

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124963.1 + phase: 1 /pseudo
(826 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
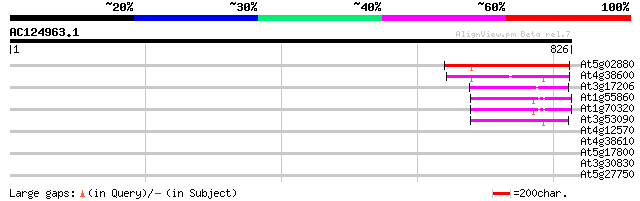
Score E
Sequences producing significant alignments: (bits) Value
At5g02880 putative protein 236 5e-62
At4g38600 putative protein 178 1e-44
At3g17206 putative ubiquitin ligase, 5'partial 94 3e-19
At1g55860 ubiquitin-protein ligase 1, putative 88 2e-17
At1g70320 hypothetical protein 85 1e-16
At3g53090 putative protein 73 7e-13
At4g12570 polyubiquitin-like protein 40 0.005
At4g38610 putative protein 35 0.17
At5g17800 MYB56 R2R3-MYB factor family member 32 1.1
At3g30830 hypothetical protein 31 3.2
At5g27750 unknown protein 30 7.2
>At5g02880 putative protein
Length = 1502
Score = 236 bits (601), Expect = 5e-62
Identities = 119/206 (57%), Positives = 149/206 (71%), Gaps = 22/206 (10%)
Query: 641 AFRNSKIEDLCLDFSLPGYDQTFLLPFSIY*ITNLYSF---------------------- 678
+F +KIEDLCL+F+LPGY L P+S + NL +
Sbjct: 1294 SFHGTKIEDLCLEFALPGYTDYDLAPYSANDMVNLDNLEEYIKGIVNATVCNGIQKQVEA 1353
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F GFNQVF IEHLR+F+EEELE +LCGE + ++ N++L H KFDHGYT+ SPP+ LL+
Sbjct: 1354 FRSGFNQVFSIEHLRIFNEEELETMLCGECDLFSMNEVLDHIKFDHGYTSSSPPVEYLLQ 1413
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCANYL 798
IL EF+ E++RAF+QFVT SPRLP GGLASL PKLT+V+K + +DTDLPSVMTCANYL
Sbjct: 1414 ILHEFDREQQRAFLQFVTGSPRLPHGGLASLSPKLTIVRKHGSDSSDTDLPSVMTCANYL 1473
Query: 799 KLPPYSSKERMKQKLLYAITEGRGCF 824
KLPPYSSKE+MK+KL+YAITEG+G F
Sbjct: 1474 KLPPYSSKEKMKEKLIYAITEGQGSF 1499
Score = 38.9 bits (89), Expect = 0.012
Identities = 17/34 (50%), Positives = 28/34 (82%), Gaps = 1/34 (2%)
Query: 1 VLILALQIVELILQKFFSDKFIKLFIEEGVYFAI 34
V+++ALQ+ E++L+K+ D F+ FI+EGV+FAI
Sbjct: 503 VIVVALQVAEVLLEKY-RDTFLNSFIKEGVFFAI 535
>At4g38600 putative protein
Length = 757
Score = 178 bits (451), Expect = 1e-44
Identities = 102/214 (47%), Positives = 129/214 (59%), Gaps = 35/214 (16%)
Query: 643 RNSKIEDLCLDFSLPGYDQTFLLPFS-IY*ITNLYSF-------------------FLVG 682
R +IEDL L+F+LPGY + L I ITNL + F G
Sbjct: 544 RGCRIEDLSLEFTLPGYPEYILRSGDEIVDITNLEEYISLVVDATVKRGVTRQIEAFRSG 603
Query: 683 FNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQE 742
FNQVF I L++F+ EL+ +LCG W L +H KFDHGY A+SP I+N +
Sbjct: 604 FNQVFDITSLQIFTPSELDYLLCGRRELWEVETLAEHIKFDHGYNAKSPAIIN---VCYP 660
Query: 743 FNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHT------------DTDLPS 790
+++++RAF QFVT +PRLPPGGLA L+PKLT+V+K S + D DLPS
Sbjct: 661 SSSDQQRAFCQFVTGAPRLPPGGLAVLNPKLTIVRKHSSTSSAAANGAGASETADDDLPS 720
Query: 791 VMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
VMTCANYLKLPPYS+KE M +KLLYAI EG+G F
Sbjct: 721 VMTCANYLKLPPYSTKEIMYKKLLYAINEGQGSF 754
>At3g17206 putative ubiquitin ligase, 5'partial
Length = 276
Score = 94.0 bits (232), Expect = 3e-19
Identities = 53/146 (36%), Positives = 83/146 (56%), Gaps = 2/146 (1%)
Query: 677 SFFLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNL 736
S FL GF Q+ P E + +F+E EL++++ G +S +DL + + GY A I
Sbjct: 129 SHFLRGFQQLIPKEWIDMFNEHELQVLISGSVDSLDIDDLRNNTNYAGGYHAGHYVIDMF 188
Query: 737 LEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCAN 796
E+++ F+ E ++ F++FVT R P G L+P + + + N + LP+ TC N
Sbjct: 189 WEVMKSFSTENQKKFLKFVTGCSRGPLLGFKYLEPAFCI--QSASNESVDRLPTSATCMN 246
Query: 797 YLKLPPYSSKERMKQKLLYAITEGRG 822
LKLPPY SKE ++ KL+YAI+ G
Sbjct: 247 LLKLPPYQSKELLETKLMYAISAEAG 272
>At1g55860 ubiquitin-protein ligase 1, putative
Length = 3891
Score = 88.2 bits (217), Expect = 2e-17
Identities = 57/152 (37%), Positives = 89/152 (58%), Gaps = 8/152 (5%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL GFN++ P E + +F+++ELEL++ G P F+DL + ++ YTA SP I E
Sbjct: 3743 FLEGFNELIPRELVSIFNDKELELLISGLPEI-DFDDLKANTEYT-SYTAGSPVIHWFWE 3800
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLD----PKLTVVQKISYNHTDTDLPSVMTC 794
+++ F+ E+ F+QFVT + ++P G +L P+ + K +Y + LPS TC
Sbjct: 3801 VVKAFSKEDMARFLQFVTGTSKVPLEGFKALQGISGPQRLQIHK-AYGAPER-LPSAHTC 3858
Query: 795 ANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
N L LP Y SKE+++++LL AI E F F
Sbjct: 3859 FNQLDLPEYQSKEQLQERLLLAIHEASEGFGF 3890
>At1g70320 hypothetical protein
Length = 3658
Score = 85.1 bits (209), Expect = 1e-16
Identities = 55/152 (36%), Positives = 87/152 (57%), Gaps = 8/152 (5%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL G N++ P E + +F+++ELEL++ G P F+DL + ++ YT SP I E
Sbjct: 3510 FLEGLNELIPRELVSIFNDKELELLISGLPEI-DFDDLKANTEYT-SYTVGSPVIRWFWE 3567
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLD----PKLTVVQKISYNHTDTDLPSVMTC 794
+++ F+ E+ F+QFVT + ++P G +L P+ + K +Y + LPS TC
Sbjct: 3568 VVKAFSKEDMARFLQFVTGTSKVPLEGFKALQGISGPQRLQIHK-AYGSPER-LPSAHTC 3625
Query: 795 ANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
N L LP Y SKE+++++LL AI E F F
Sbjct: 3626 FNQLDLPEYQSKEQVQERLLLAIHEANEGFGF 3657
>At3g53090 putative protein
Length = 1142
Score = 72.8 bits (177), Expect = 7e-13
Identities = 49/153 (32%), Positives = 73/153 (47%), Gaps = 10/153 (6%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F G + L++F+ E +L G + +DL ++ K+ GY+ S I E
Sbjct: 987 FYRGLTDLISPAWLKLFNAHEFNQLLSGGNHDIDVDDLRRNTKYTGGYSDSSRTIKIFWE 1046
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHT--------DTD-LP 789
+++ F ER ++FVT R P G L P ++ K+S + + D + LP
Sbjct: 1047 VMKGFEPSERCLLLKFVTSCSRAPLLGFKYLQPTF-IIHKVSCDTSLWAAIGGQDVERLP 1105
Query: 790 SVMTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
S TC N LKLP Y M++KLLYAIT G
Sbjct: 1106 SASTCYNTLKLPTYKRASTMREKLLYAITSNAG 1138
>At4g12570 polyubiquitin-like protein
Length = 873
Score = 40.0 bits (92), Expect = 0.005
Identities = 32/116 (27%), Positives = 55/116 (46%), Gaps = 4/116 (3%)
Query: 698 EELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEFNNEERRAFVQFVTR 757
E+L+ +L G N + +D H +++ G+ I +IL++ EE+R+ + F T
Sbjct: 747 EDLDGMLRGGENPISIDDWKAHTEYN-GFKETDRQIDWFWKILKKMTEEEQRSILFFWTS 805
Query: 758 SPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCANYLKLPPYSSKERMKQKL 813
+ +P G L KL + + N LP TC L +P Y + M+Q+L
Sbjct: 806 NKFVPVEGFRGLSSKLYIYRLYEANDR---LPLSHTCFYRLCIPRYPTITLMEQRL 858
>At4g38610 putative protein
Length = 1083
Score = 35.0 bits (79), Expect = 0.17
Identities = 17/45 (37%), Positives = 28/45 (61%), Gaps = 1/45 (2%)
Query: 1 VLILALQIVELILQKFFSDKFIKLFIEEGVYFAISHFHHLGKHLH 45
VL+ ALQ+ E++++K + F K+F+ EGV A+ +GK H
Sbjct: 646 VLVPALQVAEILMEKL-PETFSKVFVREGVVHAVDQLVLVGKPSH 689
>At5g17800 MYB56 R2R3-MYB factor family member
Length = 323
Score = 32.3 bits (72), Expect = 1.1
Identities = 45/188 (23%), Positives = 68/188 (35%), Gaps = 50/188 (26%)
Query: 639 WKAFRNSKIEDLCLDFSLPGYDQTFLLPFSIY*ITNLYSFFLVG----------FNQVFP 688
W+ ++K+++L F + NL S L+G FNQ+ P
Sbjct: 96 WRPTEDAKLKELVAQFGPQNW--------------NLISNHLLGRSGKSCRLRWFNQLDP 141
Query: 689 IEHLRVFSEEELELILCGEP---NSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEFNN 745
+ R F+EEE +L N W L + D+ + N ++
Sbjct: 142 RINKRAFTEEEEFRLLAAHRAYGNKWALISRLFPGRTDNA-------VKNHWHVIMARRT 194
Query: 746 EERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNH--------TDTDLPSVMTCANY 797
E + R + PP L S D ++TV YN D D+ +V TC
Sbjct: 195 RESQ-------RQRQQPPPTL-SRDAEMTVSSSCRYNQGKFINEEDDDDDVSAVSTCTTE 246
Query: 798 LKLPPYSS 805
L L P SS
Sbjct: 247 LSLTPPSS 254
>At3g30830 hypothetical protein
Length = 327
Score = 30.8 bits (68), Expect = 3.2
Identities = 16/51 (31%), Positives = 31/51 (60%), Gaps = 2/51 (3%)
Query: 644 NSKIEDLCLDF--SLPGYDQTFLLPFSIY*ITNLYSFFLVGFNQVFPIEHL 692
NS ++++ ++F +LPG ++ L +S++ N+Y F L NQ+F +L
Sbjct: 139 NSYVQEVVMEFYANLPGGEEGDQLAYSVFVRGNMYEFSLAITNQMFHFPNL 189
>At5g27750 unknown protein
Length = 459
Score = 29.6 bits (65), Expect = 7.2
Identities = 17/58 (29%), Positives = 28/58 (47%), Gaps = 2/58 (3%)
Query: 645 SKIEDLCLDFSLPGYDQTFLLP--FSIY*ITNLYSFFLVGFNQVFPIEHLRVFSEEEL 700
S++ D L+ PG + L F + NL S F++ N F +++ R F E+L
Sbjct: 233 SRVPDKVLEIDAPGLENMTLKEDHFDGIVVKNLTSLFMIELNIKFAVDYRRSFDPEDL 290
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.358 0.160 0.582
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,068,099
Number of Sequences: 26719
Number of extensions: 611026
Number of successful extensions: 2602
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 2580
Number of HSP's gapped (non-prelim): 14
length of query: 826
length of database: 11,318,596
effective HSP length: 108
effective length of query: 718
effective length of database: 8,432,944
effective search space: 6054853792
effective search space used: 6054853792
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC124963.1


