

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124966.3 + phase: 0
(470 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
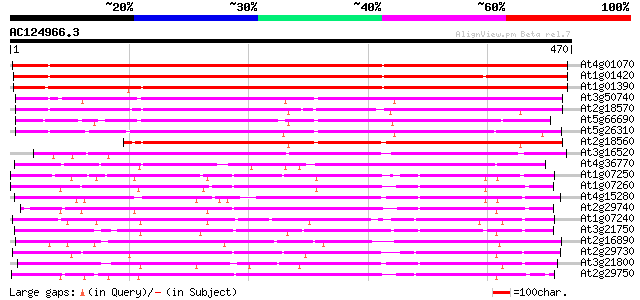
Score E
Sequences producing significant alignments: (bits) Value
At4g01070 putative flavonol glucosyltransferase 484 e-137
At1g01420 flavonol 3-o-glucosyltransferase, putative 456 e-128
At1g01390 flavonol 3-o-glucosyltransferase, putative 448 e-126
At3g50740 UTP-glucose glucosyltransferase - like protein 305 4e-83
At2g18570 putative flavonol 3-O-glucosyltransferase 301 5e-82
At5g66690 UTP-glucose glucosyltransferase 286 2e-77
At5g26310 UTP-glucose glucosyltransferase - like protein 283 1e-76
At2g18560 flavonol 3-O-glucosyltransferase like protein 271 6e-73
At3g16520 glucosyltransferase like protein 271 7e-73
At4g36770 glucosyltransferase-like protein 251 8e-67
At1g07250 putative 3-o-glucosyltransferase 226 2e-59
At1g07260 unknown protein 206 3e-53
At4g15280 UTP-glucose glucosyltransferase 205 4e-53
At2g29740 putative flavonol 3-O-glucosyltransferase 205 4e-53
At1g07240 unknown protein 202 2e-52
At3g21750 putative UDP-glucose glucosyltransferase 200 2e-51
At2g16890 glucosyltransferase like protein 200 2e-51
At2g29730 putative flavonol 3-O-glucosyltransferase 199 4e-51
At3g21800 putative UDP-glucose glucosyltransferase 198 5e-51
At2g29750 putative flavonol 3-O-glucosyltransferase 198 6e-51
>At4g01070 putative flavonol glucosyltransferase
Length = 480
Score = 484 bits (1247), Expect = e-137
Identities = 243/466 (52%), Positives = 323/466 (69%), Gaps = 3/466 (0%)
Query: 3 KTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNI 62
KT H+A++P G HL+P+++F+KRLV LH VT I G PS A +++L +LPS+I
Sbjct: 5 KTPHVAIIPSPGMGHLIPLVEFAKRLVHLH-GLTVTFVIAGEGPPSKAQRTVLDSLPSSI 63
Query: 63 NHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPL-VALVVDAFSVEV 121
+ FLPPV+ DL T +ES+I LT+T S P L + S + L ALVVD F +
Sbjct: 64 SSVFLPPVDLTDLSSSTRIESRISLTVTRSNPELRKVFDSFVEGGRLPTALVVDLFGTDA 123
Query: 122 LNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFL 181
++ E ++ YI++P+ A L++ +LPKLDE SCE+R++ EP+ +PGCVP+ G+DFL
Sbjct: 124 FDVAVEFHVPPYIFYPTTANVLSFFLHLPKLDETVSCEFRELTEPLMLPGCVPVAGKDFL 183
Query: 182 SIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPII 241
AQDR AYK L K A+G+LVN+F E+E + A++E G D PPVYPVGP++
Sbjct: 184 DPAQDRKDDAYKWLLHNTKRYKEAEGILVNTFFELEPNAIKALQEPGLDKPPVYPVGPLV 243
Query: 242 ETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWV 301
+ ECL WLD Q SVLYVSFGSGGTL+ EQ+ ELALGL S +FLWV
Sbjct: 244 NIGKQEAKQTEESECLKWLDNQPLGSVLYVSFGSGGTLTCEQLNELALGLADSEQRFLWV 303
Query: 302 LRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFL 361
+R+PS ++S+ Y + + D L FLP GFLERTK++GFVI WAPQ Q+L+ S GGFL
Sbjct: 304 IRSPSGIANSS-YFDSHSQTDPLTFLPPGFLERTKKRGFVIPFWAPQAQVLAHPSTGGFL 362
Query: 362 THCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKV 421
THCGWNSTLESVV G+PLI WPL+AEQKMNAVLLSE ++ LR ++G+V R EVA+V
Sbjct: 363 THCGWNSTLESVVSGIPLIAWPLYAEQKMNAVLLSEDIRAALRPRAGDDGLVRREEVARV 422
Query: 422 IKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
+K LMEG+EG+ +RN MKELKEAA +K+DG+STK +S +ALKW+
Sbjct: 423 VKGLMEGEEGKGVRNKMKELKEAACRVLKDDGTSTKALSLVALKWK 468
>At1g01420 flavonol 3-o-glucosyltransferase, putative
Length = 481
Score = 456 bits (1172), Expect = e-128
Identities = 227/465 (48%), Positives = 322/465 (68%), Gaps = 4/465 (0%)
Query: 4 TVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNIN 63
T H+A++P G HL+P+++ +KRL+ H F VT IP PS A +S+L +LPS+I
Sbjct: 6 TPHVAIIPSPGIGHLIPLVELAKRLLDNH-GFTVTFIIPGDSPPSKAQRSVLNSLPSSIA 64
Query: 64 HTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVA-LVVDAFSVEVL 122
FLPP + +D+P +E++I LT+T S P L + SL+ E L A LVVD F +
Sbjct: 65 SVFLPPADLSDVPSTARIETRISLTVTRSNPALRELFGSLSAEKRLPAVLVVDLFGTDAF 124
Query: 123 NIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLS 182
++ E ++ YI++ S A L + +LPKLDE SCE+R++ EP+ IPGCVP+ G+DF+
Sbjct: 125 DVAAEFHVSPYIFYASNANVLTFLLHLPKLDETVSCEFRELTEPVIIPGCVPITGKDFVD 184
Query: 183 IAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPIIE 242
QDR ++YK L VK A+G+LVNSF+++E + ++E D PPVY +GP++
Sbjct: 185 PCQDRKDESYKWLLHNVKRFKEAEGILVNSFVDLEPNTIKIVQEPAPDKPPVYLIGPLVN 244
Query: 243 TETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVL 302
+ + D + +CL WLD Q SVLYVSFGSGGTL+ EQ +ELALGL S +FLWV+
Sbjct: 245 SGSHDADVNDEYKCLNWLDNQPFGSVLYVSFGSGGTLTFEQFIELALGLAESGKRFLWVI 304
Query: 303 RAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLT 362
R+PS +SS+ Y + ++ D FLP GFL+RTKEKG V+ SWAPQ QIL+ S+GGFLT
Sbjct: 305 RSPSGIASSS-YFNPQSRNDPFSFLPQGFLDRTKEKGLVVGSWAPQAQILTHTSIGGFLT 363
Query: 363 HCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKVI 422
HCGWNS+LES+V+GVPLI WPL+AEQKMNA+LL + + LRA + E+G+V R EVA+V+
Sbjct: 364 HCGWNSSLESIVNGVPLIAWPLYAEQKMNALLLVD-VGAALRARLGEDGVVGREEVARVV 422
Query: 423 KYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
K L+EG+EG +R MKELKE + +++DG STK++++++LKW+
Sbjct: 423 KGLIEGEEGNAVRKKMKELKEGSVRVLRDDGFSTKSLNEVSLKWK 467
>At1g01390 flavonol 3-o-glucosyltransferase, putative
Length = 480
Score = 448 bits (1152), Expect = e-126
Identities = 226/466 (48%), Positives = 312/466 (66%), Gaps = 5/466 (1%)
Query: 4 TVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNIN 63
T HIA++P G HL+P ++ +KRLVQ H F VT I SPS A +S+L +LPS+I
Sbjct: 6 TPHIAIMPSPGMGHLIPFVELAKRLVQ-HDCFTVTMIISGETSPSKAQRSVLNSLPSSIA 64
Query: 64 HTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQ--GLKSLAKEIPLVALVVDAFSVEV 121
FLPP + +D+P +E++ +LT+T S P L + G S K +P V LVVD F +
Sbjct: 65 SVFLPPADLSDVPSTARIETRAMLTMTRSNPALRELFGSLSTKKSLPAV-LVVDMFGADA 123
Query: 122 LNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFL 181
++ + ++ YI++ S A L++ +LPKLD+ SCE+R + EP+KIPGCVP+ G+DFL
Sbjct: 124 FDVAVDFHVSPYIFYASNANVLSFFLHLPKLDKTVSCEFRYLTEPLKIPGCVPITGKDFL 183
Query: 182 SIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPII 241
QDR+ AYK L K A G+LVNSF+++E + A++E D P VYP+GP++
Sbjct: 184 DTVQDRNDDAYKLLLHNTKRYKEAKGILVNSFVDLESNAIKALQEPAPDKPTVYPIGPLV 243
Query: 242 ETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWV 301
T + + + + CL+WLD Q SVLY+SFGSGGTL+ EQ ELA+GL S +F+WV
Sbjct: 244 NTSSSNVNLEDKFGCLSWLDNQPFGSVLYISFGSGGTLTCEQFNELAIGLAESGKRFIWV 303
Query: 302 LRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFL 361
+R+PS SS+ Y + ++ D FLP GFL+RTKEKG V+ SWAPQ+QIL+ S GFL
Sbjct: 304 IRSPSEIVSSS-YFNPHSETDPFSFLPIGFLDRTKEKGLVVPSWAPQVQILAHPSTCGFL 362
Query: 362 THCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKV 421
THCGWNSTLES+V+GVPLI WPLFAEQKMN +LL E + LR E+GIV R EV +V
Sbjct: 363 THCGWNSTLESIVNGVPLIAWPLFAEQKMNTLLLVEDVGAALRIHAGEDGIVRREEVVRV 422
Query: 422 IKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
+K LMEG+EG+ + N +KELKE + +DG S+K+ ++ LKW+
Sbjct: 423 VKALMEGEEGKAIGNKVKELKEGVVRVLGDDGLSSKSFGEVLLKWK 468
>At3g50740 UTP-glucose glucosyltransferase - like protein
Length = 487
Score = 305 bits (781), Expect = 4e-83
Identities = 174/472 (36%), Positives = 283/472 (59%), Gaps = 22/472 (4%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLP----SN 61
H+A+ G H++P+++ KRL H F VT F+ L + + + +S P +
Sbjct: 7 HVAMFASPGMGHIIPVIELGKRLAGSH-GFDVTIFV--LETDAASAQSQFLNSPGCDAAL 63
Query: 62 INHTFLPPVNPNDLPQGTTMES-QILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
++ LP + + L + ++L+ + ++P + ++ + + AL+VD F ++
Sbjct: 64 VDIVGLPTPDISGLVDPSAFFGIKLLVMMRETIPTIRSKIEEMQHKP--TALIVDLFGLD 121
Query: 121 VLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF 180
+ +G E NML+YI+ S A LA + P LD++ E+ +P+ +PGC P+ D
Sbjct: 122 AIPLGGEFNMLTYIFIASNARFLAVALFFPTLDKDMEEEHIIKKQPMVMPGCEPVRFEDT 181
Query: 181 LSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGG----DNPPVYP 236
L D +SQ Y+ F+PF + + DG++VN++ ++E L ++++ PVYP
Sbjct: 182 LETFLDPNSQLYREFVPFGSVFPTCDGIIVNTWDDMEPKTLKSLQDPKLLGRIAGVPVYP 241
Query: 237 VGPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNT 296
+GP+ S + L+ WL+KQ SVLY+SFGSGG+LS +Q+ ELA GLE+S
Sbjct: 242 IGPLSRPVDPSKTNHPVLD---WLNKQPDESVLYISFGSGGSLSAKQLTELAWGLEMSQQ 298
Query: 297 KFLWVLRAPSSSSSSAGYLSAENDI---DTLQFLPSGFLERTKEKGFVITSWAPQIQILS 353
+F+WV+R P S+ + YLSA + T +LP GF+ RT E+GF+++SWAPQ +IL+
Sbjct: 299 RFVWVVRPPVDGSACSAYLSANSGKIRDGTPDYLPEGFVSRTHERGFMVSSWAPQAEILA 358
Query: 354 PNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRA-SVNENGI 412
+VGGFLTHCGWNS LESVV GVP+I WPLFAEQ MNA LL+E L V +R+ + G+
Sbjct: 359 HQAVGGFLTHCGWNSILESVVGGVPMIAWPLFAEQMMNATLLNEELGVAVRSKKLPSEGV 418
Query: 413 VERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGS-STKTISQIA 463
+ R E+ +++ +M +EG ++R +K+LKE A+ ++ DG + +++S+IA
Sbjct: 419 ITRAEIEALVRKIMVEEEGAEMRKKIKKLKETAAESLSCDGGVAHESLSRIA 470
>At2g18570 putative flavonol 3-O-glucosyltransferase
Length = 470
Score = 301 bits (771), Expect = 5e-82
Identities = 187/471 (39%), Positives = 281/471 (58%), Gaps = 25/471 (5%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPS-NATKSILQTLPSNINH 64
H +V G HL+PIL+ RL + N HVT T GS S T++I I
Sbjct: 5 HALLVASPGLGHLIPILELGNRLSSVL-NIHVTILAVTSGSSSPTETEAIHAAAARTICQ 63
Query: 65 -TFLPPVNPNDLPQ-GTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVL 122
T +P V+ ++L + T+ +++++ + P + +K L K P V ++VD E++
Sbjct: 64 ITEIPSVDVDNLVEPDATIFTKMVVKMRAMKPAVRDAVK-LMKRKPTV-MIVDFLGTELM 121
Query: 123 NIGKELNMLS-YIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFL 181
++ ++ M + Y+Y P+ A LA YLP LD EY DI EP+KIPGC P+ ++ +
Sbjct: 122 SVADDVGMTAKYVYVPTHAWFLAVMVYLPVLDTVVEGEYVDIKEPLKIPGCKPVGPKELM 181
Query: 182 SIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNP----PVYPV 237
DRS Q YK + + +DGVLVN++ E++ L+A++E+ + PVYP+
Sbjct: 182 ETMLDRSGQQYKECVRAGLEVPMSDGVLVNTWEELQGNTLAALREDEELSRVMKVPVYPI 241
Query: 238 GPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTK 297
GPI+ T + D N + WLD+Q+ SV++V GSGGTL+ EQ VELALGLELS +
Sbjct: 242 GPIVRTN-QHVDKPNSI--FEWLDEQRERSVVFVCLGSGGTLTFEQTVELALGLELSGQR 298
Query: 298 FLWVLRAPSSSSSSAGYLSA--ENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPN 355
F+WVLR P+S YL A +D LP GFL+RT+ G V+T WAPQ++ILS
Sbjct: 299 FVWVLRRPAS------YLGAISSDDEQVSASLPEGFLDRTRGVGIVVTQWAPQVEILSHR 352
Query: 356 SVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRAS-VNENGIVE 414
S+GGFL+HCGW+S LES+ GVP+I WPL+AEQ MNA LL+E + V +R S + ++
Sbjct: 353 SIGGFLSHCGWSSALESLTKGVPIIAWPLYAEQWMNATLLTEEIGVAVRTSELPSERVIG 412
Query: 415 RVEVAKVIKYLM--EGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIA 463
R EVA +++ +M E +EG+K+R +E++ ++ A +DGSS ++ + A
Sbjct: 413 REEVASLVRKIMAEEDEEGQKIRAKAEEVRVSSERAWSKDGSSYNSLFEWA 463
>At5g66690 UTP-glucose glucosyltransferase
Length = 481
Score = 286 bits (731), Expect = 2e-77
Identities = 173/466 (37%), Positives = 262/466 (56%), Gaps = 38/466 (8%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHT 65
H A+ G H++P+++ KRL + FHVT F+ + S +K + T ++
Sbjct: 7 HAAMFSSPGMGHVIPVIELGKRL-SANNGFHVTVFVLETDAASAQSKFLNST---GVDIV 62
Query: 66 FLPP------VNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSV 119
LP V+P+D + ++I + + ++P L + ++ ++ AL+VD F
Sbjct: 63 KLPSPDIYGLVDPDD-----HVVTKIGVIMRAAVPALRSKIAAMHQKP--TALIVDLFGT 115
Query: 120 EVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRD 179
+ L + KE NMLSY++ P+ A L Y P LD++ E+ P+ IPGC P+ D
Sbjct: 116 DALCLAKEFNMLSYVFIPTNARFLGVSIYYPNLDKDIKEEHTVQRNPLAIPGCEPVRFED 175
Query: 180 FLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNP------- 232
L Y+ F+ ADG+LVN++ E+E L ++ NP
Sbjct: 176 TLDAYLVPDEPVYRDFVRHGLAYPKADGILVNTWEEMEPKSLKSLL-----NPKLLGRVA 230
Query: 233 --PVYPVGPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALG 290
PVYP+GP+ S D L+ WL++Q SVLY+SFGSGG LS +Q+ ELA G
Sbjct: 231 RVPVYPIGPLCRPIQSSETDHPVLD---WLNEQPNESVLYISFGSGGCLSAKQLTELAWG 287
Query: 291 LELSNTKFLWVLRAPSSSSSSAGYLSAEN---DIDTLQFLPSGFLERTKEKGFVITSWAP 347
LE S +F+WV+R P S + Y+SA + +T ++LP GF+ RT ++GFV+ SWAP
Sbjct: 288 LEQSQQRFVWVVRPPVDGSCCSEYVSANGGGTEDNTPEYLPEGFVSRTSDRGFVVPSWAP 347
Query: 348 QIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASV 407
Q +ILS +VGGFLTHCGW+STLESVV GVP+I WPLFAEQ MNA LLS+ L + +R
Sbjct: 348 QAEILSHRAVGGFLTHCGWSSTLESVVGGVPMIAWPLFAEQNMNAALLSDELGIAVRLDD 407
Query: 408 NENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDG 453
+ I R ++ +++ +M EGE +R +K+L+++A ++ DG
Sbjct: 408 PKEDI-SRWKIEALVRKVMTEKEGEAMRRKVKKLRDSAEMSLSIDG 452
>At5g26310 UTP-glucose glucosyltransferase - like protein
Length = 481
Score = 283 bits (725), Expect = 1e-76
Identities = 170/467 (36%), Positives = 265/467 (56%), Gaps = 20/467 (4%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHT 65
H A+ G H++P+++ +KRL H FHVT F+ + S +K + T +N
Sbjct: 7 HAAMFSSPGMGHVLPVIELAKRLSANH-GFHVTVFVLETDAASVQSKLLNSTGVDIVN-- 63
Query: 66 FLPPVNPNDLPQGTT-MESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNI 124
LP + + L + ++I + + ++P L K +A AL++D F + L +
Sbjct: 64 -LPSPDISGLVDPNAHVVTKIGVIMREAVPTLRS--KIVAMHQNPTALIIDLFGTDALCL 120
Query: 125 GKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIA 184
ELNML+Y++ S A L Y P LDE E+ +P+ IPGC P+ D +
Sbjct: 121 AAELNMLTYVFIASNARYLGVSIYYPTLDEVIKEEHTVQRKPLTIPGCEPVRFEDIMDAY 180
Query: 185 QDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEG----GDNPPVYPVGPI 240
Y + ADG+LVN++ E+E L ++++ PVYPVGP+
Sbjct: 181 LVPDEPVYHDLVRHCLAYPKADGILVNTWEEMEPKSLKSLQDPKLLGRVARVPVYPVGPL 240
Query: 241 IETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLW 300
S D + WL+KQ SVLY+SFGSGG+L+ +Q+ ELA GLE S +F+W
Sbjct: 241 CRPIQSSTTDHPVFD---WLNKQPNESVLYISFGSGGSLTAQQLTELAWGLEESQQRFIW 297
Query: 301 VLRAPSSSSSSAGYLSAENDI---DTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSV 357
V+R P SS + Y SA+ + +T ++LP GF+ RT ++GF+I SWAPQ +IL+ +V
Sbjct: 298 VVRPPVDGSSCSDYFSAKGGVTKDNTPEYLPEGFVTRTCDRGFMIPSWAPQAEILAHQAV 357
Query: 358 GGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVE 417
GGFLTHCGW+STLESV+ GVP+I WPLFAEQ MNA LLS+ L + +R + + R +
Sbjct: 358 GGFLTHCGWSSTLESVLCGVPMIAWPLFAEQNMNAALLSDELGISVRVD-DPKEAISRSK 416
Query: 418 VAKVIKYLMEGDEGEKLRNNMKELKEAA--SNAVKEDGSSTKTISQI 462
+ +++ +M DEGE++R +K+L++ A S ++ GS+ +++ ++
Sbjct: 417 IEAMVRKVMAEDEGEEMRRKVKKLRDTAEMSLSIHGGGSAHESLCRV 463
>At2g18560 flavonol 3-O-glucosyltransferase like protein
Length = 380
Score = 271 bits (693), Expect = 6e-73
Identities = 159/375 (42%), Positives = 227/375 (60%), Gaps = 16/375 (4%)
Query: 96 LHQGLKSLAKEIPLVALVVDAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEE 155
+ +KS+ K+ P V ++VD F +L+I Y+Y PS A LA YLP LD+
Sbjct: 8 VRDAVKSM-KQKPTV-MIVDFFGTALLSITDVGVTSKYVYIPSHAWFLALIVYLPVLDKV 65
Query: 156 TSCEYRDILEPIKIPGCVPLHGRDFLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLE 215
EY DI EP+KIPGC P+ ++ L DRS Q Y+ + + +DGVLVN++ E
Sbjct: 66 MEGEYVDIKEPMKIPGCKPVGPKELLDTMLDRSDQQYRDCVQIGLEIPMSDGVLVNTWGE 125
Query: 216 IEMGPLSAMKEEGGDNP----PVYPVGPIIETETKSGDDANGLECLAWLDKQQPCSVLYV 271
++ L+A++E+ N PVYP+GPI+ T + E WLDKQ+ SV+YV
Sbjct: 126 LQGKTLAALREDIDLNRVIKVPVYPIGPIVRTNVLIEKPNSTFE---WLDKQEERSVVYV 182
Query: 272 SFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGF 331
GSGGTLS EQ +ELA GLELS FLWVLR P S + S+++D LP GF
Sbjct: 183 CLGSGGTLSFEQTMELAWGLELSCQSFLWVLRKPPSYLGA----SSKDDDQVSDGLPEGF 238
Query: 332 LERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMN 391
L+RT+ G V+T WAPQ++ILS S+GGFL+HCGW+S LES+ GVP+I WPL+AEQ MN
Sbjct: 239 LDRTRGVGLVVTQWAPQVEILSHRSIGGFLSHCGWSSVLESLTKGVPIIAWPLYAEQWMN 298
Query: 392 AVLLSEGLKVGLRAS-VNENGIVERVEVAKVIKYLM--EGDEGEKLRNNMKELKEAASNA 448
A LL+E + + +R S + ++ R EVA ++K ++ E EG K++ +E++ ++ A
Sbjct: 299 ATLLTEEIGMAIRTSELPSKKVISREEVASLVKKIVAEEDKEGRKIKTKAEEVRVSSERA 358
Query: 449 VKEDGSSTKTISQIA 463
GSS ++ + A
Sbjct: 359 WTHGGSSHSSLFEWA 373
>At3g16520 glucosyltransferase like protein
Length = 443
Score = 271 bits (692), Expect = 7e-73
Identities = 157/454 (34%), Positives = 248/454 (54%), Gaps = 22/454 (4%)
Query: 21 ILQFSKRLVQLHPNF--HVTCFIPTLGSPSNAT--KSILQTLPSNINHTFLPPVNPNDLP 76
+++ K ++ +P+ H+ P S AT S+ + PS H LP V P
Sbjct: 1 MVELGKTILSKNPSLSIHIILVPPPYQPESTATYISSVSSSFPSITFH-HLPAVTPYSSS 59
Query: 77 QGTTM--ESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNIGKELNMLSYI 134
+ ES +L L S P +H+ L SL++ + A+++D F VL+I + Y
Sbjct: 60 STSRHHHESLLLEILCFSNPSVHRTLFSLSRNFNVRAMIIDFFCTAVLDITADFTFPVYF 119
Query: 135 YFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIAQDRSSQAYKH 194
++ S A LA+ FYLP +DE T + + + IPG P+ G D +R + Y
Sbjct: 120 FYTSGAACLAFSFYLPTIDETTPGKNLKDIPTVHIPGVPPMKGSDMPKAVLERDDEVYDV 179
Query: 195 FLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGP-IIETETKSGDDANG 253
F+ F K LS + G+++N+F +E + A+ EE +YP+GP I+ + +D
Sbjct: 180 FIMFGKQLSKSSGIIINTFDALENRAIKAITEELCFRN-IYPIGPLIVNGRIEDRNDNKA 238
Query: 254 LECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAG 313
+ CL WLD Q SV+++ FGS G S+EQ++E+A+GLE S +FLWV+R P +
Sbjct: 239 VSCLNWLDSQPEKSVVFLCFGSLGLFSKEQVIEIAVGLEKSGQRFLWVVRNPPELEKT-- 296
Query: 314 YLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESV 373
++D LP GFL RT++KG V+ SWAPQ+ +L+ +VGGF+THCGWNS LE+V
Sbjct: 297 ------ELDLKSLLPEGFLSRTEDKGMVVKSWAPQVPVLNHKAVGGFVTHCGWNSILEAV 350
Query: 374 VHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKVIKYLMEGDEGE- 432
GVP++ WPL+AEQ+ N V++ + +K+ + + +E G V EV K ++ ++ GE
Sbjct: 351 CAGVPMVAWPLYAEQRFNRVMIVDEIKIAISMNESETGFVSSTEVEKRVQEII----GEC 406
Query: 433 KLRNNMKELKEAASNAVKEDGSSTKTISQIALKW 466
+R +K AA A+ E GSS ++ + W
Sbjct: 407 PVRERTMAMKNAAELALTETGSSHTALTTLLQSW 440
>At4g36770 glucosyltransferase-like protein
Length = 457
Score = 251 bits (640), Expect = 8e-67
Identities = 162/464 (34%), Positives = 253/464 (53%), Gaps = 39/464 (8%)
Query: 5 VHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINH 64
+H A+V G H VPIL+ K L+ H VT F+ T + +KS++ +
Sbjct: 3 LHGALVASPGMGHAVPILELGKHLLNHHGFDRVTVFLVT--DDVSRSKSLIGKTLMEEDP 60
Query: 65 TFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEI---PLVALVVDAFSVEV 121
F+ P D+ G + +L L + +KS E+ P V VVD E
Sbjct: 61 KFVIRFIPLDV-SGQDLSGSLLTKLAEMMRKALPEIKSSVMELEPRPRV-FVVDLLGTEA 118
Query: 122 LNIGKELNML-SYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF 180
L + KEL ++ ++ ++A LA+ Y+ LD++ + + + IPGC P
Sbjct: 119 LEVAKELGIMRKHVLVTTSAWFLAFTVYMASLDKQELYKQLSSIGALLIPGCSP------ 172
Query: 181 LSIAQDRSSQAYKHFLPFVKL------LSSADGVLVNSFLEIEMGPLSAMKEEGG----- 229
+ +R+ K+ + + +ADGV VN++ +E + + +
Sbjct: 173 --VKFERAQDPRKYIRELAESQRIGDEVITADGVFVNTWHSLEQVTIGSFLDPENLGRVM 230
Query: 230 DNPPVYPVGPII---ETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVE 286
PVYPVGP++ E K G L WLD Q SV+YV G G L+ EQ E
Sbjct: 231 RGVPVYPVGPLVRPAEPGLKHG-------VLDWLDLQPKESVVYVLLGVVGALTFEQTNE 283
Query: 287 LALGLELSNTKFLWVLRAPSSSSSSAGYLS-AENDIDTLQFLPSGFLERTKEKGFVITSW 345
LA GLEL+ +F+WV+R P+ SA +N+ + L FLP+GFL+RTK+ G V+ +W
Sbjct: 284 LAYGLELTGHRFVWVVRPPAEDDPSASMFDKTKNETEPLDFLPNGFLDRTKDIGLVVRTW 343
Query: 346 APQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRA 405
APQ +IL+ S GGF+THCGWNS LES+V+GVP++ WPL++EQKMNA ++S LK+ L+
Sbjct: 344 APQEEILAHKSTGGFVTHCGWNSVLESIVNGVPMVAWPLYSEQKMNARMVSGELKIALQI 403
Query: 406 SVNENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAV 449
+V +GIV++ +A+++K +M+ +EG+++R N+KELK+ A A+
Sbjct: 404 NV-ADGIVKKEVIAEMVKRVMDEEEGKEMRKNVKELKKTAEEAL 446
>At1g07250 putative 3-o-glucosyltransferase
Length = 479
Score = 226 bits (577), Expect = 2e-59
Identities = 154/482 (31%), Positives = 254/482 (51%), Gaps = 46/482 (9%)
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNA-----TKSIL 55
M K + +P H++ ++F+KRL+ L H T I L SPS+ +S++
Sbjct: 1 MVKETELIFIPVPSTGHILVHIEFAKRLINLDHRIH-TITILNLSSPSSPHASVFARSLI 59
Query: 56 QTLPSNINHTFLPPVN---PNDLPQGTTMESQILLTLTNSLPYLHQGLKSL-------AK 105
+ P H LPP+ P DL Q E+ I+ + + P + + S+ +
Sbjct: 60 ASQPKIRLHD-LPPIQDPPPFDLYQRAP-EAYIVKLIKKNTPLIKDAVSSIVASRRGGSD 117
Query: 106 EIPLVALVVDAFSVEVL-NIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDIL 164
+ + LV+D F ++ ++G ELN+ SYIY A L Y+P + + E+ D+
Sbjct: 118 SVQVAGLVLDLFCNSLVKDVGNELNLPSYIYLTCNARYLGMMKYIPDRHRKIASEF-DLS 176
Query: 165 ---EPIKIPGCVPLHGRDFLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPL 221
E + +PG + F+ + +AY+ ++ + A G+LVNSF E+E P
Sbjct: 177 SGDEELPVPGFINAIPTKFMPPGLF-NKEAYEAYVELAPRFADAKGILVNSFTELEPHPF 235
Query: 222 SAMKEEGGDNPPVYPVGPIIETETKSGDDANGLE---CLAWLDKQQPCSVLYVSFGSGGT 278
PPVYPVGPI+ + ++ + ++ + WLD Q SV+++ FGS G+
Sbjct: 236 DYFSHLE-KFPPVYPVGPILSLKDRASPNEEAVDRDQIVGWLDDQPESSVVFLCFGSRGS 294
Query: 279 LSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEK 338
+ + Q+ E+A LEL +FLW +R ++ ND+ LP GF+ R +
Sbjct: 295 VDEPQVKEIARALELVGCRFLWSIRTSGDVETNP------NDV-----LPEGFMGRVAGR 343
Query: 339 GFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSE- 397
G V WAPQ+++L+ ++GGF++HCGWNSTLES+ GVP+ TWP++AEQ++NA L +
Sbjct: 344 GLVC-GWAPQVEVLAHKAIGGFVSHCGWNSTLESLWFGVPVATWPMYAEQQLNAFTLVKE 402
Query: 398 -GLKVGLRASV--NENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGS 454
GL V LR + G+V E+A+ ++ LM+G G++ R +KE+ +AA A+ + GS
Sbjct: 403 LGLAVDLRMDYVSSRGGLVTCDEIARAVRSLMDG--GDEKRKKVKEMADAARKALMDGGS 460
Query: 455 ST 456
S+
Sbjct: 461 SS 462
>At1g07260 unknown protein
Length = 476
Score = 206 bits (523), Expect = 3e-53
Identities = 152/478 (31%), Positives = 239/478 (49%), Gaps = 41/478 (8%)
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFI----PTLGSPSNATKSILQ 56
M+ I V HL+ ++F+K L++ H + P KS++
Sbjct: 1 MKAEAEIIFVTYPSPGHLLVSIEFAKSLIKRDDRIHTITILYWALPLAPQAHLFAKSLVA 60
Query: 57 TLPSNINHTFLPPV-NPNDLPQG-TTMESQILLTLTNSLPYLHQGLKSLAKE------IP 108
+ P I LP V NP L E+ IL + ++P + L +L +
Sbjct: 61 SQP-RIRLLALPDVQNPPPLELFFKAPEAYILESTKKTVPLVRDALSTLVSSRKESGSVR 119
Query: 109 LVALVVDAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYR----DIL 164
+V LV+D F V ++ + ELN+ SYI+ A L+ YLP+ T+ E ++
Sbjct: 120 VVGLVIDFFCVPMIEVANELNLPSYIFLTCNAGFLSMMKYLPERHRITTSELDLSSGNVE 179
Query: 165 EPIKIPGCVPLHGRDFLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAM 224
PI PG V L ++Y+ ++ + A G+LVNS +E
Sbjct: 180 HPI--PGYVCSVPTKVLPPGLF-VRESYEAWVEIAEKFPGAKGILVNSVTCLEQNAFDYF 236
Query: 225 KEEGGDNPPVYPVGPIIETETKSGDDANGLE---CLAWLDKQQPCSVLYVSFGSGGTLSQ 281
+ PPVYPVGP++ + + + + + + WL+ Q S++Y+ FGS G + +
Sbjct: 237 ARLDENYPPVYPVGPVLSLKDRPSPNLDASDRDRIMRWLEDQPESSIVYICFGSLGIIGK 296
Query: 282 EQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFV 341
QI E+A LEL+ +FLW +R + +S LP GFL+RT KG V
Sbjct: 297 LQIEEIAEALELTGHRFLWSIRTNPTEKASP-----------YDLLPEGFLDRTASKGLV 345
Query: 342 ITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSE--GL 399
WAPQ+++L+ ++GGF++HCGWNS LES+ GVP+ TWP++AEQ++NA + + GL
Sbjct: 346 C-DWAPQVEVLAHKALGGFVSHCGWNSVLESLWFGVPIATWPMYAEQQLNAFSMVKELGL 404
Query: 400 KVGLRAS-VNENG-IVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSS 455
V LR V+ G IV+ E+A I+ LM+G++ R +KE+ EAA NA+ + GSS
Sbjct: 405 AVELRLDYVSAYGEIVKAEEIAGAIRSLMDGEDTP--RKRVKEMAEAARNALMDGGSS 460
>At4g15280 UTP-glucose glucosyltransferase
Length = 478
Score = 205 bits (522), Expect = 4e-53
Identities = 154/493 (31%), Positives = 241/493 (48%), Gaps = 63/493 (12%)
Query: 5 VHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFI-PTLGSPSNATKSI--LQTLPSN 61
+ + +P G HL P ++ +K+L+ +T I P+ +A+ I L TL +
Sbjct: 3 IELVFIPLPGIGHLRPTVKLAKQLIGSENRLSITIIIIPSRFDAGDASACIASLTTLSQD 62
Query: 62 -------INHTFLPPVN-PNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALV 113
I+ PP + P+ +P +E Q + + LA V
Sbjct: 63 DRLHYESISVAKQPPTSDPDPVPAQVYIEKQKTKVRDAVAARIVDPTRKLA------GFV 116
Query: 114 VDAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEE------------TSCEYR 161
VD F ++++ E + Y+ + S AT L ++ ++ ++ T E+
Sbjct: 117 VDMFCSSMIDVANEFGVPCYMVYTSNATFLGTMLHVQQMYDQKKYDVSELENSVTELEFP 176
Query: 162 DILEPIKIPGCVP--LHGRDFL--SIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIE 217
+ P + C+P L +++L S+AQ R + K G+LVN+ E+E
Sbjct: 177 SLTRPYPVK-CLPHILTSKEWLPLSLAQARCFRKMK-------------GILVNTVAELE 222
Query: 218 MGPLSAMKEEGGDNPPVYPVGPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGG 277
L G D P VYPVGP++ E + DD E L WLD+Q SV+++ FGS G
Sbjct: 223 PHALKMFNINGDDLPQVYPVGPVLHLENGNDDDEKQSEILRWLDEQPSKSVVFLCFGSLG 282
Query: 278 TLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTL-QFLPSGFLERTK 336
++EQ E A+ L+ S +FLW LR S + + D L + LP GFLERT
Sbjct: 283 GFTEEQTRETAVALDRSGQRFLWCLRHASPNIKT----DRPRDYTNLEEVLPEGFLERTL 338
Query: 337 EKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLS 396
++G VI WAPQ+ +L ++GGF+THCGWNS LES+ GVP++TWPL+AEQK+NA +
Sbjct: 339 DRGKVI-GWAPQVAVLEKPAIGGFVTHCGWNSILESLWFGVPMVTWPLYAEQKVNAFEMV 397
Query: 397 E--GLKVGLRASV------NENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNA 448
E GL V +R + E V ++ + I+ +ME D +RNN+KE+ E A
Sbjct: 398 EELGLAVEIRKYLKGDLFAGEMETVTAEDIERAIRRVMEQD--SDVRNNVKEMAEKCHFA 455
Query: 449 VKEDGSSTKTISQ 461
+ + GSS + +
Sbjct: 456 LMDGGSSKAALEK 468
>At2g29740 putative flavonol 3-O-glucosyltransferase
Length = 474
Score = 205 bits (522), Expect = 4e-53
Identities = 152/469 (32%), Positives = 239/469 (50%), Gaps = 44/469 (9%)
Query: 10 VPGVGYSHLVPILQFSKRLVQLHPNFHVTCFI-----PTLGSPSNATKSILQTL---PSN 61
+PG H++ ++ +KRL+ P+ T I P L P + T + L++L S
Sbjct: 16 IPG----HILATIELAKRLISHQPSRIHTITILHWSLPFL--PQSDTIAFLKSLIETESR 69
Query: 62 INHTFLPPV-NPNDLPQGT-TMESQILLTLTNSLPYLHQGLKSL------AKEIPLVALV 113
I LP V NP + ES IL + +P + L +L + + + LV
Sbjct: 70 IRLITLPDVQNPPPMELFVKASESYILEYVKKMVPLVRNALSTLLSSRDESDSVHVAGLV 129
Query: 114 VDAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDIL--EPIKIPG 171
+D F V ++++G E N+ SYI+ +A+ L YL + + ET E E I +PG
Sbjct: 130 LDFFCVPLIDVGNEFNLPSYIFLTCSASFLGMMKYLLERNRETKPELNRSSDEETISVPG 189
Query: 172 CVPLHGRDFLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDN 231
V L ++++Y+ ++ + A G+LVNSF +E +
Sbjct: 190 FVNSVPVKVLPPGLF-TTESYEAWVEMAERFPEAKGILVNSFESLERNAFDYFDRRPDNY 248
Query: 232 PPVYPVGPIIETETKSGDDANGLE-CLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALG 290
PPVYP+GPI+ + + D + + L WLD Q SV+++ FGS +L+ QI E+A
Sbjct: 249 PPVYPIGPILCSNDRPNLDLSERDRILKWLDDQPESSVVFLCFGSLKSLAASQIKEIAQA 308
Query: 291 LELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQ 350
LEL +FLW +R +S + LP GF+ R G V WAPQ++
Sbjct: 309 LELVGIRFLWSIRTDPKEYASPN-----------EILPDGFMNRVMGLGLVC-GWAPQVE 356
Query: 351 ILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRAS---V 407
IL+ ++GGF++HCGWNS LES+ GVP+ TWP++AEQ++NA + + L + L V
Sbjct: 357 ILAHKAIGGFVSHCGWNSILESLRFGVPIATWPMYAEQQLNAFTIVKELGLALEMRLDYV 416
Query: 408 NENG-IVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSS 455
+E G IV+ E+A ++ LM+G++ R +KE+ EA AV + GSS
Sbjct: 417 SEYGEIVKADEIAGAVRSLMDGEDVP--RRKLKEIAEAGKEAVMDGGSS 463
>At1g07240 unknown protein
Length = 480
Score = 202 bits (515), Expect = 2e-52
Identities = 155/483 (32%), Positives = 243/483 (50%), Gaps = 48/483 (9%)
Query: 3 KTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSP----SNATKSILQTL 58
KT + VP HL+ ++F KRL+ L + + ++ P ++A+ + L
Sbjct: 2 KTAELIFVPLPETGHLLSTIEFGKRLLNLDRRISMITIL-SMNLPYAPHADASLASLTAS 60
Query: 59 PSNINHTFLPPVN--PNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIP-------- 108
I LP ++ P T+ E+ IL + ++P L + ++ L
Sbjct: 61 EPGIRIISLPEIHDPPPIKLLDTSSETYILDFIHKNIPCLRKTIQDLVSSSSSSGGGSSH 120
Query: 109 LVALVVDAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDIL--EP 166
+ L++D F V +++IG+E+N+ SYI+ S L YLP+ T E+ + E
Sbjct: 121 VAGLILDFFCVGLIDIGREVNLPSYIFMTSNFGFLGVLQYLPERQRLTPSEFDESSGEEE 180
Query: 167 IKIPGCVPLHGRDFLSIAQ-DRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMK 225
+ IP V L D+ S Y + + L A G+LVNSF ++E P +A
Sbjct: 181 LHIPAFVNRVPAKVLPPGVFDKLS--YGSLVKIGERLHEAKGILVNSFTQVE--PYAAEH 236
Query: 226 -EEGGDNPPVYPVGPIIETETKSGD---DANGLECLAWLDKQQPCSVLYVSFGSGGTLSQ 281
+G D P VYPVGP++ ++ A E + WLD+Q SVL++ FGS G
Sbjct: 237 FSQGRDYPHVYPVGPVLNLTGRTNPGLASAQYKEMMKWLDEQPDSSVLFLCFGSMGVFPA 296
Query: 282 EQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFV 341
QI E+A LEL +F+W +R ++ AG D D + LP GF++RT +G +
Sbjct: 297 PQITEIAHALELIGCRFIWAIR-----TNMAG------DGDPQEPLPEGFVDRTMGRG-I 344
Query: 342 ITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNA--VLLSEGL 399
+ SWAPQ+ IL+ + GGF++HCGWNS ES+ +GVP+ TWP++AEQ++NA ++ GL
Sbjct: 345 VCSWAPQVDILAHKATGGFVSHCGWNSVQESLWYGVPIATWPMYAEQQLNAFEMVKELGL 404
Query: 400 KVGLRASVNENG------IVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDG 453
V +R +G IV E+A ++ LM+ D +R + E A AV + G
Sbjct: 405 AVEIRLDYVADGDRVTLEIVSADEIATAVRSLMDSD--NPVRKKVIEKSSVARKAVGDGG 462
Query: 454 SST 456
SST
Sbjct: 463 SST 465
>At3g21750 putative UDP-glucose glucosyltransferase
Length = 473
Score = 200 bits (508), Expect = 2e-51
Identities = 147/473 (31%), Positives = 234/473 (49%), Gaps = 40/473 (8%)
Query: 5 VHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINH 64
V + +P G H+ +K LV VT + +A+ S+ + +
Sbjct: 3 VELVFIPSPGVGHIRATTALAKLLVASDNRLSVTLIVIPSRVSDDASSSVYTNSEDRLRY 62
Query: 65 TFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIP------LVALVVDAFS 118
LP + Q T + S I + P + + +A ++ L +VVD F
Sbjct: 63 ILLPARD-----QTTDLVSYI----DSQKPQVRAVVSKVAGDVSTRSDSRLAGIVVDMFC 113
Query: 119 VEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSC---EYRDILEPIKIPGCV-P 174
+++I E N+ +YI++ S A+ L F++ L +E E++D +P P
Sbjct: 114 TSMIDIADEFNLSAYIFYTSNASYLGLQFHVQSLYDEKELDVSEFKDTEMKFDVPTLTQP 173
Query: 175 LHGRDFLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDN--P 232
+ S+ ++ + + + L + + G+LVNS ++E LS G+ P
Sbjct: 174 FPAKCLPSVMLNK--KWFPYVLGRARSFRATKGILVNSVADMEPQALSFFSGGNGNTNIP 231
Query: 233 PVYPVGPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLE 292
PVY VGPI++ E+ SGD+ E L WL +Q SV+++ FGS G S+EQ E+A+ LE
Sbjct: 232 PVYAVGPIMDLES-SGDEEKRKEILHWLKEQPTKSVVFLCFGSMGGFSEEQAREIAVALE 290
Query: 293 LSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQIL 352
S +FLW LR S + + E + + LP GFL+RT E G +I SWAPQ+ +L
Sbjct: 291 RSGHRFLWSLRRASPVGNKSNPPPGEFT-NLEEILPKGFLDRTVEIGKII-SWAPQVDVL 348
Query: 353 SPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRAS------ 406
+ ++G F+THCGWNS LES+ GVP+ WP++AEQ+ NA + + ++GL A
Sbjct: 349 NSPAIGAFVTHCGWNSILESLWFGVPMAAWPIYAEQQFNAFHMVD--ELGLAAEVKKEYR 406
Query: 407 ----VNENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSS 455
V E IV E+ + IK ME D K+R + E+K+ A+ + GSS
Sbjct: 407 RDFLVEEPEIVTADEIERGIKCAMEQD--SKMRKRVMEMKDKLHVALVDGGSS 457
>At2g16890 glucosyltransferase like protein
Length = 478
Score = 200 bits (508), Expect = 2e-51
Identities = 146/479 (30%), Positives = 229/479 (47%), Gaps = 46/479 (9%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLH---PNFHVTCFIPTLGSP------SNATKSILQ 56
H+ + P + H++P+LQF + L++ H P VT F P S+ + +
Sbjct: 9 HVVLFPFMSKGHIIPLLQFGRLLLRHHRKEPTITVTVFTTPKNQPFISDFLSDTPEIKVI 68
Query: 57 TLPSNINHTFLPP--VNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVV 114
+LP N T +PP N LP + + T + L + K +P V+ +V
Sbjct: 69 SLPFPENITGIPPGVENTEKLPS-----MSLFVPFTRATKLLQPFFEETLKTLPKVSFMV 123
Query: 115 -DAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCV 173
D F + N+ ++ + + + A + K + T E + EP+ +P
Sbjct: 124 SDGFLWWTSESAAKFNIPRFVSYGMNSYSAAVSISVFKHELFTEPESKSDTEPVTVPDFP 183
Query: 174 PLHGR----DFLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGG 229
+ + D + + S A + + +K +++ G LVNSF E+E + G
Sbjct: 184 WIKVKKCDFDHGTTEPEESGAALELSMDQIKSTTTSHGFLVNSFYELESAFVD-YNNNSG 242
Query: 230 DNPPVYPVGPIIETETKSGDDANGLECLAWLD--KQQPCSVLYVSFGSGGTLSQEQIVEL 287
D P + VGP+ T+ A + WLD +++ VLYV+FG+ +S +Q++EL
Sbjct: 243 DKPKSWCVGPLCLTDPPKQGSAKPA-WIHWLDQKREEGRPVLYVAFGTQAEISNKQLMEL 301
Query: 288 ALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAP 347
A GLE S FLWV R D + + GF +R +E G ++ W
Sbjct: 302 AFGLEDSKVNFLWVTRK-----------------DVEEIIGEGFNDRIRESGMIVRDWVD 344
Query: 348 QIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASV 407
Q +ILS SV GFL+HCGWNS ES+ GVPL+ WP+ AEQ +NA ++ E +KVG+R
Sbjct: 345 QWEILSHESVKGFLSHCGWNSAQESICVGVPLLAWPMMAEQPLNAKMVVEEIKVGVRVET 404
Query: 408 NE---NGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKE-DGSSTKTISQI 462
+ G V R E++ IK LMEG+ G+ R N+KE + A A+ E GSS K + I
Sbjct: 405 EDGSVKGFVTREELSGKIKELMEGETGKTARKNVKEYSKMAKAALVEGTGSSWKNLDMI 463
>At2g29730 putative flavonol 3-O-glucosyltransferase
Length = 467
Score = 199 bits (505), Expect = 4e-51
Identities = 141/476 (29%), Positives = 240/476 (49%), Gaps = 37/476 (7%)
Query: 3 KTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNA---TKSILQTLP 59
+ V + +P HLVP L+F++RL++ +T + L S+ KSI + P
Sbjct: 2 RNVELIFIPTPTVGHLVPFLEFARRLIEQDDRIRITILLMKLQGQSHLDTYVKSIASSQP 61
Query: 60 SNINHTFLPPVNPNDLPQGT-TMESQILLTLTNSLPYLHQG----LKSLAKE-IPLVALV 113
+ +P + T ++E+ + + ++P + L SLA + + + LV
Sbjct: 62 F-VRFIDVPELEEKPTLGSTQSVEAYVYDVIERNIPLVRNIVMDILTSLALDGVKVKGLV 120
Query: 114 VDAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLP-KLDEETSCEYRDILEPIKIPGC 172
VD F + ++++ K++++ Y++ + + LA YL + +TS R+ E + IPG
Sbjct: 121 VDFFCLPMIDVAKDISLPFYVFLTTNSGFLAMMQYLADRHSRDTSVFVRNSEEMLSIPGF 180
Query: 173 VPLHGRDFLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNP 232
V + L A Y ++ L + A+G+LVNS +IE ++ +E + P
Sbjct: 181 VNPVPANVLPSALF-VEDGYDAYVKLAILFTKANGILVNSSFDIEPYSVNHFLQEQ-NYP 238
Query: 233 PVYPVGPIIETETK---SGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELAL 289
VY VGPI + + + D E + WLD Q SV+++ FGS L + E+A
Sbjct: 239 SVYAVGPIFDLKAQPHPEQDLTRRDELMKWLDDQPEASVVFLCFGSMARLRGSLVKEIAH 298
Query: 290 GLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQI 349
GLEL +FLW LR + LP GFL+R +G +I W+PQ+
Sbjct: 299 GLELCQYRFLWSLRKEEVTKDD---------------LPEGFLDRVDGRG-MICGWSPQV 342
Query: 350 QILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRAS--- 406
+IL+ +VGGF++HCGWNS +ES+ GVP++TWP++AEQ++NA L+ + LK+ +
Sbjct: 343 EILAHKAVGGFVSHCGWNSIVESLWFGVPIVTWPMYAEQQLNAFLMVKELKLAVELKLDY 402
Query: 407 -VNENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQ 461
V+ + IV E+ I+Y+M+ D +R + ++ + A K GSS I +
Sbjct: 403 RVHSDEIVNANEIETAIRYVMDTD-NNVVRKRVMDISQMIQRATKNGGSSFAAIEK 457
>At3g21800 putative UDP-glucose glucosyltransferase
Length = 480
Score = 198 bits (504), Expect = 5e-51
Identities = 144/483 (29%), Positives = 230/483 (46%), Gaps = 50/483 (10%)
Query: 7 IAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFI-PTLGSPSNATKSILQTLPSNINHT 65
+ VP HL + +K LV+ ++ I P L + + + L + N
Sbjct: 6 LVFVPFPILGHLKSTAEMAKLLVEQETRLSISIIILPLLSGDDVSASAYISALSAASNDR 65
Query: 66 F-LPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIP-------LVALVVDAF 117
++ D P + L + N +P + + + L + L LVVD F
Sbjct: 66 LHYEVISDGDQPT-------VGLHVDNHIPMVKRTVAKLVDDYSRRPDSPRLAGLVVDMF 118
Query: 118 SVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEE-----TSCEYRDILEPIKIPGC 172
+ V+++ E+++ Y+++ S LA ++ L ++ + ++ D + +P
Sbjct: 119 CISVIDVANEVSVPCYLFYTSNVGILALGLHIQMLFDKKEYSVSETDFEDSEVVLDVPSL 178
Query: 173 VPLHGRDFLSIAQDRSSQAYKHFLPFV----KLLSSADGVLVNSFLEIEMGPLSAMKEEG 228
+ L A K +LP + G+LVN+F E+E L ++
Sbjct: 179 TCPYPVKCLPYGL-----ATKEWLPMYLNQGRRFREMKGILVNTFAELEPYALESL-HSS 232
Query: 229 GDNPPVYPVGPII--ETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVE 286
GD P YPVGP++ E D G + L WLD+Q P SV+++ FGS G ++EQ E
Sbjct: 233 GDTPRAYPVGPLLHLENHVDGSKDEKGSDILRWLDEQPPKSVVFLCFGSIGGFNEEQARE 292
Query: 287 LALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQ-FLPSGFLERTKEKGFVITSW 345
+A+ LE S +FLW LR S + L+ LP GF +RTK+KG VI W
Sbjct: 293 MAIALERSGHRFLWSLRRASRDIDK----ELPGEFKNLEEILPEGFFDRTKDKGKVI-GW 347
Query: 346 APQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRA 405
APQ+ +L+ ++GGF+THCGWNS LES+ GVP+ WPL+AEQK NA ++ E L + ++
Sbjct: 348 APQVAVLAKPAIGGFVTHCGWNSILESLWFGVPIAPWPLYAEQKFNAFVMVEELGLAVKI 407
Query: 406 SVNENG---------IVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSST 456
G IV E+ + I+ LME D +RN +KE+ + A+K+ GSS
Sbjct: 408 RKYWRGDQLVGTATVIVTAEEIERGIRCLMEQD--SDVRNRVKEMSKKCHMALKDGGSSQ 465
Query: 457 KTI 459
+
Sbjct: 466 SAL 468
>At2g29750 putative flavonol 3-O-glucosyltransferase
Length = 481
Score = 198 bits (503), Expect = 6e-51
Identities = 147/480 (30%), Positives = 242/480 (49%), Gaps = 46/480 (9%)
Query: 2 EKTVHIAVVPGVGYSHLVPILQFSKRLV-QLHPNFHVTCFI----PTLGSPSNATKSILQ 56
++ + ++P H++ ++ +KRL+ Q +P H + P + P T + L+
Sbjct: 4 QEDAELVIIPFPFSGHILATIELAKRLISQDNPRIHTITILYWGLPFI--PQADTIAFLR 61
Query: 57 TLPSN---INHTFLPPVNPNDLPQGTTM----ESQILLTLTNSLPYLHQGLKSLAKE--- 106
+L N I LP V D P ES IL + +P + + L +L
Sbjct: 62 SLVKNEPRIRLVTLPEVQ--DPPPMELFVEFAESYILEYVKKMVPIIREALSTLLSSRDE 119
Query: 107 ---IPLVALVVDAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEY-RD 162
+ + LV+D F V ++++G E N+ SYI+ +A L YLP+ E E+ R
Sbjct: 120 SGSVRVAGLVLDFFCVPMIDVGNEFNLPSYIFLTCSAGFLGMMKYLPERHREIKSEFNRS 179
Query: 163 ILEPIK-IPGCVPLHGRDFLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPL 221
E + IPG V L + Y+ ++ + A G+LVNS+ +E
Sbjct: 180 FNEELNLIPGYVNSVPTKVLPSGLFMK-ETYEPWVELAERFPEAKGILVNSYTALEPNGF 238
Query: 222 SAMKEEGGDNPPVYPVGPIIETETKSGDDANGLE-CLAWLDKQQPCSVLYVSFGSGGTLS 280
+ P +YP+GPI+ + + D++ + + WLD Q SV+++ FGS LS
Sbjct: 239 KYFDRCPDNYPTIYPIGPILCSNDRPNLDSSERDRIITWLDDQPESSVVFLCFGSLKNLS 298
Query: 281 QEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGF 340
QI E+A LE+ + KF+W R +S + LP GF++R ++G
Sbjct: 299 ATQINEIAQALEIVDCKFIWSFRTNPKEYASP-----------YEALPHGFMDRVMDQG- 346
Query: 341 VITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLK 400
++ WAPQ++IL+ +VGGF++HCGWNS LES+ GVP+ TWP++AEQ++NA + + L
Sbjct: 347 IVCGWAPQVEILAHKAVGGFVSHCGWNSILESLGFGVPIATWPMYAEQQLNAFTMVKELG 406
Query: 401 VGLRAS---VNENG-IVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSST 456
+ L V+E+G IV+ E+A ++ LM+G + K + +KE+ EA AV DG S+
Sbjct: 407 LALEMRLDYVSEDGDIVKADEIAGTVRSLMDGVDVPK--SKVKEIAEAGKEAV--DGGSS 462
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.135 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,680,938
Number of Sequences: 26719
Number of extensions: 465614
Number of successful extensions: 1853
Number of sequences better than 10.0: 127
Number of HSP's better than 10.0 without gapping: 120
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1367
Number of HSP's gapped (non-prelim): 156
length of query: 470
length of database: 11,318,596
effective HSP length: 103
effective length of query: 367
effective length of database: 8,566,539
effective search space: 3143919813
effective search space used: 3143919813
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC124966.3


