

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124952.12 + phase: 2 /pseudo
(313 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
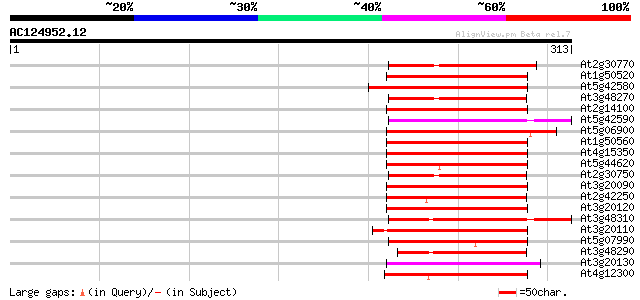
Sequences producing significant alignments: (bits) Value
At2g30770 putative cytochrome P450 74 1e-13
At1g50520 cytochrome P450 like protein 71 7e-13
At5g42580 cytochrome P450 70 1e-12
At3g48270 cytochrome P450 - like protein 70 1e-12
At2g14100 putative cytochrome P450 70 1e-12
At5g42590 cytochrome P450 69 3e-12
At5g06900 cytochrome P450 69 3e-12
At1g50560 cytochrome P450, putative 68 6e-12
At4g15350 cytochrome P450 like protein 66 3e-11
At5g44620 flavonoid 3',5'-hydroxylase-like; cytochrome P450 65 5e-11
At2g30750 cytochrome P450 like protein 65 6e-11
At3g20090 cytochrome P450 like protein 64 8e-11
At2g42250 putative cytochrome P450 64 8e-11
At3g20120 cytochrome P450 - like protein 64 1e-10
At3g48310 cytochrome P450 - like protein 63 2e-10
At3g20110 cytochrome P450, putative 63 2e-10
At5g07990 flavonoid 3'-hydroxylase - like protein 62 4e-10
At3g48290 cytochrome P450 - like protein 62 4e-10
At3g20130 cytochrome P450, putative 62 4e-10
At4g12300 flavonoid 3',5'-hydroxylase -like protein 61 7e-10
>At2g30770 putative cytochrome P450
Length = 497
Score = 73.9 bits (180), Expect = 1e-13
Identities = 35/83 (42%), Positives = 55/83 (66%), Gaps = 2/83 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
KIK+ S D +D+V+ EH + D K DF+DILL ++ D + F++ +ND+K ++++
Sbjct: 237 KIKEVSRGFSDLMDKVVQEHLEASND--KADFVDILLSIEKDKNSGFQVQRNDIKFMILD 294
Query: 272 MFLGGSDTTSTTLEWAMAACKKS 294
MF+GG+ TTST LEW M +S
Sbjct: 295 MFIGGTSTTSTLLEWTMTELIRS 317
>At1g50520 cytochrome P450 like protein
Length = 533
Score = 71.2 bits (173), Expect = 7e-13
Identities = 29/79 (36%), Positives = 55/79 (68%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
K+I + S+ D+ L+++I EH+ + +D +D+LL++ D E ++T+N +KAL++
Sbjct: 250 KEIMEVSQRYDELLEKIIKEHEEDPNKKEDRDMMDVLLEVCADDKAEVKITRNQIKALIV 309
Query: 271 NMFLGGSDTTSTTLEWAMA 289
+FLGG+DT++ T++W MA
Sbjct: 310 ELFLGGTDTSAQTIQWIMA 328
>At5g42580 cytochrome P450
Length = 499
Score = 70.5 bits (171), Expect = 1e-12
Identities = 32/89 (35%), Positives = 58/89 (64%)
Query: 201 VGLMFSLAKLKKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFEL 260
V L FS K I S E D+FL+R++VEH+ + + +D +D LL+ + E+++
Sbjct: 222 VKLFFSNMFRKDIMGVSREFDEFLERILVEHEENLEGDQDRDMIDHLLEAYRNEEAEYKI 281
Query: 261 TQNDLKALLMNMFLGGSDTTSTTLEWAMA 289
T+ +K+L++ +FLGG+D+++ T++W MA
Sbjct: 282 TRKQIKSLIVEIFLGGTDSSAQTIQWTMA 310
>At3g48270 cytochrome P450 - like protein
Length = 489
Score = 70.5 bits (171), Expect = 1e-12
Identities = 31/77 (40%), Positives = 54/77 (69%), Gaps = 2/77 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
+++ ++ ++D F +RV+ +H RD DF+D+LL +Q D FE+ + +KA++MN
Sbjct: 230 QLEKTANDVDKFFERVVQDHVDGNRD--MTDFVDVLLAIQRDKTVGFEINRVSIKAIVMN 287
Query: 272 MFLGGSDTTSTTLEWAM 288
+F+GG+DT+ST +EWAM
Sbjct: 288 VFVGGTDTSSTLMEWAM 304
>At2g14100 putative cytochrome P450
Length = 518
Score = 70.1 bits (170), Expect = 1e-12
Identities = 31/79 (39%), Positives = 52/79 (65%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
K+ D S + D+ L+R+IVEH+ D +D+LL + DG E+++T++ LK+L +
Sbjct: 248 KETLDVSRKFDELLERIIVEHEEKTDYDHGMDLMDVLLAVYRDGKAEYKITRDHLKSLFV 307
Query: 271 NMFLGGSDTTSTTLEWAMA 289
+ LGG+DT++ T+EW MA
Sbjct: 308 ELILGGTDTSAQTIEWTMA 326
>At5g42590 cytochrome P450
Length = 497
Score = 69.3 bits (168), Expect = 3e-12
Identities = 33/102 (32%), Positives = 61/102 (59%), Gaps = 3/102 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
K++ +++ DF+++V+ EH+ + D + DF+D+LL +Q D + +L ++DLK ++
Sbjct: 236 KLEKITKQFGDFIEKVLQEHEDTTADKETPDFVDMLLTIQRDETAQCQLDKSDLKVIIFE 295
Query: 272 MFLGGSDTTSTTLEWAMAACKKSSHNEESTGGGKENCRGINK 313
MFLG + TTS +EWAM + N E ++ R ++K
Sbjct: 296 MFLGSTTTTSAVIEWAMT---RLMRNPECLKKLQDEIRSVSK 334
>At5g06900 cytochrome P450
Length = 507
Score = 68.9 bits (167), Expect = 3e-12
Identities = 33/103 (32%), Positives = 66/103 (64%), Gaps = 8/103 (7%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPK-KKDFLDILLQLQDDGLTEFELTQNDLKALL 269
K++K++ ++ D ++R++ EH+ S+++ +++ LD+LL + +D E +LT+ ++KA +
Sbjct: 239 KRLKNARDKYDVIIERIMEEHESSKKNATGERNMLDVLLDIYEDKNAEMKLTRENIKAFI 298
Query: 270 MNMFLGGSDTTSTTLEWAMA-------ACKKSSHNEESTGGGK 305
MN++ GG+DT++ T+EWA+A KK+ E G K
Sbjct: 299 MNIYGGGTDTSAITVEWALAELINHPEIMKKAQQEIEQVVGNK 341
>At1g50560 cytochrome P450, putative
Length = 519
Score = 68.2 bits (165), Expect = 6e-12
Identities = 27/79 (34%), Positives = 56/79 (70%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
K+I + S+ D+ L+++I EH+ + + + +D +D+LL++ D EF++++N +KAL +
Sbjct: 251 KEIMEVSQRYDELLEKIIKEHEENPNNGEDRDMMDVLLEVCADDNAEFKISRNQIKALFV 310
Query: 271 NMFLGGSDTTSTTLEWAMA 289
+FL G+DT++ T++W +A
Sbjct: 311 EIFLAGTDTSAQTIQWILA 329
>At4g15350 cytochrome P450 like protein
Length = 509
Score = 65.9 bits (159), Expect = 3e-11
Identities = 27/80 (33%), Positives = 55/80 (68%), Gaps = 1/80 (1%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKK-KDFLDILLQLQDDGLTEFELTQNDLKALL 269
K+I S + D+ L++++VEH+ + + D +D+LL+ D E+++T+N +K++
Sbjct: 240 KEIMSVSRKFDELLEKILVEHEEKMEEHHQGTDMMDVLLEAYRDENAEYKITRNHIKSMF 299
Query: 270 MNMFLGGSDTTSTTLEWAMA 289
+++F+ G+DT+STT++W MA
Sbjct: 300 VDLFIAGTDTSSTTIQWIMA 319
>At5g44620 flavonoid 3',5'-hydroxylase-like; cytochrome P450
Length = 519
Score = 65.1 bits (157), Expect = 5e-11
Identities = 33/82 (40%), Positives = 55/82 (66%), Gaps = 3/82 (3%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDP---KKKDFLDILLQLQDDGLTEFELTQNDLKA 267
K+++ ++ MD DR+I + RD + DFLD+LL+++D+ + +LT ND+KA
Sbjct: 251 KRMRRPAQRMDQMFDRIINQRLGMDRDSSDGRAVDFLDVLLKVKDEEAEKTKLTMNDVKA 310
Query: 268 LLMNMFLGGSDTTSTTLEWAMA 289
+LM+M LGG+DT+ +E+AMA
Sbjct: 311 VLMDMVLGGTDTSLHVIEFAMA 332
>At2g30750 cytochrome P450 like protein
Length = 503
Score = 64.7 bits (156), Expect = 6e-11
Identities = 29/77 (37%), Positives = 52/77 (66%), Gaps = 2/77 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
+IK+ S+ D +D+V+ EH + K+DF+DILL ++ + F+ ++D+K ++++
Sbjct: 243 RIKEVSQGFSDLMDKVVQEHLEAGNH--KEDFVDILLSIESEKSIGFQAQRDDIKFMILD 300
Query: 272 MFLGGSDTTSTTLEWAM 288
MF+GG+ T+ST LEW M
Sbjct: 301 MFIGGTSTSSTLLEWIM 317
>At3g20090 cytochrome P450 like protein
Length = 386
Score = 64.3 bits (155), Expect = 8e-11
Identities = 29/80 (36%), Positives = 52/80 (64%), Gaps = 1/80 (1%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMS-RRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALL 269
K+I S+ D+ L+R+IVEHK ++ + D +D+LL D E+++T+N +K+
Sbjct: 110 KEIMGVSDRFDELLERIIVEHKDKLEKEHQVMDMMDVLLAAYRDKNAEYKITRNHIKSFF 169
Query: 270 MNMFLGGSDTTSTTLEWAMA 289
+++F+GG+DT+ T +W MA
Sbjct: 170 VDLFVGGTDTSVQTTQWTMA 189
>At2g42250 putative cytochrome P450
Length = 514
Score = 64.3 bits (155), Expect = 8e-11
Identities = 29/80 (36%), Positives = 53/80 (66%), Gaps = 2/80 (2%)
Query: 211 KKIKDSSEEMDDFLDRVIVEH--KMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKAL 268
KK+ E+ D ++R++ E K ++D +KD LDILL+ D E ++T+ND+K+
Sbjct: 244 KKLVAVMEKYDLLVERIMKEREAKAKKKDGTRKDILDILLETYRDPTAEMKITRNDMKSF 303
Query: 269 LMNMFLGGSDTTSTTLEWAM 288
L+++F+ G+DT++ ++WAM
Sbjct: 304 LLDVFMAGTDTSAAAMQWAM 323
>At3g20120 cytochrome P450 - like protein
Length = 378
Score = 63.9 bits (154), Expect = 1e-10
Identities = 27/79 (34%), Positives = 50/79 (63%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
K+I S+ D+ L+RV+VEH+ D + +D +D+LL D E ++++N +KA +
Sbjct: 110 KEIMCVSDSFDEVLERVLVEHEQKLDDHQDRDMMDVLLAAYGDENAEHKISRNHIKAFFV 169
Query: 271 NMFLGGSDTTSTTLEWAMA 289
+F G+DT++ +++W MA
Sbjct: 170 ELFFAGTDTSAQSIQWTMA 188
>At3g48310 cytochrome P450 - like protein
Length = 490
Score = 63.2 bits (152), Expect = 2e-10
Identities = 31/102 (30%), Positives = 63/102 (61%), Gaps = 5/102 (4%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
++K + ++D+FL++V+ +H+ D ++ DF+D+LL++Q + FE+ + +KA++++
Sbjct: 231 QLKKTGNDLDEFLEKVVQDHEDG--DAQRTDFVDVLLRIQREKSVGFEIDRLSIKAIILD 288
Query: 272 MFLGGSDTTSTTLEWAMAACKKSSHNEESTGGGKENCRGINK 313
+ +GG+DT+ +EWAM + H E +E R I K
Sbjct: 289 VVVGGTDTSYALMEWAMT---ELLHRPECLNRLQEEVRTICK 327
>At3g20110 cytochrome P450, putative
Length = 510
Score = 63.2 bits (152), Expect = 2e-10
Identities = 31/87 (35%), Positives = 54/87 (61%), Gaps = 1/87 (1%)
Query: 203 LMFSLAKLKKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQ 262
L SL K K+I S D+ L+R+IVE + + + +D+LL+ +D E ++T+
Sbjct: 237 LRISLFK-KEIMYVSNSFDELLERIIVEREKKPNEHQGTYLMDVLLEAYEDEKAEHKITR 295
Query: 263 NDLKALLMNMFLGGSDTTSTTLEWAMA 289
N +K+L + + LGG+DT++ T++W MA
Sbjct: 296 NHIKSLFVELLLGGTDTSAQTIQWTMA 322
>At5g07990 flavonoid 3'-hydroxylase - like protein
Length = 513
Score = 62.0 bits (149), Expect = 4e-10
Identities = 31/80 (38%), Positives = 50/80 (61%), Gaps = 2/80 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEF--ELTQNDLKALL 269
K+K + D FL ++ EH+M+ +D K D L L+ L+ L LT ++KALL
Sbjct: 237 KMKRLHKRFDAFLSSILKEHEMNGQDQKHTDMLSTLISLKGTDLDGDGGSLTDTEIKALL 296
Query: 270 MNMFLGGSDTTSTTLEWAMA 289
+NMF G+DT+++T++WA+A
Sbjct: 297 LNMFTAGTDTSASTVDWAIA 316
>At3g48290 cytochrome P450 - like protein
Length = 488
Score = 62.0 bits (149), Expect = 4e-10
Identities = 29/72 (40%), Positives = 48/72 (66%), Gaps = 2/72 (2%)
Query: 217 SEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMNMFLGG 276
S ++D+FL+RV+ +H D K DF+D LL ++ + FE+ + +KA+++++F+G
Sbjct: 237 SNDLDEFLERVVQDHVDG--DGHKNDFVDFLLTIEREKSVGFEIDRLSIKAIILDVFVGD 294
Query: 277 SDTTSTTLEWAM 288
DTT T LEWAM
Sbjct: 295 MDTTYTLLEWAM 306
>At3g20130 cytochrome P450, putative
Length = 515
Score = 62.0 bits (149), Expect = 4e-10
Identities = 28/86 (32%), Positives = 50/86 (57%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
K I S + D L++V+VEH+ + LD+LL D E+++T+N +KA +
Sbjct: 247 KDIMGVSNKFDVLLEKVLVEHREKPEKDQGTVMLDVLLAAYGDENAEYKITKNHIKAFFV 306
Query: 271 NMFLGGSDTTSTTLEWAMAACKKSSH 296
++F+G +DT+ T++W MA ++H
Sbjct: 307 DLFIGATDTSVQTIQWTMAEIMNNTH 332
>At4g12300 flavonoid 3',5'-hydroxylase -like protein
Length = 516
Score = 61.2 bits (147), Expect = 7e-10
Identities = 35/83 (42%), Positives = 56/83 (67%), Gaps = 3/83 (3%)
Query: 210 LKKIKDSSEEMDDFLDRVIVEHK--MSRRDPKKKDFLDILLQLQD-DGLTEFELTQNDLK 266
+K++ + E+D LDR I + K R D + KDFL L++L+D +G +E +T N +K
Sbjct: 246 VKRMGVCARELDAVLDRAIEQMKPLRGRDDDEVKDFLQYLMKLKDQEGDSEVPITINHVK 305
Query: 267 ALLMNMFLGGSDTTSTTLEWAMA 289
ALL +M +GG+DT++ T+E+AMA
Sbjct: 306 ALLTDMVVGGTDTSTNTIEFAMA 328
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.353 0.156 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,126,129
Number of Sequences: 26719
Number of extensions: 218091
Number of successful extensions: 1526
Number of sequences better than 10.0: 166
Number of HSP's better than 10.0 without gapping: 133
Number of HSP's successfully gapped in prelim test: 33
Number of HSP's that attempted gapping in prelim test: 1312
Number of HSP's gapped (non-prelim): 179
length of query: 313
length of database: 11,318,596
effective HSP length: 99
effective length of query: 214
effective length of database: 8,673,415
effective search space: 1856110810
effective search space used: 1856110810
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 60 (27.7 bits)
Medicago: description of AC124952.12


