

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124958.16 + phase: 0
(248 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
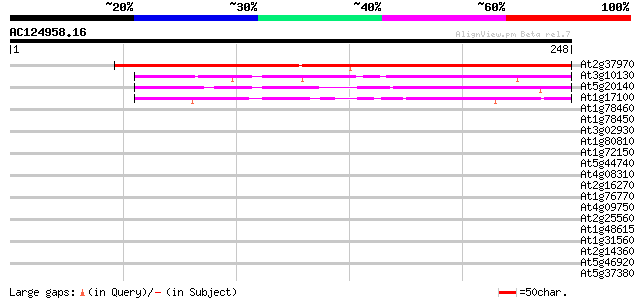
Score E
Sequences producing significant alignments: (bits) Value
At2g37970 unknown protein (At2g37970) 299 1e-81
At3g10130 unknown protein 99 3e-21
At5g20140 unknown protein 87 8e-18
At1g17100 SOUL-like protein 50 1e-06
At1g78460 unknown protein 39 0.003
At1g78450 unknown protein 33 0.19
At3g02930 unknown protein 32 0.32
At1g80810 unknown protein 30 0.92
At1g72150 putative cytosolic factor protein 30 1.2
At5g44740 translesion synthesis polymerase RAD30 like protein 30 1.6
At4g08310 unknown protein (At4g08310) 29 2.7
At2g16270 unknown protein 29 2.7
At1g76770 putative heat shock protein 29 2.7
At4g09750 putative protein 28 3.5
At2g25560 putative DnaJ protein 28 3.5
At1g48615 unknown protein 28 3.5
At1g31560 putative protein 28 3.5
At2g14360 pseudogene 28 4.6
At5g46920 unknown protein 28 6.0
At5g37380 unknown protein 28 6.0
>At2g37970 unknown protein (At2g37970)
Length = 215
Score = 299 bits (765), Expect = 1e-81
Identities = 151/216 (69%), Positives = 172/216 (78%), Gaps = 15/216 (6%)
Query: 47 MGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIG 106
MGMVFGKI VETPKY VTK+ YEIR Y P+VAAEVTYD S+FKG+KDGGF +LA YIG
Sbjct: 1 MGMVFGKIAVETPKYTVTKSGDGYEIREYPPAVAAEVTYDASEFKGDKDGGFQLLAKYIG 60
Query: 107 ALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGE--------------RNK 152
G P+N KPEKIAMTAPVITK EKIAMTAPV+TK SE+ E R K
Sbjct: 61 VFGKPENEKPEKIAMTAPVITK-EGEKIAMTAPVITKESEKIEMTSPVVTKEGGGEGRKK 119
Query: 153 MVTMQFILPSSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLS 212
+VTMQF+LPS Y+KAEEAP+PTDERVVI+EEG RKYGV+KFSG+AS+ VV EKV+KL
Sbjct: 120 LVTMQFLLPSMYKKAEEAPRPTDERVVIKEEGGRKYGVIKFSGIASESVVSEKVKKLSSH 179
Query: 213 LERDGFKVIGDFLLGRYNPPWTLPMFRTNEVMIPIE 248
LE+DGFK+ GDF+L RYNPPWTLP FRTNEVMIP+E
Sbjct: 180 LEKDGFKITGDFVLARYNPPWTLPPFRTNEVMIPVE 215
>At3g10130 unknown protein
Length = 309
Score = 98.6 bits (244), Expect = 3e-21
Identities = 70/202 (34%), Positives = 104/202 (50%), Gaps = 19/202 (9%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGG---FMVLANYIGALGNPQ 112
+ET + V T YEIR P AE T P + + G F VLA Y+ +
Sbjct: 114 LETMNFRVLFRTDKYEIRQVEPYFVAE-TIMPGETGFDSYGASKSFNVLAEYLFG----K 168
Query: 113 NTKPEKIAMTAPVITK---GSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
NT EK+ MT PV+T+ EK+ MT PV+T +++ + +M F++PS Y
Sbjct: 169 NTIKEKMEMTTPVVTRKVQSVGEKMEMTTPVITSKAKDQNQWRM---SFVMPSKY--GSN 223
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGD---FLL 226
P P D V I++ + VV FSG +DE ++ + +LR +L+ D + D F +
Sbjct: 224 LPLPKDPSVKIQQVPRKIVAVVAFSGYVTDEEIERRERELRRALQNDKKFRVRDGVSFEV 283
Query: 227 GRYNPPWTLPMFRTNEVMIPIE 248
+YNPP+TLP R NEV + +E
Sbjct: 284 AQYNPPFTLPFMRRNEVSLEVE 305
>At5g20140 unknown protein
Length = 378
Score = 87.0 bits (214), Expect = 8e-18
Identities = 62/194 (31%), Positives = 98/194 (49%), Gaps = 26/194 (13%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
+ETPKY++ K T +YE+R Y P + E D K + GF +A YI +N+
Sbjct: 205 LETPKYQILKRTANYEVRNYEPFIVVETIGD----KLSGSSGFNNVAGYIFG----KNST 256
Query: 116 PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
EKI MT PV T+ + +++ V++Q ++PS + + P P +
Sbjct: 257 MEKIPMTTPVFTQTTDTQLSSD----------------VSVQIVIPSGKDLSS-LPMPNE 299
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPW-T 234
E+V +++ VKFSG +++VV+ K +LR SL +DG + +L RYN P T
Sbjct: 300 EKVNLKKLEGGFAAAVKFSGKPTEDVVQAKENELRSSLSKDGLRAKKGCMLARYNDPGRT 359
Query: 235 LPMFRTNEVMIPIE 248
NEV+I +E
Sbjct: 360 WNFIMRNEVIIWLE 373
>At1g17100 SOUL-like protein
Length = 232
Score = 50.1 bits (118), Expect = 1e-06
Identities = 57/208 (27%), Positives = 86/208 (40%), Gaps = 38/208 (18%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSV--AAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQN 113
+E P YE+ + YEIR Y +V + E D S + F + A G +N
Sbjct: 45 IECPSYELVHSGNGYEIRRYNNTVWVSTEPIPDISLVDATRTAFFQLFAYIQG-----KN 99
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
+KI MTAPVI++ S P S T+ F +P +K + P P
Sbjct: 100 EYHQKIEMTAPVISQVSPSD----GPFCESS---------FTVSFYVP---KKNQPDPAP 143
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSL-------------ERDGFKV 220
+ E + I++ R V +FSG SD+ + E+ L SL E G
Sbjct: 144 S-ENLHIQKWNSRYVAVRQFSGFVSDDSIGEQAAALDSSLKGTAWANAIAKSKEDGGVGS 202
Query: 221 IGDFLLGRYNPPWTLPMFRTNEVMIPIE 248
+ + +YN P+ R NE+ +P E
Sbjct: 203 DSAYTVAQYNSPFEF-SGRVNEIWLPFE 229
>At1g78460 unknown protein
Length = 219
Score = 38.9 bits (89), Expect = 0.003
Identities = 46/199 (23%), Positives = 76/199 (38%), Gaps = 35/199 (17%)
Query: 57 ETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPS-QFKGNKDGGFMVLANYIGALGNPQNTK 115
E P Y++ + +EIR+Y ++ + PS GF L YI N
Sbjct: 44 ECPTYKLVEAGYGFEIRMYDAALWISTSPIPSLSMTQATKTGFRRLNRYIEG----DNKS 99
Query: 116 PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
K+ MTAPVI + + + + T+ LP +K ++ P D
Sbjct: 100 NVKMNMTAPVIAQATPGR------------------SVYTVSLYLP---KKNQQNPPQAD 138
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKV--------IGDFLLG 227
+ + +R V + G S+ V K++ L SL + + + + L
Sbjct: 139 D-LHVRSTKPTYVAVRQIGGYVSNNVAKDEAAALMESLRDSNWILPIEKSKGKLPAYFLA 197
Query: 228 RYNPPWTLPMFRTNEVMIP 246
YNPP NE+M+P
Sbjct: 198 VYNPPSHTTARVINEIMVP 216
>At1g78450 unknown protein
Length = 225
Score = 32.7 bits (73), Expect = 0.19
Identities = 39/162 (24%), Positives = 63/162 (38%), Gaps = 33/162 (20%)
Query: 57 ETPKYEVTKTTQDYEIRIYAPSV--AAEVTYDPS--QFKGNKDGGFMVLANYIGALGNPQ 112
E P YEV YEI Y +V + E D S + GN G+ L++Y+ N
Sbjct: 35 ECPSYEVVHAGNGYEIHRYNTTVWISTEPIQDISLNEASGN---GWNQLSDYM----NGN 87
Query: 113 NTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPK 172
N ++I + P IT+ S FI+ KA +
Sbjct: 88 NDYHQRIEIALPYITQVSQN----------------------LSTFIVSFFVPKAFQPDP 125
Query: 173 PTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLE 214
P + ++ R V + SG +D + ++V +L+ SL+
Sbjct: 126 PPGNNLHVQRWDSRYVAVKQISGYVADHKIGKQVAELKASLQ 167
>At3g02930 unknown protein
Length = 806
Score = 32.0 bits (71), Expect = 0.32
Identities = 27/106 (25%), Positives = 51/106 (47%), Gaps = 8/106 (7%)
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKS----SEEGERNKMVTMQFILPSSYEKAEE 169
T PEK + A +++ + + + + + S E E+ K + L + ++AEE
Sbjct: 70 TPPEKTQIRAVRVSESQPQSVQIKEDLKKANELIASLENEKAKALDQ---LKEARKEAEE 126
Query: 170 APKPTDERVVIREEGERKYGVVKFSGV-ASDEVVKEKVEKLRLSLE 214
A + DE + +++ + + KF V A E V+ K E+L+ LE
Sbjct: 127 ASEKLDEALEAQKKSLENFEIEKFEVVEAGIEAVQRKEEELKKELE 172
>At1g80810 unknown protein
Length = 826
Score = 30.4 bits (67), Expect = 0.92
Identities = 30/115 (26%), Positives = 44/115 (38%), Gaps = 11/115 (9%)
Query: 86 DPSQFKGNKDGGFMVLANYIGALGNPQNTKPEKIAMTAPVITKG----SAEKIAMT---- 137
+P+Q K G L Q T +K + A ++ SA +A
Sbjct: 441 EPNQEDDRKIGNSSKQTRSKNGLEKSQKTAKKKPVVEAKIVNSSGKRLSARSVAKRRNLE 500
Query: 138 -APVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPK--PTDERVVIREEGERKYG 189
AP+ T + +R KMV+ + E EE PK PT R V +E +G
Sbjct: 501 RAPLDTLVPQSSKRKKMVSQVAARQLANESEEETPKSHPTRRRTVRKEVESDGFG 555
>At1g72150 putative cytosolic factor protein
Length = 573
Score = 30.0 bits (66), Expect = 1.2
Identities = 28/102 (27%), Positives = 41/102 (39%), Gaps = 3/102 (2%)
Query: 112 QNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAP 171
+ T K+ P + EK + APV TKS E+ E VT + S+ E +
Sbjct: 146 ETTTEVKVEEEKPAVPAAEEEKSSEAAPVETKSEEKPEEKAEVTTE-KASSAEEDGTKTV 204
Query: 172 KPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSL 213
+ +E +V E V VA E E VE +S+
Sbjct: 205 EAIEESIVSVSPPESAVAPVVVETVAVAEA--EPVEPEEVSI 244
>At5g44740 translesion synthesis polymerase RAD30 like protein
Length = 672
Score = 29.6 bits (65), Expect = 1.6
Identities = 36/169 (21%), Positives = 64/169 (37%), Gaps = 15/169 (8%)
Query: 43 SRSIMGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKG-NKDGGFMVL 101
SRS G V G + + E + E + P V VTY F+ +KD + L
Sbjct: 458 SRSADGCVQGNVAMTASASEGCSEQRSTETQAAMPEVDTGVTYTLPNFENQDKD---IDL 514
Query: 102 ANYIGALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILP 161
+ + P N + V T+ + K T + K + E+N+ + +
Sbjct: 515 VSEKDVVSCPSNEATD-------VSTQSESNKGTQTKKIGRKMNNSKEKNRGMPSIVDIF 567
Query: 162 SSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLR 210
+Y + + T E + +R K S + + V ++VE+ R
Sbjct: 568 KNYNATPPSKQETQEDSTVSSASKR----AKLSSSSHNSQVNQEVEESR 612
>At4g08310 unknown protein (At4g08310)
Length = 504
Score = 28.9 bits (63), Expect = 2.7
Identities = 20/61 (32%), Positives = 34/61 (54%), Gaps = 5/61 (8%)
Query: 159 ILPSSYEKAEEAPKPTDERVVIREEGE--RKYGVVKFSGVASDEVVKEKVEKLRLSLERD 216
I PS Y KA++AP+ E ++I+E E K G+ S S++ +KE ++ + E +
Sbjct: 361 ISPSVYRKAKQAPEEKREEILIKELKELLAKEGL---SANPSEKEIKEVKKRKERTKELE 417
Query: 217 G 217
G
Sbjct: 418 G 418
>At2g16270 unknown protein
Length = 759
Score = 28.9 bits (63), Expect = 2.7
Identities = 22/67 (32%), Positives = 33/67 (48%), Gaps = 7/67 (10%)
Query: 163 SYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDG----F 218
S E++EE K E VV+ EE E V + + +E+V E VE+ + + F
Sbjct: 254 SVEESEEQEKDGAEEVVVEEETE---DVEQSEAESDEEMVCESVEETTSQVPKQSGSRKF 310
Query: 219 KVIGDFL 225
K +G FL
Sbjct: 311 KFLGWFL 317
>At1g76770 putative heat shock protein
Length = 244
Score = 28.9 bits (63), Expect = 2.7
Identities = 27/99 (27%), Positives = 47/99 (47%), Gaps = 4/99 (4%)
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVV-TKSSEEGERNKMVTMQFILPSSYEKAEEAPK 172
T P+K+ + + E+ M P+V K+ E+ E + + + E+AEE +
Sbjct: 124 TMPKKVKGITGLKIEEEDEEEEMKEPIVEEKTEEKTEPEEEIKEETKPEEENEEAEEPQR 183
Query: 173 PTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRL 211
+E VV EEG R + K + D+ K++ +K RL
Sbjct: 184 EEEEEVV--EEGTRDHEGKKEEEI-EDKPRKKRRKKFRL 219
>At4g09750 putative protein
Length = 322
Score = 28.5 bits (62), Expect = 3.5
Identities = 21/90 (23%), Positives = 39/90 (43%), Gaps = 1/90 (1%)
Query: 60 KYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTKPEKI 119
K + + Q+ + + S E+ S F +KD VL N G L N + T PE
Sbjct: 86 KIQTSTGNQNVYLEVCDLSSVNEIKSFASSF-ASKDVPVHVLVNNAGLLENKRTTTPEGF 144
Query: 120 AMTAPVITKGSAEKIAMTAPVVTKSSEEGE 149
++ V G+ + P++ K++ + +
Sbjct: 145 ELSFAVNVLGTYTMTELMLPLLEKATPDAK 174
>At2g25560 putative DnaJ protein
Length = 656
Score = 28.5 bits (62), Expect = 3.5
Identities = 13/35 (37%), Positives = 21/35 (59%)
Query: 188 YGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIG 222
YGV+ + A DE+V+++ KL + L D K +G
Sbjct: 68 YGVLGLNPEADDEIVRKRYRKLAVMLHPDRNKSVG 102
>At1g48615 unknown protein
Length = 212
Score = 28.5 bits (62), Expect = 3.5
Identities = 18/63 (28%), Positives = 31/63 (48%), Gaps = 12/63 (19%)
Query: 102 ANYIGALGNPQNTKPE------------KIAMTAPVITKGSAEKIAMTAPVVTKSSEEGE 149
++ IGA+ +TKP ++ A V +G A++ +TA VVT ++ EG
Sbjct: 58 SSQIGAVSAKASTKPSGRPKRNVAQAVPSTSVAAAVKKRGRAKRSTVTAAVVTTATGEGS 117
Query: 150 RNK 152
R +
Sbjct: 118 RKR 120
>At1g31560 putative protein
Length = 1431
Score = 28.5 bits (62), Expect = 3.5
Identities = 28/105 (26%), Positives = 45/105 (42%), Gaps = 10/105 (9%)
Query: 104 YIGALGNPQNTKPEKIAMTAPVITKGSAEKIA-MTAPVVTKSSEEGERNKMVTMQFILPS 162
YIG + PQ+T E+ T+ SA++ T PVV + +EE K +
Sbjct: 833 YIGTI-EPQDTAAEQ--------TEPSAKQTEPQTEPVVDQQAEEDLLQKENPLVTDKDG 883
Query: 163 SYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVE 207
E+ R + + +R+ G K +A D V+E+VE
Sbjct: 884 LNEEEAVGEIELSGRTEPQAKRKRQRGPTKMKNIAKDPTVRERVE 928
>At2g14360 pseudogene
Length = 221
Score = 28.1 bits (61), Expect = 4.6
Identities = 14/49 (28%), Positives = 26/49 (52%)
Query: 128 KGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTDE 176
K S+ +++ + PV ++ EG+ +V + + EKA E P+ DE
Sbjct: 18 KSSSGEVSTSEPVTSEIENEGDAADLVPTEPAETAEIEKAMEEPRVGDE 66
>At5g46920 unknown protein
Length = 735
Score = 27.7 bits (60), Expect = 6.0
Identities = 15/41 (36%), Positives = 21/41 (50%), Gaps = 2/41 (4%)
Query: 133 KIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
K A+ PVVT E+GE+ K ++ AE+ PKP
Sbjct: 272 KSALVTPVVTSKVEDGEKKKTKKRKY--QKKRVLAEDEPKP 310
>At5g37380 unknown protein
Length = 431
Score = 27.7 bits (60), Expect = 6.0
Identities = 15/35 (42%), Positives = 18/35 (50%)
Query: 188 YGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIG 222
YG++ S DE +K K KL L L D K IG
Sbjct: 68 YGILNASPRDDDETLKRKYRKLALMLHPDKNKSIG 102
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.133 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,588,720
Number of Sequences: 26719
Number of extensions: 239179
Number of successful extensions: 532
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 506
Number of HSP's gapped (non-prelim): 26
length of query: 248
length of database: 11,318,596
effective HSP length: 97
effective length of query: 151
effective length of database: 8,726,853
effective search space: 1317754803
effective search space used: 1317754803
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC124958.16


