

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124970.4 + phase: 0 /pseudo
(204 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
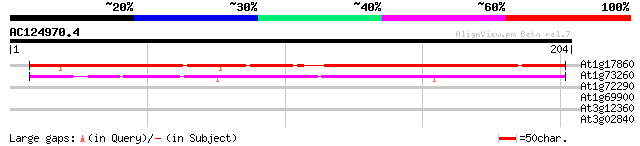
At1g17860 Unknown protein (F2H15.9) 143 5e-35
At1g73260 putative trypsin inhibitor (At1g73260) 84 7e-17
At1g72290 drought induced protein, putative 37 0.009
At1g69900 unknown protein 28 3.3
At3g12360 unknown protein 27 5.7
At3g02840 unknown protein 27 7.4
>At1g17860 Unknown protein (F2H15.9)
Length = 196
Score = 143 bits (361), Expect = 5e-35
Identities = 79/200 (39%), Positives = 122/200 (60%), Gaps = 16/200 (8%)
Query: 8 LFLLLTLFTT-KPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKT 66
+FLLL +F + + + E V+D +G + +NY+ILP G M+ L T +T
Sbjct: 7 IFLLLAVFISHRGVTTEAAVEPVKDINGKSLLTGVNYYILPVIRGRGGGLTMSNLKT-ET 65
Query: 67 CPLDVVEEE----EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVD 122
CP V++++ + + F P++ K I VSTD+N+ S PT+ +W++ D
Sbjct: 66 CPTSVIQDQFEVSQGLPVKFSPYD-KSRTIPVSTDVNIKFS-PTS-------IWELANFD 116
Query: 123 VATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIFLDE 182
T Q F++T GV+GNPG++TVDNWFKI++FE YK+ FCPTVC C+V+C+D+G+F+ +
Sbjct: 117 ETTKQWFISTCGVEGNPGQKTVDNWFKIDKFEKDYKIRFCPTVCNFCKVICRDVGVFVQD 176
Query: 183 NRNTRFVLSDFPFGVKFQRA 202
+ R LSD P V F+RA
Sbjct: 177 GKR-RLALSDVPLKVMFKRA 195
>At1g73260 putative trypsin inhibitor (At1g73260)
Length = 215
Score = 83.6 bits (205), Expect = 7e-17
Identities = 64/200 (32%), Positives = 92/200 (46%), Gaps = 13/200 (6%)
Query: 8 LFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTC 67
L L LT GA V D GN + + +Y++LP G +A + C
Sbjct: 14 LVLALTAVLASNAYGA-----VVDIDGNAMFHE-SYYVLPVIRGRGGGLTLAGRG-GQPC 66
Query: 68 PLDVVEE----EEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDV 123
P D+V+E +E + F + K + S +LN+ S T W+V + D
Sbjct: 67 PYDIVQESSEVDEGIPVKFSNWRLKVAFVPESQNLNIETDVGATICIQS-TYWRVGEFDH 125
Query: 124 ATSQRFVTTGGVQGNPGRETVDNWFKIERF-ESGYKLVFCPTVCRECEVVCKDIGIFLDE 182
Q FV G G++++ ++FKIE+ E YK VFCP C C D+GIF+DE
Sbjct: 126 ERKQYFVVAGPKPEGFGQDSLKSFFKIEKSGEDAYKFVFCPRTCDSGNPKCSDVGIFIDE 185
Query: 183 NRNTRFVLSDFPFGVKFQRA 202
R LSD PF V F++A
Sbjct: 186 LGVRRLALSDKPFLVMFKKA 205
>At1g72290 drought induced protein, putative
Length = 215
Score = 36.6 bits (83), Expect = 0.009
Identities = 42/166 (25%), Positives = 67/166 (40%), Gaps = 20/166 (12%)
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQC--GTRCEM 58
MK + FL++ LF +P V+DT+GN + YFI P + G
Sbjct: 1 MKNPSVISFLIILLFAATICTHGNEP--VKDTAGNPLNTREQYFIQPVKTESKNGGGLVP 58
Query: 59 ALLNTNKTCPLDVVEE----EEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTN----CST 110
A + CPL + + + + SFV + S+ +N+ F +N C
Sbjct: 59 AAITVLPFCPLGITQTLLPYQPGLPVSFVLALGVGSTVMTSSAVNI--EFKSNIWPFCKE 116
Query: 111 SSVTVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESG 156
S W+VD A + + GG G+ ++ FKIE+ G
Sbjct: 117 FS-KFWEVDDSSSAPKEPSILIGGKMGDR-----NSSFKIEKAGEG 156
>At1g69900 unknown protein
Length = 397
Score = 28.1 bits (61), Expect = 3.3
Identities = 11/25 (44%), Positives = 14/25 (56%)
Query: 99 NVIHSFPTNCSTSSVTVWKVDKVDV 123
+V H P +T S VW VD VD+
Sbjct: 369 SVTHDLPNRSATQSSVVWDVDVVDI 393
>At3g12360 unknown protein
Length = 590
Score = 27.3 bits (59), Expect = 5.7
Identities = 20/56 (35%), Positives = 26/56 (45%), Gaps = 1/56 (1%)
Query: 89 KGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRETV 144
K V +S +L +H N +T+SVTV V VA + F GG N G V
Sbjct: 398 KNVHNISKELRKLHREGINNATNSVTVVAVLFATVAFAAIFTVPGG-DNNDGSAVV 452
>At3g02840 unknown protein
Length = 379
Score = 26.9 bits (58), Expect = 7.4
Identities = 21/65 (32%), Positives = 30/65 (45%), Gaps = 10/65 (15%)
Query: 56 CEMALLNTNKTCPLDVVEEEEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTV 115
CE +L+ N TC + +E+ VP KK ++RVS T CS S +
Sbjct: 252 CERSLVVLNATCDNEQGKEDVLRNALIVPLLVKK-ILRVS-------DLATQCSVS--IL 301
Query: 116 WKVDK 120
WK+ K
Sbjct: 302 WKLWK 306
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.139 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,730,427
Number of Sequences: 26719
Number of extensions: 192476
Number of successful extensions: 505
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 498
Number of HSP's gapped (non-prelim): 7
length of query: 204
length of database: 11,318,596
effective HSP length: 95
effective length of query: 109
effective length of database: 8,780,291
effective search space: 957051719
effective search space used: 957051719
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124970.4


