

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124953.18 + phase: 0
(391 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
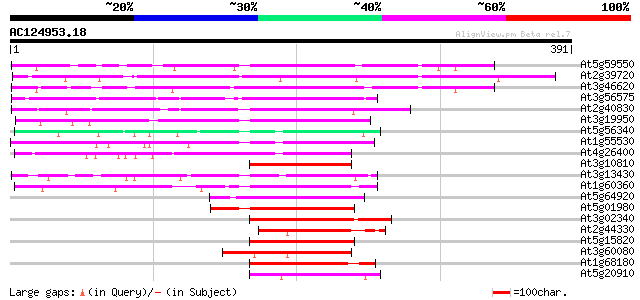
Score E
Sequences producing significant alignments: (bits) Value
At5g59550 unknown protein 280 1e-75
At2g39720 unknown protein 279 2e-75
At3g46620 unknown protein 276 1e-74
At3g56575 unknown protein 129 4e-30
At2g40830 unknown protein 128 5e-30
At3g19950 unknown protein 124 7e-29
At5g56340 unknown protein (At5g56340) 104 7e-23
At1g55530 unknown protein 104 1e-22
At4g26400 unknown protein 100 2e-21
At3g10810 putative RING zinc finger protein 98 9e-21
At3g13430 unknown protein 97 1e-20
At1g60360 95 8e-20
At5g64920 COP1-interacting protein CIP8 90 2e-18
At5g01980 unknown protein 90 2e-18
At3g02340 unknown protein 88 9e-18
At2g44330 putative protein 87 1e-17
At5g15820 putative protein 86 3e-17
At3g60080 unknown protein 85 6e-17
At1g68180 unknown protein 85 6e-17
At5g20910 ABI3-interacting protein 2 84 1e-16
>At5g59550 unknown protein
Length = 407
Score = 280 bits (716), Expect = 1e-75
Identities = 161/367 (43%), Positives = 216/367 (57%), Gaps = 64/367 (17%)
Query: 2 SSHWCYRCNKFVRVWR------LGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAA 55
+S+WCY C +FV VW +G CP CD GF+E + S+ +A + P +
Sbjct: 16 ASYWCYSCTRFVSVWADQGTTTVGSVACPHCDGGFIEQINDSSSAAT-----ELTIPAST 70
Query: 56 AMYMIGHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENNE-- 113
+ I NN ++ RR R G FNP+I+++GG G EREE E
Sbjct: 71 EVRSI----NNNRRSVIRRRR-----SGRRPSFNPVIVLQGGAG-------EREEGEEGD 114
Query: 114 -------FELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEAN-----RSGNEG 161
FE +Y+DG+GSGLR LP +SE+++GSGFER++EQLS +EA+ RSGN
Sbjct: 115 AARDRRAFEFYYDDGSGSGLRPLPDSVSEILMGSGFERLLEQLSQIEASATGIGRSGNP- 173
Query: 162 HNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECIL 221
PA KSA+E LP +EI++ H+ E++CAVC E FE AREMPCKH++H++CI+
Sbjct: 174 ------PASKSAIESLPRVEISDCHIGSEANCAVCTEIFETETEAREMPCKHLFHDDCIV 227
Query: 222 PWLAIQNSCPVCRHELPCESPQINNEISNSNEDEN-VGLTIWRLPGGGFAVGRFSGDGGG 280
PWL+I+NSCPVCR ELP E N SN+NE++N VG+TIWRLPGGGFAVGRF+
Sbjct: 228 PWLSIRNSCPVCRFELPSEP----NRRSNNNEEDNAVGMTIWRLPGGGFAVGRFNAAMRD 283
Query: 281 GENRMEHPIVYTEVDGA--FNNVGEPRRISW-------SLTSSRGGIGRSRGGAFRRMLS 331
GE + P+V TE+DG N+ PRRISW S+ G G GG RRM+
Sbjct: 284 GERVL--PVVLTEMDGGGIGNSQDGPRRISWVRSHGTLESDSNGTGSGSGSGGRLRRMVR 341
Query: 332 NLFGCLR 338
+ +R
Sbjct: 342 GMVSLMR 348
>At2g39720 unknown protein
Length = 401
Score = 279 bits (714), Expect = 2e-75
Identities = 171/407 (42%), Positives = 224/407 (55%), Gaps = 41/407 (10%)
Query: 3 SHWCYRCNKFVRVWRLGMPICPDCDSGFLEDVEQS---------THSANTVGGRRMRFPM 53
S+WCY C++FV W CPDCD GFLE +++ T + T RFP
Sbjct: 5 SYWCYSCSRFV--WVSDSISCPDCDGGFLELIQEPLDFTPSDSFTTTTTTQHRSPTRFPP 62
Query: 54 AAAMYMIG----HRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGS-SEGTSRER 108
++ H +N+ R R N SP NP+I++RG + S E
Sbjct: 63 PSSSSSTPSASMHADNSPTPTIVTRTRSNR------SP-NPVIVLRGSAAAPSSDVVSEG 115
Query: 109 EENNEFELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLP 168
+ + F+++Y+DG SGLR LPP M+E +LGSGF+R+++Q+S +E N + N + +H P
Sbjct: 116 LDRSAFQMYYDDGTDSGLRPLPPSMTEFLLGSGFDRLLDQISQIELNTNRNL-RSCEHPP 174
Query: 169 ALKSAVELLPTIEINESHM--NVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAI 226
A KSA+E LP IEI+ +H+ + +SHCAVCKE F L SAREMPC HIYH +CILPWLAI
Sbjct: 175 ASKSAIEALPLIEIDPTHLLSDSQSHCAVCKENFVLKSSAREMPCNHIYHPDCILPWLAI 234
Query: 227 QNSCPVCRHELPCE--------SPQINNEISNSNEDENVGLTIWRLPGGGFAVGRFSGDG 278
+NSCPVCRHELP E + + ED GLTIWRLPGGGFAVGR G
Sbjct: 235 RNSCPVCRHELPAEDLTDGTGAALTAVTATAEEEEDSAAGLTIWRLPGGGFAVGRIPGGW 294
Query: 279 GGGENRMEHPIVYTEVDGA-FNNVGEPRRISW-SLTSSR-GGIGRSRGGAFRRMLSNLFG 335
GG+ M P+VYTEVDG + PRR++W S R GG R RGG F + LFG
Sbjct: 295 RGGDRMM--PVVYTEVDGGRLGDERLPRRVAWGSRRGGRDGGGSRERGGGFAGRIMRLFG 352
Query: 336 CLRG--GGVRNQHSPFTTREFPQPMTMRNNSASHTNENPSLRSRRTW 380
C G G + + + +T R S S + S RR W
Sbjct: 353 CFSGSSGSIAAAAAASSGSGSRIRVTRRTRSFSMFSTASSSSRRRNW 399
>At3g46620 unknown protein
Length = 395
Score = 276 bits (707), Expect = 1e-74
Identities = 155/355 (43%), Positives = 211/355 (58%), Gaps = 42/355 (11%)
Query: 2 SSHWCYRCNKFVRVWR-----LGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAA 56
+S+WCY C +F+ VW G+ +CP C+ GF+E++E S++S TV P +
Sbjct: 29 TSYWCYSCTRFISVWEDQDANAGV-LCPYCNGGFIEEIEDSSNS--TVAAIPASTPEVRS 85
Query: 57 MYMIGHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENN---- 112
+ +++ RR R N FNP+I++ GGGG G E EE +
Sbjct: 86 V-------EETHRSIIRRRRSNRRTS-----FNPVIVLHGGGGGGAGERVENEEGDGATR 133
Query: 113 ---EFELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPA 169
+E +Y+DG+GSGLR LP +SE+++GSGFER++EQLS +EA SGN + PA
Sbjct: 134 ERRAYEFYYDDGSGSGLRPLPDSVSEILMGSGFERLLEQLSQIEA--SGNGIGRSGNPPA 191
Query: 170 LKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNS 229
KSA+E LP +EI++ H E++CAVC E FE GI REMPCKHI+H +CI+PWL+I+NS
Sbjct: 192 SKSAIESLPRVEISDCHTKAEANCAVCTEVFEAGIEGREMPCKHIFHGDCIVPWLSIRNS 251
Query: 230 CPVCRHELPCESPQINNEISNSNEDENVGLTIWRLPGGGFAVGRFSGDGGGGENRMEHPI 289
CPVCR ELP + Q +NE E+ VG+TIWRLPGGGFAVGRF+ GE + P+
Sbjct: 252 CPVCRFELPSDPIQRSNE-----EEHAVGMTIWRLPGGGFAVGRFNAGVREGERIL--PV 304
Query: 290 VYTEVDGAFNNVGE-PRRISW-----SLTSSRGGIGRSRGGAFRRMLSNLFGCLR 338
V TE+DG E PRRISW + SR G GG RR + + +R
Sbjct: 305 VLTEMDGGGLGSNEGPRRISWVRAHETPEMSRNGGRSGNGGRLRRAVRGMVSFMR 359
>At3g56575 unknown protein
Length = 320
Score = 129 bits (323), Expect = 4e-30
Identities = 83/258 (32%), Positives = 129/258 (49%), Gaps = 18/258 (6%)
Query: 2 SSHWCYRCNKFVRVW-RLGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAAMYMI 60
++HWC+RC + VW R +C C GF+E+++ A+ R F + A
Sbjct: 6 NTHWCHRCQR--AVWLRARDAVCSYCGGGFVEEIDIGPSRAHRDVERDPTFDLMEAFSAF 63
Query: 61 GHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENNEFELFYED 120
+ + ++ R + F+ + + GG + +N+ E F +
Sbjct: 64 --MRSRLAERSYDREISGRLGSAGSESFSNLAPLLIFGGQAP-FRLAGGDNSSVEAFV-N 119
Query: 121 GAGSGLR-ALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPT 179
GA G+ A + G G E ++EQLS SG H++ PA KS+++ LPT
Sbjct: 120 GAAPGIGIARGTNAGDYFFGPGLEELIEQLS------SGT--HHRGPPPAPKSSIDALPT 171
Query: 180 IEINESHM-NVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELP 238
I+I + H+ + +SHC VCK+ FEL A++MPC HIYH++CI+PWL NSCPVCR ELP
Sbjct: 172 IKITQKHLKSSDSHCPVCKDEFELKSEAKQMPCHHIYHSDCIVPWLVQHNSCPVCRKELP 231
Query: 239 CESPQINNEISNSNEDEN 256
+ + S+ N N
Sbjct: 232 SRGSSSSTQ-SSQNRSTN 248
>At2g40830 unknown protein
Length = 328
Score = 128 bits (322), Expect = 5e-30
Identities = 92/307 (29%), Positives = 137/307 (43%), Gaps = 45/307 (14%)
Query: 2 SSHWCYRCNKFVRVWRLGMPICPDCDSGFLEDVEQST-------HSANTVGGRRMRFPMA 54
++HWC+RC + VR+ P+C C GF+E+++ + S V R F +
Sbjct: 6 NTHWCHRCQRAVRLHGQE-PVCFYCGGGFVEELDMAQASPFDMFRSHRGVVERDQTFDLM 64
Query: 55 AAMYMIGHRNNNYNQNTFRRHRRNNVNGGDIS-PFNPIIMIRGGGGSSEGTSREREENNE 113
A + + RN ++ R R ++ G + P ++I GG T +N
Sbjct: 65 DA-FSVFMRNRLAERSHDREIRGRTISSGPENFPGLAPLLIFGGQVPYRLTG-----DNA 118
Query: 114 FELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLPALKSA 173
E + +G G+ + G G E + EQLS R PA +SA
Sbjct: 119 VEALF-NGGSPGIGITRGNTGDYFFGPGLEELFEQLSAGTTRRGPP--------PAPRSA 169
Query: 174 VELLPTIEINESHM-NVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPV 232
++ LPTI+I + H+ + +S+C VCK+ FELG A++MPC HIYH++CI+PWL NSCPV
Sbjct: 170 IDALPTIKIAQRHLRSSDSNCPVCKDEFELGSEAKQMPCNHIYHSDCIVPWLVQHNSCPV 229
Query: 233 CRHELP--------------------CESPQINNEISNSNEDENVGLTIWRLPGGGFAVG 272
CR ELP S +N N NE N + W G +
Sbjct: 230 CRQELPSASGPSSSQNRTTPTRNYRSSSSSSSSNSRENGNERRNPFSSFWPFRSSGSSSS 289
Query: 273 RFSGDGG 279
GG
Sbjct: 290 STQNRGG 296
>At3g19950 unknown protein
Length = 328
Score = 124 bits (312), Expect = 7e-29
Identities = 76/266 (28%), Positives = 118/266 (43%), Gaps = 32/266 (12%)
Query: 5 WCYRCNKFVRVWRLGM--PICPDCDSGFLEDVEQSTHSAN---TVGGRRMRFPMA----- 54
+CY+CN+ V + P CP C+ GFLE+ E + + FPMA
Sbjct: 22 FCYQCNQTVTISISSSADPFCPICNQGFLEEYEDPNPNQSLNFNPNSSDSFFPMADPFST 81
Query: 55 --------AAMYMIGHRNNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSR 106
+A G + + + R+ F+P ++
Sbjct: 82 LLPLIFGSSAAAPSGMDFMSLFGPSMQPQARSTQQNPQSDAFDPFTFLQNH------LQT 135
Query: 107 EREENNEFELFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQH 166
R FE E+ +P + G G E++++QL+ + NR G
Sbjct: 136 LRSSGTHFEFVIENHPSDPGNRMPGNFGDYFFGPGLEQLIQQLAENDPNRYGTP------ 189
Query: 167 LPALKSAVELLPTIEINESHMNVE-SHCAVCKEPFELGISAREMPCKHIYHNECILPWLA 225
PA KSA++ LPT+++ + + E + CAVC + FE G ++MPCKH++H +C+LPWL
Sbjct: 190 -PASKSAIDALPTVKVTKDMLKSEMNQCAVCMDEFEDGSDVKQMPCKHVFHQDCLLPWLE 248
Query: 226 IQNSCPVCRHELPCESPQINNEISNS 251
+ NSCPVCR ELP + P N S
Sbjct: 249 LHNSCPVCRFELPTDDPDYENRSQGS 274
>At5g56340 unknown protein (At5g56340)
Length = 396
Score = 104 bits (260), Expect = 7e-23
Identities = 92/345 (26%), Positives = 133/345 (37%), Gaps = 104/345 (30%)
Query: 4 HWCYRCNKFVRVWRLGMPICPDCDSGFLE---------------DVEQSTHSANTVGGRR 48
+WC+ C++ V CP C SGF+E DV+ S T
Sbjct: 9 YWCHMCSQMVNPVMESEIKCPFCQSGFIEEMSGNGGGGGGRGIRDVQDSETDFGTDRALS 68
Query: 49 MRFPMAAAMYMI------------------------------GHRNNNYNQNTFRRHRRN 78
+ P+ M G+ +NN N N +R HR
Sbjct: 69 LWAPILLGMMSSPRRRRRFRRSEFGEENDDNGDDLTTNADGNGNDSNNSNNNVYRHHRAR 128
Query: 79 NVNGGDI---------------SPFNPIIMIRG--GGGSSEGTSREREENNEFE------ 115
+GG+I S N + +++G G +SE S + +N E
Sbjct: 129 R-HGGEIDLDREFESILRRRRRSSGNILQLLQGIRAGIASEYESSDNNWDNSRERDRVIM 187
Query: 116 -------LFYEDGAGSGLRALPPRMSELILGSGFERVMEQLSHVEANRSGNEGHNQQHLP 168
L +L + + +G G + +++ L+ + NR G P
Sbjct: 188 INPYNQSLVVPSDQNQNHPSLTS-LGDYFIGPGLDLLLQHLAENDPNRQGTP-------P 239
Query: 169 ALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQN 228
A K AVE LPT++I E C+VC + FE G A+EMPCKH +H CI+PWL + +
Sbjct: 240 ARKEAVEALPTVKIMEP-----LQCSVCLDDFEKGTEAKEMPCKHKFHVRCIVPWLELHS 294
Query: 229 SCPVCRHELPCESPQIN---------------NEISNSNEDENVG 258
SCPVCR ELP + + E SN N ENVG
Sbjct: 295 SCPVCRFELPSSADDDDETKTDSERVLRTRNVRETSNGNVVENVG 339
>At1g55530 unknown protein
Length = 351
Score = 104 bits (259), Expect = 1e-22
Identities = 75/291 (25%), Positives = 134/291 (45%), Gaps = 49/291 (16%)
Query: 1 MSSHWCYRCNKFVRVWRLGMPICPDCDSGFLEDVEQST-HSANTVGGRRMRFPMAAAMYM 59
++ +WC+ C++ V CP C SGF+E++E H ++ R + A + M
Sbjct: 6 VTRYWCHMCSQTVNPVMEAEIKCPFCQSGFVEEMEDDDDHDSSDPADVRANNSLWAPILM 65
Query: 60 ------IGHRNNNY------NQNTFRRHRRNNVNGGDIS-PFNPII------------MI 94
+ R N NQN + + D+ I+ ++
Sbjct: 66 ELMNDPVRRRRNQSVESVEDNQNEVQTENNEDDGENDLDWQLQEILRRRRRHSAAVLQLL 125
Query: 95 RG--GGGSSEGTSREREENNEFELFYEDGAG--------SGLRALPP-RMSELILGSGFE 143
+G G S E S NN + + + + + ++P + + +G GFE
Sbjct: 126 QGIRAGLSVESESTGNGGNNPGRVILINTSNQTITVQNSADMDSVPAGSLGDYFIGPGFE 185
Query: 144 RVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPFELG 203
++++L+ + NR G PA K AVE L T++I E+ C+VC + FE+G
Sbjct: 186 MLLQRLAENDPNRYGTP-------PAKKEAVEALATVKIEET-----LQCSVCLDDFEIG 233
Query: 204 ISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCESPQINNEISNSNED 254
A+ MPC H +H++C+LPWL + +SCPVCR++LP + + ++ + S+ +
Sbjct: 234 TEAKLMPCTHKFHSDCLLPWLELHSSCPVCRYQLPADEAKTDSVTTTSDNN 284
>At4g26400 unknown protein
Length = 356
Score = 99.8 bits (247), Expect = 2e-21
Identities = 79/289 (27%), Positives = 129/289 (44%), Gaps = 61/289 (21%)
Query: 4 HWCYRCNKFVRVWRLGMPI-CPDCDSGFLEDVEQSTHSANTVGGRRMRFP-------MAA 55
+WC+ C++ V +G I CP C SGF+E++ + + ++ R ++ P A
Sbjct: 5 YWCHMCSQMVNPV-IGAEIKCPFCQSGFVEEMSREINGGSSSSLREVQDPEIDFGTDRAL 63
Query: 56 AMY---MIGHRNNNYNQNTFRR----------------------------HRRNN--VNG 82
+++ ++G +N + FRR HRR+ G
Sbjct: 64 SLWGPILLGMMSNPRRRRRFRRTEFGVDNDEVNVSDDVDGNDSVDIDRHHHRRHRHRQQG 123
Query: 83 GDIS---PFNPIIMIRGGG---------GSSEGTSREREENNEFELFYEDGAGSGLRALP 130
+I F I+ R G G + E E ++ +L + G +L
Sbjct: 124 REIDLDREFESILRRRRRSSATILQLLQGIRAGIASEYESSDR-DLLNQSAVVQGSTSLN 182
Query: 131 PRMSELILGS-GFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNV 189
+ L + G V L H+ + + N+ LPA K V+ LPT++I+ES
Sbjct: 183 QNRNNTSLSAIGDYFVGSSLDHLLEHLADNDSIRHGSLPARKEVVDNLPTVKISES---- 238
Query: 190 ESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELP 238
C++C + F+ G A+EMPCKH +H CI+PWL + +SCPVCR+ELP
Sbjct: 239 -LQCSICLDDFDKGSEAKEMPCKHKFHIRCIVPWLELHSSCPVCRYELP 286
>At3g10810 putative RING zinc finger protein
Length = 684
Score = 97.8 bits (242), Expect = 9e-21
Identities = 39/71 (54%), Positives = 54/71 (75%)
Query: 168 PALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQ 227
PA +A+ L I+I + H+ ++ +C VC++ FE+G AR+MPCKHIYH+ECILPWL +
Sbjct: 96 PASLAAINSLQKIKIRQKHLGLDPYCPVCQDQFEIGSDARKMPCKHIYHSECILPWLVQR 155
Query: 228 NSCPVCRHELP 238
N+CPVCR ELP
Sbjct: 156 NTCPVCRKELP 166
>At3g13430 unknown protein
Length = 315
Score = 97.4 bits (241), Expect = 1e-20
Identities = 81/313 (25%), Positives = 144/313 (45%), Gaps = 87/313 (27%)
Query: 2 SSHWCYRCNKFVRVWRLGMPICPD-------CDSGFLEDVEQST-HSANTVGGRRMRFPM 53
+S+WC+ C++ V +P+ D C SGF+E+++ + H A + P+
Sbjct: 7 TSYWCHMCSRSV------IPLIQDEIIKCNFCQSGFVEEMDNNDDHQA----ADSLWTPI 56
Query: 54 AAAMYMIGHRNNNYNQNTFRRHRRNNV-----------------NGGDI----------- 85
M NNN++Q++ + ++ N G+I
Sbjct: 57 LMEMM-----NNNHDQHSTNQEDSESILEDEDEDEDDGDDGDQNNDGEIDITHQLEEIRR 111
Query: 86 -------SPFNPIIMIRGGGGSSEGTSREREENNEFELF----------YEDGAGSGLRA 128
+ N + IR G + + +N+E + ++D + +
Sbjct: 112 IRTRHSTAIVNLLQGIRAGLLIESENNEDNPDNSELVVLINSFNQRIRVHQDSVDT--TS 169
Query: 129 LPP-RMSELILGSGFERVMEQLSHVEAN-RSGNEGHNQQHLPALKSAVELLPTIEINESH 186
+P + + +G GFE ++++L+ + N R G PA K AVE L ++I +S
Sbjct: 170 VPSGSLGDYFIGPGFETLLQRLAENDLNNRYGTP-------PATKEAVEALAMVKIEDSL 222
Query: 187 MNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPC---ESPQ 243
+ C+VC + FE+G+ A+EMPCKH +H++C+LPWL + +SCPVCR+ LP + P+
Sbjct: 223 LQ----CSVCLDDFEIGMEAKEMPCKHKFHSDCLLPWLELHSSCPVCRYLLPTGDDDEPK 278
Query: 244 INNEISNSNEDEN 256
+ E S N+D N
Sbjct: 279 TDAETSR-NDDNN 290
>At1g60360
Length = 327
Score = 94.7 bits (234), Expect = 8e-20
Identities = 76/289 (26%), Positives = 118/289 (40%), Gaps = 66/289 (22%)
Query: 4 HWCYRCNKFVRVWRLGMP--ICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAAMYMIG 61
+WCY C++ VR+ CP C F+ ++E T F + ++
Sbjct: 24 YWCYHCDRMVRIASSNPSEIACPRCLRQFVVEIETRQRPRFTFNHATPPFDASPEARLLE 83
Query: 62 HRNNNYNQNTF----------------------------RRHRRNNVNGGDIS-PFNPII 92
+ + T RRH +NVN G + P +
Sbjct: 84 ALSLMFEPATIGRFGADPFLRARSRNILEPESRPRPQHRRRHSLDNVNNGGLPLPRRTYV 143
Query: 93 MIRGGGGSSEGTSREREENNEFELFYEDGAGSGLRALPPR---MSELILG-SGFERVMEQ 148
++R +S + N PPR + G S E+++EQ
Sbjct: 144 ILRPNNPTSPLGNIIAPPNQA----------------PPRHVNSHDYFTGASSLEQLIEQ 187
Query: 149 LSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHM-NVESHCAVCKEPFELGISAR 207
L+ + +R G PA + + LP+++I H+ N S C VC E F +G A
Sbjct: 188 LT--QDDRPGPP-------PASEPTINSLPSVKITPQHLTNDMSQCTVCMEEFIVGGDAT 238
Query: 208 EMPCKHIYHNECILPWLAIQNSCPVCRHELPCESPQINNEISNSNEDEN 256
E+PCKHIYH +CI+PWL + NSCP+CR +LP + N ++ S E N
Sbjct: 239 ELPCKHIYHKDCIVPWLRLNNSCPICRRDLP-----LVNTVAESRERSN 282
>At5g64920 COP1-interacting protein CIP8
Length = 334
Score = 90.1 bits (222), Expect = 2e-18
Identities = 40/108 (37%), Positives = 64/108 (59%), Gaps = 3/108 (2%)
Query: 140 SGFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEP 199
+G+E +++ L+ + G G + PA KSA+E L T E++ S + CAVCK+
Sbjct: 207 AGYEALLQNLAEGDG---GGGGGRRGAPPAAKSAIEALETFEVSSSEGEMVMVCAVCKDG 263
Query: 200 FELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCESPQINNE 247
+G + +++PC H YH +CI+PWL +NSCPVCR +L + + E
Sbjct: 264 MVMGETGKKLPCGHCYHGDCIVPWLGTRNSCPVCRFQLETDDAEYEEE 311
Score = 39.7 bits (91), Expect = 0.003
Identities = 16/35 (45%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Query: 2 SSHWCYRCNKFVRVWRL-GMPICPDCDSGFLEDVE 35
+SHWCY CNK V V L +C +C+ GF+E ++
Sbjct: 13 ASHWCYHCNKRVVVETLDDFVVCCECNKGFVESIQ 47
>At5g01980 unknown protein
Length = 493
Score = 89.7 bits (221), Expect = 2e-18
Identities = 42/100 (42%), Positives = 60/100 (60%), Gaps = 7/100 (7%)
Query: 141 GFERVMEQLSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPF 200
GF+ ++EQL+ + +R G PA S V LP + I E H+ CA+CKE F
Sbjct: 305 GFDELLEQLAESDNSRRGAP-------PASVSCVRNLPRVIIAEEHVMKGLVCAICKELF 357
Query: 201 ELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCE 240
L ++PC H+YH CI+PWL+ +NSCP+CR+ELP +
Sbjct: 358 SLRNETTQLPCLHLYHAHCIVPWLSARNSCPLCRYELPTD 397
>At3g02340 unknown protein
Length = 409
Score = 87.8 bits (216), Expect = 9e-18
Identities = 42/100 (42%), Positives = 61/100 (61%), Gaps = 3/100 (3%)
Query: 168 PALKSAVELLPTIEINESHMNVESH-CAVCKEPFELGISAREMPCKHIYHNECILPWLAI 226
PA KS ++ LP +E+ ++ ++ CAVCK+ + R +PC H YH ECI+PWL I
Sbjct: 309 PAAKSVIQDLPVVELAVEELDKGNNVCAVCKDEMLVEEKVRRLPCSHFYHGECIIPWLGI 368
Query: 227 QNSCPVCRHELPCESPQINNEISNSNEDENVGLTIWRLPG 266
+N+CPVCR+ELP + + E S+E + GL LPG
Sbjct: 369 RNTCPVCRYELPTD--DLEYERHKSSERGDTGLARNVLPG 406
>At2g44330 putative protein
Length = 180
Score = 87.4 bits (215), Expect = 1e-17
Identities = 42/95 (44%), Positives = 60/95 (62%), Gaps = 17/95 (17%)
Query: 174 VELLPTIEINESHMNVESH------CAVCKEPFELGISAREMPCKHIYHNECILPWLAIQ 227
+E LPTI+I+ S ++ S CA+C+E F +G SAR +PC H+YHN+CI+PWL
Sbjct: 71 MESLPTIKISSSMLSSASSDDSALPCAICREDFVVGESARRLPCNHLYHNDCIIPWLTSH 130
Query: 228 NSCPVCRHELPCESPQINNEISNSNEDENVGLTIW 262
NSCP+CR ELP +++S +D GL +W
Sbjct: 131 NSCPLCRVELP---------VASSEDDS--GLDMW 154
>At5g15820 putative protein
Length = 348
Score = 86.3 bits (212), Expect = 3e-17
Identities = 37/74 (50%), Positives = 49/74 (66%), Gaps = 1/74 (1%)
Query: 168 PALKSAVELLPTIEIN-ESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAI 226
PA KS V+ LP +E+ E +V CA+CK+ + +PCKH YH ECI+PWL I
Sbjct: 265 PASKSVVDGLPDVELTIEELSSVSIVCAICKDEVVFKEKVKRLPCKHYYHGECIIPWLGI 324
Query: 227 QNSCPVCRHELPCE 240
+N+CPVCRHELP +
Sbjct: 325 RNTCPVCRHELPTD 338
>At3g60080 unknown protein
Length = 306
Score = 85.1 bits (209), Expect = 6e-17
Identities = 43/101 (42%), Positives = 63/101 (61%), Gaps = 11/101 (10%)
Query: 149 LSHVEANRSGNEGHNQQHLPA--LKSA-VELLPTIEINESHMNVESH--------CAVCK 197
L H+ ++ SG+ + + LKS+ ++ +PTI+I+ S + CAVCK
Sbjct: 114 LRHLASDNSGSSSSSSSSSSSSLLKSSDIDSIPTIQISSSLLCSTDDSDPDSVLLCAVCK 173
Query: 198 EPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELP 238
E F +G SAR +PC HIYH++CI+PWL+ NSCP+CR ELP
Sbjct: 174 EDFIIGESARRLPCSHIYHSDCIVPWLSDHNSCPLCRFELP 214
>At1g68180 unknown protein
Length = 145
Score = 85.1 bits (209), Expect = 6e-17
Identities = 38/88 (43%), Positives = 55/88 (62%), Gaps = 7/88 (7%)
Query: 168 PALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQ 227
PA +SA+E + T+ I + + E CA+CKE FE+G +E+ C H+YH+ CI+ WL I
Sbjct: 10 PASQSAIEAVRTVIITDEDLVKEKVCAICKEEFEVGEEGKELKCLHLYHSSCIVSWLNIH 69
Query: 228 NSCPVCRHELPCESPQINNEISNSNEDE 255
N+CP+CR E +N +S SN DE
Sbjct: 70 NTCPICRFE-------VNLGVSESNVDE 90
>At5g20910 ABI3-interacting protein 2
Length = 310
Score = 84.0 bits (206), Expect = 1e-16
Identities = 40/96 (41%), Positives = 52/96 (53%), Gaps = 5/96 (5%)
Query: 168 PALKSAVELLPTIEINESHMN---VESHCAVCKEPFELGISAREMPCKHIYHNECILPWL 224
PA K VE LP I E + E+ C +CKE +G +E+PCKH +H C+ PWL
Sbjct: 202 PASKEVVEKLPVIIFTEELLKKFGAEAECCICKENLVIGDKMQELPCKHTFHPPCLKPWL 261
Query: 225 AIQNSCPVCRHELPCESPQINN--EISNSNEDENVG 258
NSCP+CRHELP + + N E E+E G
Sbjct: 262 DEHNSCPICRHELPTDDQKYENWKEREKEAEEERKG 297
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,171,705
Number of Sequences: 26719
Number of extensions: 496639
Number of successful extensions: 2256
Number of sequences better than 10.0: 277
Number of HSP's better than 10.0 without gapping: 218
Number of HSP's successfully gapped in prelim test: 59
Number of HSP's that attempted gapping in prelim test: 1874
Number of HSP's gapped (non-prelim): 364
length of query: 391
length of database: 11,318,596
effective HSP length: 101
effective length of query: 290
effective length of database: 8,619,977
effective search space: 2499793330
effective search space used: 2499793330
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC124953.18


