

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124953.2 + phase: 0
(53 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
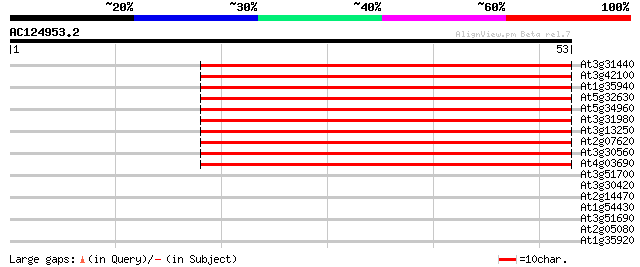
Score E
Sequences producing significant alignments: (bits) Value
At3g31440 hypothetical protein 52 4e-08
At3g42100 putative protein 52 7e-08
At1g35940 hypothetical protein 51 1e-07
At5g32630 putative protein 50 2e-07
At5g34960 putative protein 49 4e-07
At3g31980 hypothetical protein 49 5e-07
At3g13250 hypothetical protein 48 8e-07
At2g07620 putative helicase 48 8e-07
At3g30560 hypothetical protein 44 1e-05
At4g03690 hypothetical protein 42 6e-05
At3g51700 unknown protein 35 0.007
At3g30420 hypothetical protein 30 0.30
At2g14470 pseudogene 30 0.30
At1g54430 hypothetical protein 30 0.30
At3g51690 putative protein 29 0.39
At2g05080 putative helicase 26 3.3
At1g35920 hypothetical protein 26 3.3
>At3g31440 hypothetical protein
Length = 536
Score = 52.4 bits (124), Expect = 4e-08
Identities = 24/35 (68%), Positives = 29/35 (82%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLKILI D+NG TTN+V+KEVFQN+
Sbjct: 501 RVTSKKGLKILILDKNGKLQKQTTNIVFKEVFQNI 535
>At3g42100 putative protein
Length = 1752
Score = 51.6 bits (122), Expect = 7e-08
Identities = 24/35 (68%), Positives = 30/35 (85%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLKILI D++G+ TTNVV+KEVFQN+
Sbjct: 1717 RVTSKKGLKILILDKDGNMQKQTTNVVFKEVFQNI 1751
>At1g35940 hypothetical protein
Length = 1678
Score = 50.8 bits (120), Expect = 1e-07
Identities = 24/35 (68%), Positives = 29/35 (82%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLKILI D++G TTNVV+KEVFQN+
Sbjct: 1643 RVTSKKGLKILILDKDGKLQKQTTNVVFKEVFQNI 1677
>At5g32630 putative protein
Length = 856
Score = 50.4 bits (119), Expect = 2e-07
Identities = 23/35 (65%), Positives = 29/35 (82%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+ GLKILI D++GD TTNVV+KE+FQN+
Sbjct: 821 RVTSKSGLKILILDKDGDIQKQTTNVVFKELFQNI 855
>At5g34960 putative protein
Length = 1033
Score = 49.3 bits (116), Expect = 4e-07
Identities = 23/35 (65%), Positives = 28/35 (79%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLK LI D++G TTNVV+KEVFQN+
Sbjct: 998 RVTSKKGLKFLILDKDGKLQKQTTNVVFKEVFQNI 1032
>At3g31980 hypothetical protein
Length = 1099
Score = 48.9 bits (115), Expect = 5e-07
Identities = 21/35 (60%), Positives = 28/35 (80%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLK+LI D+ G+ T NVV+KE+FQN+
Sbjct: 1063 RVTSKKGLKVLIVDKEGNTQSQTMNVVFKEIFQNI 1097
>At3g13250 hypothetical protein
Length = 1419
Score = 48.1 bits (113), Expect = 8e-07
Identities = 23/35 (65%), Positives = 27/35 (76%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+ GLKILI D+ G TTNVV+KEVFQN+
Sbjct: 1385 RVTSKTGLKILILDKEGKIQKQTTNVVFKEVFQNI 1419
>At2g07620 putative helicase
Length = 1241
Score = 48.1 bits (113), Expect = 8e-07
Identities = 21/35 (60%), Positives = 28/35 (80%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS+KGLK+LI D+ G+ T NVV+KE+FQN+
Sbjct: 1205 RVTSKKGLKVLIVDKEGNTQSQTMNVVFKEIFQNL 1239
>At3g30560 hypothetical protein
Length = 1473
Score = 44.3 bits (103), Expect = 1e-05
Identities = 20/35 (57%), Positives = 27/35 (77%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV S+ GLK+LITD G + TTNVV+KE+F+N+
Sbjct: 1439 RVKSKGGLKVLITDSKGKQKNETTNVVFKEIFRNL 1473
>At4g03690 hypothetical protein
Length = 570
Score = 42.0 bits (97), Expect = 6e-05
Identities = 19/35 (54%), Positives = 25/35 (71%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RV S+ LK+LITD G TTNV++KE+FQN+
Sbjct: 535 RVKSKARLKVLITDSKGKQKKETTNVIFKEIFQNL 569
>At3g51700 unknown protein
Length = 344
Score = 35.0 bits (79), Expect = 0.007
Identities = 18/33 (54%), Positives = 25/33 (75%), Gaps = 1/33 (3%)
Query: 19 RVTSRKGLKILITDENG-DCIDNTTNVVYKEVF 50
+V SR GLK+LITD++G + T NVV+KE+F
Sbjct: 300 KVKSRAGLKVLITDKDGKPDQEETKNVVFKELF 332
>At3g30420 hypothetical protein
Length = 837
Score = 29.6 bits (65), Expect = 0.30
Identities = 16/35 (45%), Positives = 22/35 (62%), Gaps = 1/35 (2%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS KGL +L T + + TN+VY+EVF +
Sbjct: 796 RVTSPKGLTVLDTSKKKEG-KYVTNIVYREVFNGL 829
>At2g14470 pseudogene
Length = 1265
Score = 29.6 bits (65), Expect = 0.30
Identities = 13/17 (76%), Positives = 16/17 (93%)
Query: 19 RVTSRKGLKILITDENG 35
RVTS+KGLKILI D++G
Sbjct: 1247 RVTSKKGLKILILDKDG 1263
>At1g54430 hypothetical protein
Length = 1639
Score = 29.6 bits (65), Expect = 0.30
Identities = 16/35 (45%), Positives = 22/35 (62%), Gaps = 1/35 (2%)
Query: 19 RVTSRKGLKILITDENGDCIDNTTNVVYKEVFQNV 53
RVTS KGL +L T + + TN+VY+EVF +
Sbjct: 1598 RVTSPKGLTVLDTSKKKEG-KYVTNIVYREVFNGL 1631
>At3g51690 putative protein
Length = 374
Score = 29.3 bits (64), Expect = 0.39
Identities = 15/37 (40%), Positives = 22/37 (58%)
Query: 14 FWADYRVTSRKGLKILITDENGDCIDNTTNVVYKEVF 50
F A +V SR GLK+LITD++G+ + N + F
Sbjct: 268 FVAISKVKSRAGLKVLITDKDGNPQEEAKNYPFTLAF 304
>At2g05080 putative helicase
Length = 1219
Score = 26.2 bits (56), Expect = 3.3
Identities = 11/17 (64%), Positives = 14/17 (81%)
Query: 19 RVTSRKGLKILITDENG 35
RVTS+ GLKILI ++ G
Sbjct: 1164 RVTSKSGLKILIVNDEG 1180
>At1g35920 hypothetical protein
Length = 567
Score = 26.2 bits (56), Expect = 3.3
Identities = 10/28 (35%), Positives = 18/28 (63%)
Query: 8 QALILFFWADYRVTSRKGLKILITDENG 35
Q IL W Y +R+ ++++I+DE+G
Sbjct: 21 QVKILHSWKQYTSNTRETIELVISDEHG 48
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.140 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,129,581
Number of Sequences: 26719
Number of extensions: 30143
Number of successful extensions: 101
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 83
Number of HSP's gapped (non-prelim): 17
length of query: 53
length of database: 11,318,596
effective HSP length: 29
effective length of query: 24
effective length of database: 10,543,745
effective search space: 253049880
effective search space used: 253049880
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC124953.2


