

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124952.2 - phase: 0
(880 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
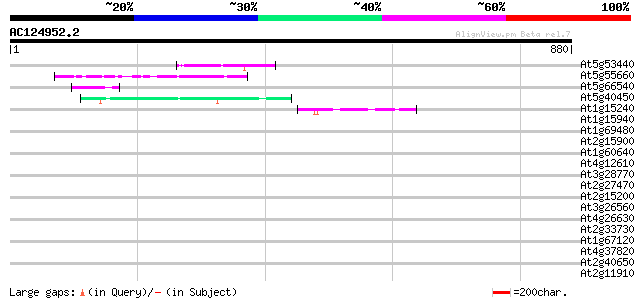
Score E
Sequences producing significant alignments: (bits) Value
At5g53440 putative protein 45 2e-04
At5g55660 putative protein 44 3e-04
At5g66540 unknown protein 44 5e-04
At5g40450 unknown protein 44 5e-04
At1g15240 hypothetical protein 43 9e-04
At1g15940 T24D18.4 42 0.001
At1g69480 hypothetical protein 42 0.002
At2g15900 hypothetical protein 41 0.003
At1g60640 unknown protein 41 0.003
At4g12610 putative protein 41 0.003
At3g28770 hypothetical protein 40 0.004
At2g27470 putative CCAAT-binding transcription factor subunit 40 0.004
At2g15200 putative protein on transposon FARE2.7 (cds2) 40 0.004
At3g26560 ATP-dependent RNA helicase, putative 40 0.006
At4g26630 unknown protein 40 0.007
At2g33730 putative U5 small nuclear ribonucleoprotein, an RNA he... 39 0.010
At1g67120 hypothetical protein 39 0.010
At4g37820 unknown protein 39 0.013
At2g40650 unknown protein 39 0.013
At2g11910 unknown protein 39 0.013
>At5g53440 putative protein
Length = 1181
Score = 45.1 bits (105), Expect = 2e-04
Identities = 47/170 (27%), Positives = 73/170 (42%), Gaps = 18/170 (10%)
Query: 262 EETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNIT 321
E T+D DR+ E RDRDRDRDR+RDRD + D D D + D + T
Sbjct: 321 ERTRD-RDRD-YERDRDRDRDRDRERDRDRRDYEHDRYHDRDWDRDRSRDRDRDHERDRT 378
Query: 322 DGGSEGKGNRYSKNDEAGASGDAQREN-LDLDNFEFKSDQFCDNRD-----------DVS 369
+ + Y +D + D +R+N D+ + +S ++ D RD DV
Sbjct: 379 HDREKDRSRDY-YHDGKRSKSDRERDNDRDVSRLDDQSGRYKDRRDGRRSPDYQDYQDVI 437
Query: 370 TSNVSVHVE---NVDAKSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKS 416
T + S VE ++ + +V NG + + ++S V E S
Sbjct: 438 TGSRSSRVEPDGDMTRPERQLSSSVVQEENGNASDQITKGASSREVAELS 487
Score = 32.7 bits (73), Expect = 0.92
Identities = 58/312 (18%), Positives = 114/312 (35%), Gaps = 46/312 (14%)
Query: 76 TFDDSNRKSFQYGSSGLELYG----DEGDELSMTGLDSSELIGNNGIEESDGNENGGEVG 131
T S R+ + G SG + + D+G+ T + S + + +E DG E G
Sbjct: 69 TSSSSKRRKGKSGESGSDRWNGKDDDKGESSKKTKVSSEK---SRKRDEGDGEETKKSSG 125
Query: 132 ERE-----------IETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFY 180
+ + + ++E++ ++ EG F G D ++++ S K E+ + +
Sbjct: 126 KSDGKHRESSRRESKDVDKEKDRKYKEGKSDKFYDG---DDHHKSKAGSDKTESKAQDHA 182
Query: 181 SSRSVSLYEEGEVRNENPLLMNSSVAF-GSHDLDDFLLQNGPVSVVSDLFYNPRESNNRV 239
S Y E R + S D+ D +L +G + D + +S ++
Sbjct: 183 RSPGTENYTEKRSRRKRDDHGTGDKHHDNSDDVGDRVLTSGD-DYIKDGKHKGEKSRDKY 241
Query: 240 EEHGVSSVRKEE---------------KDSLIVNDEVEETKDIGDRE-------ALEEVR 277
E K++ D + DE ++ D + L+
Sbjct: 242 REDKEEEDIKQKGDKQRDDRPTKEHLRSDEKLTRDESKKKSKFQDNDHGHEPDSELDGYH 301
Query: 278 DRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKND- 336
+R+R+RD DR+ D + D D + + D + + + +RY D
Sbjct: 302 ERERNRDYDRESDRNERDRERTRDRDRDYERDRDRDRDRDRERDRDRRDYEHDRYHDRDW 361
Query: 337 EAGASGDAQREN 348
+ S D R++
Sbjct: 362 DRDRSRDRDRDH 373
>At5g55660 putative protein
Length = 759
Score = 44.3 bits (103), Expect = 3e-04
Identities = 64/304 (21%), Positives = 123/304 (40%), Gaps = 44/304 (14%)
Query: 71 ASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEV 130
+S +T D + K Q +SG E+ + +E TGL+ E+ +E +E E
Sbjct: 18 SSLQKTSDAISGKEVQENASGKEVQESKKEE--DTGLEKMEID-----DEGKQHEGESET 70
Query: 131 GEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEE 190
G++E+E EEE+++ G+D D D +++ ++K ++ S
Sbjct: 71 GDKEVEVTEEEKKDV--GEDKEQPEADKMDEDTDDK--NLKADDGVS------------- 113
Query: 191 GEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKE 250
V E +M SV S D D G +ES E G ++ +E
Sbjct: 114 -GVATEEDAVMKESVE--SADNKDAENPEGE---------QEKESKEEKLEGGKANGNEE 161
Query: 251 --EKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDL 308
++ L+ D+ + D+ + E +E V + D++ +A ++ E + +
Sbjct: 162 GDTEEKLVGGDKGD---DVDEAEKVENVDEDDKEEALKEKNEAELAEEEETNKGEEVKEA 218
Query: 309 LPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDV 368
E+D + + + E K +E + ++E+ D E +D D ++D+
Sbjct: 219 NKEDDVEADTKVAEPEVEDKKTESKDENE---DKEEEKEDEKEDEKEESNDDKEDKKEDI 275
Query: 369 STSN 372
SN
Sbjct: 276 KKSN 279
>At5g66540 unknown protein
Length = 524
Score = 43.5 bits (101), Expect = 5e-04
Identities = 29/78 (37%), Positives = 41/78 (52%), Gaps = 12/78 (15%)
Query: 97 DEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYG 156
D+ DE+ M G DS ++ + ES+ +E E E E E EEEEEEE E +
Sbjct: 109 DDIDEMDMDGFDSDDVDDEDKEIESNDSEGEDEEEEEEDEEEEEEEEEEEEEEK------ 162
Query: 157 SGCDGDN---ENEFYSMK 171
DGDN E++F+ +K
Sbjct: 163 ---DGDNEGIEDKFFKIK 177
>At5g40450 unknown protein
Length = 2910
Score = 43.5 bits (101), Expect = 5e-04
Identities = 67/343 (19%), Positives = 136/343 (39%), Gaps = 26/343 (7%)
Query: 111 ELIGNNGIEESDGNENGGEVGEREIETEEE-----EEEEFSEGDDSMFNYGSGCDGDNEN 165
E I + EE ++ G E G ++T++E E E E C + +
Sbjct: 1532 EKIEDKPKEEVTLHQEGREEGSYGLDTKDEAVSVLESRELGEQPQQE----ELCLANEQE 1587
Query: 166 EFYSMKGENVSSEFYSSRSVSLYEEGEVRN-ENPLLMNSSVAFGSHDLDDFLLQNGPVSV 224
++ E V + VS ++ V N ++ SS + + +++ +
Sbjct: 1588 NETKLQEEQVDKHEPTKEEVSNDQQSPVEEISNEVIQVSSASLSEGPEYETVVEAEKIGE 1647
Query: 225 --VSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRD 282
V+D E+ VE H EEK+ V+++ ++ K + D E + ++R+R D
Sbjct: 1648 EQVADKIQKSFETGEIVEAHSSLPSSSEEKEHETVSEKTDDEK-VKDAEPIGDMRERGLD 1706
Query: 283 RDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGG----SEGKGNRYSKNDEA 338
P + + D + D Q+ + T+ G SE + ++ ++ ++
Sbjct: 1707 IAETTHLSLPSVDQKEDVDEIHIPSVALPLDEQEKVTSTEKGETKSSEAEDDKPDEHVDS 1766
Query: 339 GASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSFKNLKPIVLPSNGG 398
S +N + S+ C +++ T V E+ + KS + + I
Sbjct: 1767 STSPMLSEKNDNETQTSKTSEDVCMQQEESGTLEVPKPEESKEDKSQEISETI------- 1819
Query: 399 TRKILERSSTSTNVLEKSHVISKIEDSELSEFYDEVVQEMEEI 441
+ +E +S T +E SH + + +SE D+ +++EEI
Sbjct: 1820 --EEIEATSDQTLPIETSHTDNTLSSELVSEQDDQSPKKVEEI 1860
Score = 33.5 bits (75), Expect = 0.54
Identities = 84/389 (21%), Positives = 147/389 (37%), Gaps = 82/389 (21%)
Query: 119 EESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSE 178
+ES G+E EV +E E EE+ D +M + SG E+E Y ++S
Sbjct: 1427 DESQGSEESVEVKSKETVQGESSEEK----DVNMLDVQSG-----ESEKYQENEPDIS-- 1475
Query: 179 FYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRES--- 235
VS E G+ E P ++ + G + SVV P E
Sbjct: 1476 -----LVSKTENGDKFEEIPSVVEGA---GLDETTHNQTLLDVESVVKQSLDTPSEEETS 1527
Query: 236 ---NNRVEEHGVSSV------RKEEKDSLIVNDE---VEETKDIGDREALEEV------R 277
+ ++E+ V R+E L DE V E++++G++ EE+
Sbjct: 1528 KTIDEKIEDKPKEEVTLHQEGREEGSYGLDTKDEAVSVLESRELGEQPQQEELCLANEQE 1587
Query: 278 DRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDE 337
+ + ++ D+ P EV SP EE + + ++ S +G Y E
Sbjct: 1588 NETKLQEEQVDKHEPTKEEVSNDQQSP-----VEEISNEVIQVS-SASLSEGPEYETVVE 1641
Query: 338 AGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVENV----DAKSFKNLKPIVL 393
A G+ Q D F++ + + + +S+ E V D + K+ +PI
Sbjct: 1642 AEKIGEEQ--VADKIQKSFETGEIVEAHSSLPSSSEEKEHETVSEKTDDEKVKDAEPI-- 1697
Query: 394 PSNGGTRKILERSSTSTNVLEKSHVISKIEDSELSEFYDEVVQEMEEILLESMDSPAARF 453
G R+ ++ E +H+ D + ++++EI + S+ P
Sbjct: 1698 ---GDMRE------RGLDIAETTHLSLPSVDQK---------EDVDEIHIPSVALP---- 1735
Query: 454 SVGNRMFDPQQSVPSRDGGLTASTSSTDD 482
D Q+ V S + G T S+ + DD
Sbjct: 1736 ------LDEQEKVTSTEKGETKSSEAEDD 1758
Score = 32.3 bits (72), Expect = 1.2
Identities = 42/191 (21%), Positives = 80/191 (40%), Gaps = 10/191 (5%)
Query: 97 DEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYG 156
D+ ++ T ++ + IG + E N+N E +E ++EE+E+E S +++ N
Sbjct: 1104 DDATKIHETRVEQARDIGPSLTEICSINQNQPEEQVKEACSKEEQEKEISTNSENIVNET 1163
Query: 157 SGCDG-DNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDF 215
+ E + GE++ + +++SV L EVR E + A D +
Sbjct: 1164 YALHSVEAAEEETATNGESL-DDVETTKSVLL----EVRKEEEEAEMKTDAEPRLDAIEK 1218
Query: 216 LLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEE 275
+VV D E E + +EK++ + VE T+++ D E +
Sbjct: 1219 EELETVKTVVQDAKIVNNEETTAHESESLKGDNHQEKNA----EPVEATQNLDDAEQISR 1274
Query: 276 VRDRDRDRDRD 286
D +R+ D
Sbjct: 1275 EVTVDTEREAD 1285
Score = 31.2 bits (69), Expect = 2.7
Identities = 61/320 (19%), Positives = 126/320 (39%), Gaps = 22/320 (6%)
Query: 100 DELSMTGLDSSELIGNNGIEESDG-NENGGEVGEREIETEEEEEEEFSEGDDSMFNYGS- 157
+EL+ ++ ++ G +EE G + NG E + + E EE S+ + S
Sbjct: 799 EELNDEKINVDQVDGTQIMEEPIGLDSNGAEAEQIDQNITNETEEILVAKPVSLLDVKSV 858
Query: 158 ------GCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAF--GS 209
+ +E + K + E V+LY+EG+V L S
Sbjct: 859 EQMQKPKLESPSEVSEETSKTVDEKIEEKPEEEVTLYQEGQVDGSYGLETKEETVSVPES 918
Query: 210 HDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRK---EEKDSLIVNDEVEETKD 266
+L++ + V + L ES + V E +V + E+ DS+ + + +E +
Sbjct: 919 IELEEQPQEERSVIDPTPLQKPTLESPSEVLEESSKTVDEKIEEKTDSIELGEIAQEERS 978
Query: 267 IGDREALEEVRDRDRDRDRDR--DRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGG 324
+ D L+E + +++++ ++ P + EV+ + L P ++ + +
Sbjct: 979 VTDLTPLQEESSQPNEQEKETKLEKHEPTNEEVKSDEVIEVLSASPSKELEGETVVEAEN 1038
Query: 325 SEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVDAKS 384
E +N+E A+ Q+ + E S + + +V+V E VD K+
Sbjct: 1039 IE----NIKENEEEQAAEKIQKSLETVQTVESPSSLLFSSEEQ---DHVTVAEEIVDEKA 1091
Query: 385 FKNLKPIVLPSNGGTRKILE 404
+ + + + + KI E
Sbjct: 1092 KEEVPMLQIKNEDDATKIHE 1111
>At1g15240 hypothetical protein
Length = 906
Score = 42.7 bits (99), Expect = 9e-04
Identities = 56/209 (26%), Positives = 88/209 (41%), Gaps = 47/209 (22%)
Query: 452 RFSVGNRMFDPQQSVPSRDGGLTAS------TSSTD-------------DACLLVKRPRR 492
R + + D S S DG LT S TSST + LLV +
Sbjct: 553 RSRISGHIIDDDDSDDSEDGSLTRSYSGMSATSSTSYVSAAESDLPNAPKSSLLVDSFAK 612
Query: 493 IDRIEVVGARQKRGDVSFSERLVGVKEYTVYKIKVWSGKDQ-WEVEKRYRDFLTLHRCMK 551
+ R EV+GA +G K + VY + V + W +++R+R F LHR +K
Sbjct: 613 L-RCEVLGANIVKGSS---------KMFAVYSVAVTDESNHSWSIKRRFRHFEELHRRLK 662
Query: 552 SLFNEQGWTLPLPWSSVEKESKIFRSASLDI--IAKRSVLIQECLQSILCSRFFSSPPRA 609
+F E LP K F S +DI I +R VL+ E +++ S FS
Sbjct: 663 -VFPEYKLHLP---------PKHFLSTGVDIPVIQERCVLLDEYIKTYAFSSSFS----- 707
Query: 610 LVWFLSPEDSHPSSPVSNSPVSLSSFTRG 638
++ L+ + + +S V+ + S++ G
Sbjct: 708 IIETLTVKPVNKTSTVATNIASMTQAAPG 736
>At1g15940 T24D18.4
Length = 1012
Score = 42.4 bits (98), Expect = 0.001
Identities = 52/221 (23%), Positives = 95/221 (42%), Gaps = 11/221 (4%)
Query: 249 KEEKDSLIVNDEVEETKD-IGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLD 307
K+E+DSL + E + D + D + L E + + D + +++A + N + +
Sbjct: 778 KDEEDSLKLGKESDAEPDRMEDHQELPENHNVETKTDGE-EQEAAKEPTAESKTNGEEPN 836
Query: 308 LLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDD 367
PE D ++ ++ + +E K + + EA +A+ + D +N E + + + D
Sbjct: 837 AEPETDGKEHKSLKEPNAEPKSD--GEEQEAAKEPNAELKT-DGENQEAAKELTAERKTD 893
Query: 368 VSTSNVSVHVENVDAKSFKNLKPIVLPSNGGTRKILERSSTS--TNVLEKSHVISKIEDS 425
V+ VE K N++P + G +K +E + T V EK + D+
Sbjct: 894 EEEHKVADEVEQKSQKE-TNVEP---EAEGEEQKSVEEPNAEPKTKVEEKESAKEQTADT 949
Query: 426 ELSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQSV 466
+L E D + EEI E+ S VGN + Q V
Sbjct: 950 KLIEKEDMSKTKGEEIDKETYSSIPETGKVGNEAEEDDQRV 990
>At1g69480 hypothetical protein
Length = 777
Score = 41.6 bits (96), Expect = 0.002
Identities = 43/177 (24%), Positives = 79/177 (44%), Gaps = 16/177 (9%)
Query: 124 NENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVS--SEFYS 181
+E G++ ++ I+ + +EE + ++ F S G+ E F+ EN++ ++FY
Sbjct: 68 SERAGDIEDQVIKVDTVQEEGSRKLYETKFLKKSEEGGEFEESFFKKLDENLNKVNKFYR 127
Query: 182 SRSVSLYEEGEVRNEN-PLLMNSSVAFGSHDLDDFLLQNGPVSVV----SDLFYNPRESN 236
+ + EE + ++ L+ V D+D+ L+ P V SD + +
Sbjct: 128 DKVKEVIEEAALLDKQMDALIALRVKMQKPDVDNLNLEKHPSDKVVVDTSDNTMRTQGTA 187
Query: 237 NRVEEHGVSSVR-KEEKDSLIVNDEV--------EETKDIGDREALEEVRDRDRDRD 284
N HG+ EE+ S I+ D V EE IGD++ L E+ +R + D
Sbjct: 188 NTDMVHGIERTNIPEEEASHIMADIVPVSHTNGDEEEASIGDKQDLREILERVKMND 244
>At2g15900 hypothetical protein
Length = 952
Score = 41.2 bits (95), Expect = 0.003
Identities = 28/86 (32%), Positives = 47/86 (54%), Gaps = 11/86 (12%)
Query: 516 GVKEYTVYKIKVWSGKDQ-WEVEKRYRDFLTLHRCMKSLFNEQGWTLPLPWSSVEKESKI 574
G K + VY I V +++ W V++RY +F LHR +K + N + L LP +I
Sbjct: 541 GSKSFAVYSIAVTDVENKTWFVKRRYSNFERLHRQLKEIPN---YNLQLP------PKRI 591
Query: 575 FRSASLD-IIAKRSVLIQECLQSILC 599
F S++ D + +R + + + LQ +LC
Sbjct: 592 FSSSTEDAFVHRRCIQLDKYLQDLLC 617
>At1g60640 unknown protein
Length = 340
Score = 41.2 bits (95), Expect = 0.003
Identities = 75/286 (26%), Positives = 116/286 (40%), Gaps = 65/286 (22%)
Query: 97 DEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGE----REIETEEEEEEEFSEGDDSM 152
D GD+ S + +++ ++G + SD ++ GG + + R +E EEEEE
Sbjct: 52 DSGDDESPAAGEDADV--DDGDDNSDADDYGGTLEKMSMNRFLEEPPEEEEE-------- 101
Query: 153 FNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDL 212
NY GC ++ ++V Y V L NEN M + V+ H
Sbjct: 102 -NYILGC---------MIQSKSVYKCRYCPTVVCL-------NENT--MQAHVSSKKHAR 142
Query: 213 DDFLLQNGPVSV----VSDLFYNPRES----NNRVEEHGVSSVRKEEKDSLIVNDEVEET 264
+ L++ G + V DL +E N R + G S +K+EKDSL N E EE
Sbjct: 143 MEKLVKEGKIRTDDEEVDDLETASQEKEKKGNRRSQRQGKRS-QKQEKDSLTKNGENEEV 201
Query: 265 KD----------IGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDP 314
+D G+R A E+R R + +++D + EV DNS P +
Sbjct: 202 EDPETPSQEKQIKGNRRARRELR-RSQKQEKDSSTKHGENEEV---DNSG----TPSQGK 253
Query: 315 QKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQ 360
Q N S + R + S EN ++DN E S +
Sbjct: 254 QIKEN-----SRARRQRKRLEKQGKGSLTKHGENEEVDNPETPSQE 294
>At4g12610 putative protein
Length = 649
Score = 40.8 bits (94), Expect = 0.003
Identities = 54/265 (20%), Positives = 104/265 (38%), Gaps = 20/265 (7%)
Query: 75 RTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGERE 134
+T D R + ++G L+G+ +E G G + S G+E G V +R
Sbjct: 183 KTADGYQRWMMKAANNGPALFGEVDNEKESGGTSGGG--GRGRKKSSGGDEEEGNVSDR- 239
Query: 135 IETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVR 194
+E+EEEE S N S D D E +G ++ + +E E+
Sbjct: 240 --GDEDEEEEASRKSRLGLNRKSNDDDDEEGP----RGGDLDMDDDDIEKGDDWEHEEIF 293
Query: 195 NENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDS 254
++ + +V + +D L P ++ + + N EE G+S KE K
Sbjct: 294 TDD----DEAVGNDPEEREDLLAPEIPAP--PEIKQDEDDEENEEEEGGLSKSGKELKKL 347
Query: 255 LIVNDEVEETKDIGDREALEEVRDR----DRDRDRDRDRDAPV-SCEVQCADNSPDLDLL 309
L + ++E+ + D ++ +E + ++ ++ PV + + A + P
Sbjct: 348 LGKANGLDESDEDDDDDSDDEEETNYGTVTNSKQKEAAKEEPVDNAPAKPAPSGPPRGTP 407
Query: 310 PEEDPQKSLNITDGGSEGKGNRYSK 334
P + + + DG S+ + K
Sbjct: 408 PAKPSKGKRKLNDGDSKKPSSSVQK 432
>At3g28770 hypothetical protein
Length = 2081
Score = 40.4 bits (93), Expect = 0.004
Identities = 82/404 (20%), Positives = 160/404 (39%), Gaps = 67/404 (16%)
Query: 64 DFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELI----GNNGIE 119
+ +++ S+ + N +S Q G+E G G ++SM L +++ GNN +E
Sbjct: 200 EHNNVTTGSNMVETNGENSESTQEKGDGVE--GSNGGDVSMENLQGNKVEDLKEGNNVVE 257
Query: 120 ESDGNENGGE----------------VGEREIETEEEEEEEFSEGDDSMFNYGS------ 157
+ EN GE +G+ IE E +E+ ++ N GS
Sbjct: 258 NGETKENNGENVESNNEKEVEGQGESIGDSAIEKNLESKEDVKSEVEAAKNDGSSMTENL 317
Query: 158 ----GCDG----DNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGS 209
G +G DNE E +GE++ S +L + +V++E N+ + +
Sbjct: 318 GEAQGNNGVSTIDNEKEVEG-QGESIED---SDIEKNLESKEDVKSEVEAAKNAGSSM-T 372
Query: 210 HDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSV------RKEEKDSLIVND-EVE 262
L++ NG VS + + S + V++ +KE K+ N+ E
Sbjct: 373 GKLEEAQRNNG-VSTNETMNSENKGSGESTNDKMVNATTNDEDHKKENKEETHENNGESV 431
Query: 263 ETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITD 322
+ +++ ++ EE + ++ + + + V+ N+ E+ Q+S +
Sbjct: 432 KGENLENKAGNEESMKGENLENKVGNEELKGNASVEAKTNNESSKEEKREESQRSNEVYM 491
Query: 323 GGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVST-----------S 371
KG + N + + GD+ ++N L+N E + N D ++ +
Sbjct: 492 NKETTKGE--NVNIQGESIGDSTKDN-SLENKEDVKPKVDANESDGNSTKERHQEAQVNN 548
Query: 372 NVSVHVENVD----AKSFKNLKPIVLPSNGGTRKILERSSTSTN 411
VS +N+D + KN K + + +N G +R T N
Sbjct: 549 GVSTEDKNLDNIGADEQKKNDKSVEVTTNDGDHTKEKREETQGN 592
Score = 38.9 bits (89), Expect = 0.013
Identities = 82/427 (19%), Positives = 159/427 (37%), Gaps = 69/427 (16%)
Query: 71 ASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGG-- 128
A S T D+ S + E G++ E D E +E + NENGG
Sbjct: 796 AKSVETKDNKKLSSTENRDEAKERSGEDNKE------DKEESKDYQSVEAKEKNENGGVD 849
Query: 129 ------------------EVGEREIETEEEEEEEFSEGDDS----MFNYGSGCD-----G 161
EV + E+ +++ EE D S + ++ + D G
Sbjct: 850 TNVGNKEDSKDLKDDRSVEVKANKEESMKKKREEVQRNDKSSTKEVRDFANNMDIDVQKG 909
Query: 162 DNENEFYSM--KGENVSSEFYSSRSVSLYEEG-EVRNENPLLMNSSVAFGSHDLDDFLLQ 218
E+ Y K E E + + S ++G + + + NS++ D +++
Sbjct: 910 SGESVKYKKDEKKEGNKEENKDTINTSSKQKGKDKKKKKKESKNSNMKKKEEDKKEYV-- 967
Query: 219 NGPVSVVSDLFYNPRESNNRV--EEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEV 276
N + D +S N EE+ + +KE +DS N E +E ++ + E
Sbjct: 968 NNELKKQEDNKKETTKSENSKLKEENKDNKEKKESEDSASKNREKKEYEEKKSKTKEEAK 1027
Query: 277 RDRDRDRDRDR-DRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGK------- 328
+++ + +D+ R ++D+ + + S DL +E+ K ++ K
Sbjct: 1028 KEKKKSQDKKREEKDSEERKSKKEKEESRDLKAKKKEEETKEKKESENHKSKKKEDKKEH 1087
Query: 329 ----------GNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVE 378
+ K E S + + D++ E ++ ++D + S HV+
Sbjct: 1088 EDNKSMKKEEDKKEKKKHEESKSRKKEEDKKDMEKLEDQNSN--KKKEDKNEKKKSQHVK 1145
Query: 379 NVDAKSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKSHVIS-----KIEDSELSEFYDE 433
V +S K K K +E S + N ++K S K ++ E+ E ++
Sbjct: 1146 LVKKESDKKEKK--ENEEKSETKEIESSKSQKNEVDKKEKKSSKDQQKKKEKEMKESEEK 1203
Query: 434 VVQEMEE 440
+++ EE
Sbjct: 1204 KLKKNEE 1210
Score = 38.1 bits (87), Expect = 0.022
Identities = 81/402 (20%), Positives = 161/402 (39%), Gaps = 47/402 (11%)
Query: 103 SMTGLDSSELIGNNGIE-----ESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGS 157
SMTG E NNG+ S+ +G ++ + +E+ E +
Sbjct: 370 SMTG-KLEEAQRNNGVSTNETMNSENKGSGESTNDKMVNATTNDEDHKKENKEET----- 423
Query: 158 GCDGDNENEFYSMKGENVSSEFYSSRSVSLYE-EGEVRNENPLLMNSSVAFGSHDLDDFL 216
+EN S+KGEN+ ++ + S+ E +V NE L N+SV +++
Sbjct: 424 -----HENNGESVKGENLENKAGNEESMKGENLENKVGNEE-LKGNASVEAKTNNESSKE 477
Query: 217 LQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEV 276
+ ++++ N + S+ KD+ + N E D + +
Sbjct: 478 EKREESQRSNEVYMNKETTKGENVNIQGESIGDSTKDNSLENKE--------DVKPKVDA 529
Query: 277 RDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKND 336
+ D + ++R ++A V+ V D + D ++ +E + ++ ++G + + +
Sbjct: 530 NESDGNSTKERHQEAQVNNGVSTEDKNLD-NIGADEQKKNDKSVEVTTNDGDHTKEKREE 588
Query: 337 EAGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSN---VSVHVENVDAKSFKNLKPIVL 393
G +G++ + N +L+N E K + D T+N + E ++ ++
Sbjct: 589 TQGNNGESVK-NENLENKEDKKELKDDESVGAKTNNETSLEEKREQTQKGHDNSINSKIV 647
Query: 394 PSNGGT------RKILERSSTSTNVLE-----KSHVISKIED--SELSEFYDEVVQE-ME 439
+ GG +++ ST+ N +E KS V K D SE E E ++ ME
Sbjct: 648 DNKGGNADSNKEKEVHVGDSTNDNNMESKEDTKSEVEVKKNDGSSEKGEEGKENNKDSME 707
Query: 440 EILLESMDSPAARFSVGNRMFDPQQSVPSRDGGLTASTSSTD 481
+ LE+ +S S ++ D +Q GG + S +
Sbjct: 708 DKKLENKESQTD--SKDDKSVDDKQEEAQIYGGESKDDKSVE 747
Score = 36.2 bits (82), Expect = 0.083
Identities = 68/370 (18%), Positives = 138/370 (36%), Gaps = 55/370 (14%)
Query: 97 DEGDELSMTGLDSS---ELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMF 153
+E E + G D+S +++ N G E VG+ + E +E+ +
Sbjct: 628 EEKREQTQKGHDNSINSKIVDNKGGNADSNKEKEVHVGDSTNDNNMESKEDTKSEVEVKK 687
Query: 154 NYGSGCDGD-----NENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFG 208
N GS G+ N++ K EN S+ S S+ ++ E + +G
Sbjct: 688 NDGSSEKGEEGKENNKDSMEDKKLENKESQTDSKDDKSVDDKQE----------EAQIYG 737
Query: 209 SHDLDDFLL----------QNGPVSVVSDLFYNPRES---NNRVEEHGVSSVRKEEKDSL 255
DD + +N + N E+ N + E +KE KD+
Sbjct: 738 GESKDDKSVEAKGKKKESKENKKTKTNENRVRNKEENVQGNKKESEKVEKGEKKESKDAK 797
Query: 256 IVNDEVEETKDIGDREALEEVRDR---DRDRDRDRDRD-APVSCEVQCADNSPDLDLLPE 311
V E ++ K + E +E ++R D D++ +D V + + + D ++ +
Sbjct: 798 SV--ETKDNKKLSSTENRDEAKERSGEDNKEDKEESKDYQSVEAKEKNENGGVDTNVGNK 855
Query: 312 EDPQ-----KSLNITDGGSEG---KGNRYSKNDEAGASGDAQ-RENLDLDNFEFKSDQFC 362
ED + +S+ + E K +ND++ N+D+D + +
Sbjct: 856 EDSKDLKDDRSVEVKANKEESMKKKREEVQRNDKSSTKEVRDFANNMDIDVQKGSGESVK 915
Query: 363 DNRDDVSTSNVSVHVENVDAKSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKSHVISKI 422
+D+ N + + ++ S G +K ++ S ++N+ +K +
Sbjct: 916 YKKDEKKEGNKEENKDTINTSS---------KQKGKDKKKKKKESKNSNMKKKEEDKKEY 966
Query: 423 EDSELSEFYD 432
++EL + D
Sbjct: 967 VNNELKKQED 976
Score = 32.0 bits (71), Expect = 1.6
Identities = 64/280 (22%), Positives = 118/280 (41%), Gaps = 57/280 (20%)
Query: 89 SSGLELYGDEGDELSMTGLDSSELIGNNGIEES--DGNENGGEV---GEREIETEEEEE- 142
+ G E+ +EG + DS+ + N G E+S +G+E+G V G E+ TEE +
Sbjct: 1623 NGGKEVSTEEGSK------DSNIVERNGGKEDSIKEGSEDGKTVEINGGEELSTEEGSKD 1676
Query: 143 ---EEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPL 199
EE EG ++ GS D E M+G+ S++ S ++G++ NE
Sbjct: 1677 GKIEEGKEGKENSTKEGSKDDKIEE----GMEGKENSTKESS-------KDGKI-NEIHG 1724
Query: 200 LMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGV--SSVRKEEKDSLIV 257
+++ GS D G S D + + VE +GV S++ + K+ I
Sbjct: 1725 DKEATMEEGSKD-------GGTNSTGKD-----SKDSKSVEINGVKDDSLKDDSKNGDI- 1771
Query: 258 NDEVEETKDIGDREALEEVRDRDRD-------------RDRDRDRDAPVSCEVQCADNSP 304
+E+ K+ ++ + E++ D D ++D + E Q N
Sbjct: 1772 -NEINNGKEDSVKDNVTEIQGNDNSLTNSTSSEPNGDKLDTNKDSMKNNTMEAQGGSNGD 1830
Query: 305 DLDLLPEEDPQKSLNITDGGSEGKG-NRYSKNDEAGASGD 343
+ EE + ++++ + + G N S N++ +GD
Sbjct: 1831 STNGETEETKESNVSMNNQNMQDVGSNENSMNNQTTGTGD 1870
>At2g27470 putative CCAAT-binding transcription factor subunit
Length = 275
Score = 40.4 bits (93), Expect = 0.004
Identities = 32/91 (35%), Positives = 41/91 (44%), Gaps = 15/91 (16%)
Query: 97 DEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEE-------EEEEFSEGD 149
DE D+ +++E GN+ + + E G E E E EE E EE D
Sbjct: 187 DENDD------ENTEENGNDEENDDENTEENGNDEENEKEDEENSMEENGNESEESGNED 240
Query: 150 DSMFNYGSGCDGDNENEFYSM--KGENVSSE 178
SM GSG DNENE S+ GE V S+
Sbjct: 241 HSMEENGSGVGEDNENEDGSVSGSGEEVESD 271
Score = 33.1 bits (74), Expect = 0.70
Identities = 27/104 (25%), Positives = 46/104 (43%), Gaps = 5/104 (4%)
Query: 115 NNGIEESDGN----ENGG-EVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYS 169
N+ EE +GN ENG E E + E EE + D++ G+ + + E+E S
Sbjct: 166 NDNTEEENGNDEEDENGNDEEDENDDENTEENGNDEENDDENTEENGNDEENEKEDEENS 225
Query: 170 MKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLD 213
M+ SE + S+ E G E+ + SV+ +++
Sbjct: 226 MEENGNESEESGNEDHSMEENGSGVGEDNENEDGSVSGSGEEVE 269
>At2g15200 putative protein on transposon FARE2.7 (cds2)
Length = 650
Score = 40.4 bits (93), Expect = 0.004
Identities = 51/210 (24%), Positives = 87/210 (41%), Gaps = 23/210 (10%)
Query: 107 LDSSELIGNNGIEESDGNENGGEVGEREIETEEE-EEEEFSEGDDSMFNYGSGCDGDNEN 165
L+ E G+ G E+ + + G E E+E EEE +EEE E ++ Y G +G +
Sbjct: 444 LEKVEYRGDEGKEKQEIPKQGDE----EMEGEEEKQEEEGKEEEEEKVEY-RGDEGKEKQ 498
Query: 166 EFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVV 225
E E + E EEG+ E +L +V HD D ++
Sbjct: 499 EIPKQGDEEMEGEEEKQE-----EEGKEEEEEKVLKEENVE--EHDEHDETEDQEAYVIL 551
Query: 226 SDLFYN---PRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDR--- 279
SD N P E ++ ++ + V KEE ++ +D+ +ET+D L + D
Sbjct: 552 SDDEDNGTTPTEKESQPQKEETTEVPKEE--NVEEHDKHDETEDQEAYVILSDDEDNGTA 609
Query: 280 --DRDRDRDRDRDAPVSCEVQCADNSPDLD 307
+++ R ++ V E + D + D
Sbjct: 610 PTEKESQRQKEETTEVPRETKKDDEDVEKD 639
Score = 31.2 bits (69), Expect = 2.7
Identities = 35/174 (20%), Positives = 65/174 (37%), Gaps = 7/174 (4%)
Query: 119 EESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSE 178
EE + E E E E + EE E+ E+ GD+ GD E E K E E
Sbjct: 423 EEDEKKEQEEEKQEEEGKEEELEKVEY-RGDEGKEKQEIPKQGDEEMEGEEEKQEEEGKE 481
Query: 179 FYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNR 238
+ +EG+ + E P + + ++ + V+ + + ++
Sbjct: 482 EEEEKVEYRGDEGKEKQEIPKQGDEEMEGEEEKQEEEGKEEEEEKVLKEENVEEHDEHDE 541
Query: 239 VEEHGVSSVRKEEKDSLIVNDE------VEETKDIGDREALEEVRDRDRDRDRD 286
E+ + +++D+ E EET ++ E +EE D D++
Sbjct: 542 TEDQEAYVILSDDEDNGTTPTEKESQPQKEETTEVPKEENVEEHDKHDETEDQE 595
>At3g26560 ATP-dependent RNA helicase, putative
Length = 1168
Score = 40.0 bits (92), Expect = 0.006
Identities = 40/181 (22%), Positives = 74/181 (40%), Gaps = 17/181 (9%)
Query: 163 NENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPV 222
NE E + E V +EF + L E +E + ++ ++ D+ +++
Sbjct: 20 NELETHLGSAEKVLAEFI----IDLGRHSETVDE----FDKNLKEAGAEMPDYFVRSLLT 71
Query: 223 SVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRD 282
++ Y P+ + + +E G K L + D ++ K++ ++E E +R R+
Sbjct: 72 TIHG--IYPPKPKSEKKKEEGDDQKFK----GLAIKDTKDKVKEL-EKEIEREAEERRRE 124
Query: 283 RDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASG 342
DR+RDRD S + D + D D D + G EG+ R + G
Sbjct: 125 EDRNRDRDRRESGRDRDRDRNRDRD--DRRDRHRDRERNRGDEEGEDRRSDRRHRERGRG 182
Query: 343 D 343
D
Sbjct: 183 D 183
>At4g26630 unknown protein
Length = 763
Score = 39.7 bits (91), Expect = 0.007
Identities = 38/147 (25%), Positives = 64/147 (42%), Gaps = 11/147 (7%)
Query: 245 SSVRKEEKDSLI--VNDEVEETKDIGDREALEEVRDRDRDRDRDRD----RDAPVSCEVQ 298
S V+K E ++ + ++VE TKD G EA D D +++ D D + V+
Sbjct: 72 SEVKKNEDNAETQKMEEKVEVTKDEGQAEATNMDEDADGKKEQTDDGVSVEDTVMKENVE 131
Query: 299 CADNSPDLDLLPEEDPQKSLNIT--DGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEF 356
DN+ D +E K +IT D GK + + D+A + D ++ +
Sbjct: 132 SKDNNYAKD---DEKETKETDITEADHKKAGKEDIQHEADKANGTKDGNTGDIKEEGTLV 188
Query: 357 KSDQFCDNRDDVSTSNVSVHVENVDAK 383
D+ D + V + + VENV+ K
Sbjct: 189 DEDKGTDMDEKVENGDENKQVENVEGK 215
>At2g33730 putative U5 small nuclear ribonucleoprotein, an RNA
helicase
Length = 733
Score = 39.3 bits (90), Expect = 0.010
Identities = 31/115 (26%), Positives = 45/115 (38%), Gaps = 9/115 (7%)
Query: 235 SNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVS 294
S RV + E +D + N + KD + RDRDR+RDR RDRD
Sbjct: 46 SEQRVRREQLGRPEPETED--VSNGDTNRDKDRDRDRDRDRERDRDRERDRGRDRD---- 99
Query: 295 CEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENL 349
+ D D D E D ++ D + + + +E A A+ E L
Sbjct: 100 ---RDRDRDRDRDRERERDRERDRRERDREPDRRNREKEREEEVKAREKARVEKL 151
>At1g67120 hypothetical protein
Length = 5138
Score = 39.3 bits (90), Expect = 0.010
Identities = 84/459 (18%), Positives = 164/459 (35%), Gaps = 76/459 (16%)
Query: 39 CSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDE 98
C + + S+ +V D F S DD ++ + G +
Sbjct: 4258 CYLKEFLAISKTVSLITHVLASVLADLFTKGFGISKNEEDDDSKVDKSEAAEGTGMGDGV 4317
Query: 99 GDELSMTGLDSSELIGNN-GIEESDGNENGGEVGEREIETEEEEEE-------------- 143
G + + +++G N GIE SD +G E E E E++E+E
Sbjct: 4318 GAKDEEEEKEQDDVLGKNKGIEMSD-EFDGKEYSVSEDEEEDKEDEGSEDEPLDNGIGDV 4376
Query: 144 --EFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLM 201
+ + D+ +N + +N NE + G ++ + SR + ++G + P
Sbjct: 4377 GSDAEKADEKPWNKDEEDEEENMNE-KNESGPSIVDKDTRSRELRAKDDGVETADEPEES 4435
Query: 202 NSS--------VAFGSHDLDDFLLQNGPVSVVSDLF--YNPRESNNRVEEHGVSSVRKEE 251
N+S D DD + + P N ++++ + + EE
Sbjct: 4436 NTSDKPEEGNDENVEQDDFDDTDNLEEKIQTKEEALGGLTPDVDNEQIDD-DMEMDKTEE 4494
Query: 252 KDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCA-----DNSPDL 306
+ N + E + D++ EE + + + + E +C + DL
Sbjct: 4495 VEKEDANQQEEPCSE--DQKHPEEGENDQEETQEPSEENMEAEAEDRCGSPQKEEPGNDL 4552
Query: 307 DLLPEEDPQKSLNI--------------TDGGSEGKGNRYSKNDEAGASGDAQRENLDLD 352
+ PE +P + + G G N + N GA A +ENL
Sbjct: 4553 EQEPETEPIEGKEVMSEDMMKPNFRNDNISGVESGSQNPHGSN-VLGAGSTAPQENLSAT 4611
Query: 353 NFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSFK----NLKPIVLPSNGGT--------- 399
+ +D+ D+ D S+SN +++ + + + NL + P N +
Sbjct: 4612 DV---TDELTDSMDLPSSSNTEMNLMMTNMANGETLTDNLPKMEFPQNQSSTAQQTKVNP 4668
Query: 400 --------RKILERSSTSTNVLEKSHVISKIEDSELSEF 430
++ ER S+++ EK +++ED + SE+
Sbjct: 4669 YRNVGDALKEWKERVRISSDLGEKQEAENEMEDPDASEY 4707
>At4g37820 unknown protein
Length = 532
Score = 38.9 bits (89), Expect = 0.013
Identities = 85/400 (21%), Positives = 145/400 (36%), Gaps = 54/400 (13%)
Query: 72 SSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVG 131
+S DD + G L +E ++ D G + S ENG E+
Sbjct: 49 TSKIVVDDIDNTVVNLGRKDLRPRIEETKDVKDEVEDEEGSKNEGGGDVSTDKENGDEIV 108
Query: 132 EREIETE--EEEEEEFSEGDDSMFNYGSGCDGDNE---------NEF-------YSMKGE 173
ERE E + EE E+ +EG + +G++E NE + KG+
Sbjct: 109 EREEEEKAVEENNEKEAEGTGNEEGNEDSNNGESEKVVDESEGGNEISNEEAREINYKGD 168
Query: 174 NVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPV--SVVSDLFYN 231
+ SSE E+ EV E+ NS+ H+ D+ +N + SV+ ++ N
Sbjct: 169 DASSEVMHGTEEKSNEKVEVEGESK--SNSTENVSVHE-DESGPKNEVLEGSVIKEVSLN 225
Query: 232 PRESNNRVEEHGVSSVRKEEKDSLI-------VNDEVEETKDIGDREALEEVRDRDRDRD 284
E+ + + G K E DS N E+ ET ++ A E D
Sbjct: 226 TTENGS---DDGEQQETKSELDSKTGEKGFSDSNGELPET-NLSTSNATETTESSGSDES 281
Query: 285 RDRDRDAPVSCEVQCADNSPDLDLLPEEDPQK--------SLNITDGGSEGKGNRYSKND 336
+ D + EE K + + D E K R K +
Sbjct: 282 GSSGKSTGYQQTKNEEDEKEKVQSSEEESKVKESGKNEKDASSSQDESKEEKPER-KKKE 340
Query: 337 EAGASGDA------QRENLDLDNFEFKSDQFCDNRD-DVSTSNVSVHVENVDAKSFKNLK 389
E+ + G+ +RE D + E ++ +N++ + S+S ++ + K K
Sbjct: 341 ESSSQGEGKEEEPEKREKEDSSSQEESKEEEPENKEKEASSSQEENEIKETEIKE----K 396
Query: 390 PIVLPSNGGTRKILERSSTSTNVLEKSHVISKIEDSELSE 429
G K E+ S+ + E ++ KIE E ++
Sbjct: 397 EESSSQEGNENKETEKKSSESQRKENTNSEKKIEQVESTD 436
Score = 33.9 bits (76), Expect = 0.41
Identities = 54/277 (19%), Positives = 100/277 (35%), Gaps = 29/277 (10%)
Query: 78 DDSNRKSFQYGSSGLELYGDEGDELSMTGLDSS---ELIGNNGIEESDGNENGGEVGERE 134
+ KS +G + + D EL T L +S E ++G +ES +G G ++
Sbjct: 235 EQQETKSELDSKTGEKGFSDSNGELPETNLSTSNATETTESSGSDES--GSSGKSTGYQQ 292
Query: 135 IETEEEEEEEFSEGDDSMFNYGSGCD----GDNENEFYSMKGENVSSEFYSSRSVSLYEE 190
+ EE+E+E+ ++ SG + +++E K E E SS+ EE
Sbjct: 293 TKNEEDEKEKVQSSEEESKVKESGKNEKDASSSQDESKEEKPERKKKEESSSQGEGKEEE 352
Query: 191 GEVRNENPLLMN---------SSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEE 241
E R + + S ++ ++ + + N E+
Sbjct: 353 PEKREKEDSSSQEESKEEEPENKEKEASSSQEENEIKETEIKEKEESSSQEGNENKETEK 412
Query: 242 HGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCAD 301
S RKE +S ++VE T ++ E+ D + R+ D ++
Sbjct: 413 KSSESQRKENTNSEKKIEQVESTDSSNTQKGDEQKTDESK---RESGNDT--------SN 461
Query: 302 NSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEA 338
+ D E +K N +G +E N + A
Sbjct: 462 KETEDDSSKTESEKKEENNRNGETEETQNEQEQTKSA 498
>At2g40650 unknown protein
Length = 355
Score = 38.9 bits (89), Expect = 0.013
Identities = 28/79 (35%), Positives = 37/79 (46%), Gaps = 7/79 (8%)
Query: 216 LLQNGPV----SVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDRE 271
L QNG + SV+ D F E + E G++ ++E D + E E +D R
Sbjct: 171 LEQNGLLEPRKSVLEDDF---EEEEEKEENEGIADGSEDEMDQRRKSPERERERDRDRRR 227
Query: 272 ALEEVRDRDRDRDRDRDRD 290
RDRD DRD D DRD
Sbjct: 228 DSHRHRDRDYDRDYDMDRD 246
Score = 31.6 bits (70), Expect = 2.0
Identities = 20/59 (33%), Positives = 35/59 (58%), Gaps = 6/59 (10%)
Query: 239 VEEHGVSSVRKEE-KDSLIVNDEVEETKDIGD--REALEEVR---DRDRDRDRDRDRDA 291
+E++G+ RK +D +E EE + I D + +++ R +R+R+RDRDR RD+
Sbjct: 171 LEQNGLLEPRKSVLEDDFEEEEEKEENEGIADGSEDEMDQRRKSPERERERDRDRRRDS 229
Score = 30.4 bits (67), Expect = 4.5
Identities = 20/54 (37%), Positives = 27/54 (49%), Gaps = 7/54 (12%)
Query: 269 DREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDL------DLLPEEDPQK 316
DR+ E RDRDR+R RDR+RD + S D D+ E+P+K
Sbjct: 266 DRDHYRE-RDRDRERGRDRERDRRDRARRRSRSRSRDRKRHETDDVRDREEPKK 318
>At2g11910 unknown protein
Length = 168
Score = 38.9 bits (89), Expect = 0.013
Identities = 25/89 (28%), Positives = 40/89 (44%), Gaps = 7/89 (7%)
Query: 64 DFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDG 123
D D D DD+N + F G GDEG+E D G G ++ D
Sbjct: 78 DDDDEDADEDDDDEDDANDEDFSGGE------GDEGEE-EADPEDDPVTNGGGGSDDEDD 130
Query: 124 NENGGEVGEREIETEEEEEEEFSEGDDSM 152
++ G+ + + + E+EEE++ E DD +
Sbjct: 131 DDEEGDNDDEDEDNEDEEEDDDEEDDDDV 159
Score = 35.8 bits (81), Expect = 0.11
Identities = 28/112 (25%), Positives = 48/112 (42%), Gaps = 9/112 (8%)
Query: 248 RKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLD 307
R K+ +EE KD D + ++ D D D D + D + + + + D
Sbjct: 53 RLSVKEGYKRGSRIEENKDASDSDDDDDDEDADEDDDDEDDANDEDFSGGEGDEGEEEAD 112
Query: 308 LLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSD 359
PE+DP +T+GG G + +DE G + D +N D + + + D
Sbjct: 113 --PEDDP-----VTNGG--GGSDDEDDDDEEGDNDDEDEDNEDEEEDDDEED 155
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.133 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,678,289
Number of Sequences: 26719
Number of extensions: 1050851
Number of successful extensions: 5706
Number of sequences better than 10.0: 228
Number of HSP's better than 10.0 without gapping: 66
Number of HSP's successfully gapped in prelim test: 167
Number of HSP's that attempted gapping in prelim test: 4292
Number of HSP's gapped (non-prelim): 674
length of query: 880
length of database: 11,318,596
effective HSP length: 108
effective length of query: 772
effective length of database: 8,432,944
effective search space: 6510232768
effective search space used: 6510232768
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 65 (29.6 bits)
Medicago: description of AC124952.2


