLVDLSVSRGSLDDI 369
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.137 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,276,349
Number of Sequences: 26719
Number of extensions: 416667
Number of successful extensions: 3412
Number of sequences better than 10.0: 106
Number of HSP's better than 10.0 without gapping: 81
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 2987
Number of HSP's gapped (non-prelim): 207
length of query: 397
length of database: 11,318,596
effective HSP length: 101
effective length of query: 296
effective length of database: 8,619,977
effective search space: 2551513192
effective search space used: 2551513192
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC124971.1


