

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124217.1 - phase: 0 /pseudo
(1312 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
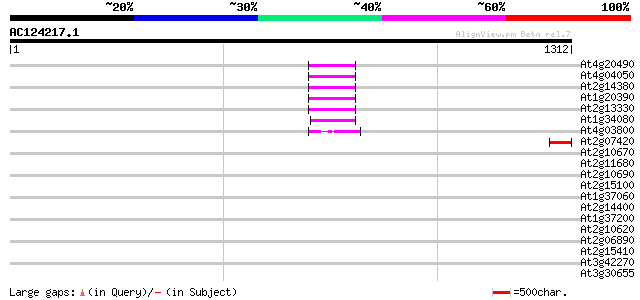
Score E
Sequences producing significant alignments: (bits) Value
At4g20490 putative protein 66 1e-10
At4g04050 putative transposon protein 63 1e-09
At2g14380 putative retroelement pol polyprotein 63 1e-09
At1g20390 hypothetical protein 63 1e-09
At2g13330 F14O4.9 60 1e-08
At1g34080 putative protein 53 1e-06
At4g03800 49 1e-05
At2g07420 putative retroelement pol polyprotein 46 2e-04
At2g10670 pseudogene 43 0.001
At2g11680 putative retroelement pol polyprotein 42 0.002
At2g10690 putative retroelement pol polyprotein 39 0.026
At2g15100 putative retroelement pol polyprotein 38 0.044
At1g37060 Athila retroelment ORF 1, putative 38 0.044
At2g14400 putative retroelement pol polyprotein 37 0.075
At1g37200 hypothetical protein 36 0.13
At2g10620 putative Athila retroelement ORF1 protein 35 0.22
At2g06890 putative retroelement integrase 35 0.28
At2g15410 putative retroelement pol polyprotein 35 0.37
At3g42270 putative protein 34 0.63
At3g30655 hypothetical protein, 3' partial 33 0.83
>At4g20490 putative protein
Length = 853
Score = 66.2 bits (160), Expect = 1e-10
Identities = 34/108 (31%), Positives = 62/108 (56%)
Query: 700 HLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSG 759
H L IK + + ++L+D G VNL+ L K+G+SES + P ++ + ++
Sbjct: 477 HNDALVIKLVMEDFDVERILIDTGSSVNLMFLKTLFKMGISESKIMPKIRPMTGYDGEAK 536
Query: 760 SSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVPST 807
S G +++ + VG + + T F+V+ S+ YN +LG+ WI+ + +PST
Sbjct: 537 MSIGEIKLQVQVGGITQKTKFVVIDSEPIYNAILGSPWIYSMKAIPST 584
>At4g04050 putative transposon protein
Length = 681
Score = 63.2 bits (152), Expect = 1e-09
Identities = 36/112 (32%), Positives = 58/112 (51%)
Query: 698 KIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERK 757
K H L I+ V L+ +++D G V++L K++G +S L+ L+
Sbjct: 533 KPHDDALVIRIDVGNYELSHIMIDTGSSVDVLFYDAFKRMGHLDSELQGRKTPLTGFAGD 592
Query: 758 SGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVPSTMH 809
+ S G +++ + V+R T FLVV KA +N +LG W+H + VPST H
Sbjct: 593 TTFSLGTIQLPTIARGVRRLTSFLVVNKKAPFNAILGRPWLHAMKAVPSTYH 644
>At2g14380 putative retroelement pol polyprotein
Length = 764
Score = 63.2 bits (152), Expect = 1e-09
Identities = 31/110 (28%), Positives = 62/110 (56%)
Query: 700 HLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSG 759
H PL I+ + + + ++L+D G VN++ + L+K+ + + ++KP L+ + +
Sbjct: 344 HNDPLVIELHIGESEVTRILIDTGSSVNVVFKDVLQKMKVHDRHIKPSVRPLTGFDGNTM 403
Query: 760 SSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVPSTMH 809
+ G +++ + +G F+VV YN++LGT WIHD+ +PS+ H
Sbjct: 404 MTNGTIKLPIYLGGAATWHKFVVVDKPTIYNIILGTPWIHDMQAIPSSYH 453
Score = 38.9 bits (89), Expect = 0.020
Identities = 19/47 (40%), Positives = 29/47 (61%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFIS 1311
LG+++ K+GIE N + +AI + KKE+Q G+I L RFI+
Sbjct: 697 LGYIVTKRGIEANPKQIRAILDLQSPRNKKEVQRLTGRIAGLNRFIA 743
>At1g20390 hypothetical protein
Length = 1791
Score = 63.2 bits (152), Expect = 1e-09
Identities = 31/110 (28%), Positives = 61/110 (55%)
Query: 700 HLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSG 759
H PL ++ ++ + +VL+D G V+L+ + L + +++ +KP + L+ +
Sbjct: 567 HNDPLVVELIISDSRVTRVLIDTGSSVDLIFKDVLTAMNITDRQIKPVSKPLAGFDGDFV 626
Query: 760 SSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVPSTMH 809
+ G +++ + VG + F+V+ A YN++LGT WIH + +PST H
Sbjct: 627 MTIGTIKLPIFVGGLIAWVKFVVIGKPAVYNVILGTPWIHQMQAIPSTYH 676
Score = 36.6 bits (83), Expect = 0.098
Identities = 19/47 (40%), Positives = 27/47 (57%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFIS 1311
LG+V+ K+GIE N + +AI +E+Q G+I L RFIS
Sbjct: 977 LGYVVTKRGIEANPKQIRAILELPSPRNAREVQRLTGRIAALNRFIS 1023
>At2g13330 F14O4.9
Length = 889
Score = 59.7 bits (143), Expect = 1e-08
Identities = 36/112 (32%), Positives = 57/112 (50%)
Query: 698 KIHLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERK 757
K H L I+ V L+ ++VD G V++L K+ G +S L+ L+
Sbjct: 59 KPHDDALVIRIDVGNYELSCIMVDTGSSVDVLFYDAFKRTGHLDSKLQGRKTPLTGFAGD 118
Query: 758 SGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVPSTMH 809
+ S G +++ + V++ T FLVV KA +N +LG W+H + VPST H
Sbjct: 119 TTFSIGTIQLPTIARGVRQLTNFLVVDKKAPFNAILGRPWLHVMKAVPSTYH 170
>At1g34080 putative protein
Length = 811
Score = 53.1 bits (126), Expect = 1e-06
Identities = 30/106 (28%), Positives = 54/106 (50%)
Query: 704 LFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSGSSFG 763
L I V + ++L+D G V+L+ L ++G+S+ ++K L ++ S G
Sbjct: 477 LVISLDVGNCEVQRILIDTGSSVDLIFLDTLVRMGISKKDIKGAPSPLVSFTIETSMSLG 536
Query: 764 VVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVPSTMH 809
+ + + V + F V A YN++LGT W++++ VPST H
Sbjct: 537 TITLPVTAQGVVKMIEFTVFDRPAAYNIILGTPWLYEMKAVPSTYH 582
>At4g03800
Length = 637
Score = 49.3 bits (116), Expect = 1e-05
Identities = 34/121 (28%), Positives = 58/121 (47%), Gaps = 14/121 (11%)
Query: 700 HLKPLFIKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSG 759
H L I V ++++LVD G V+L+ L P + ++++ +S
Sbjct: 199 HDDALIITLDVANFKISRILVDTGSSVDLI-------------FLGPPSPLVAFTS-ESA 244
Query: 760 SSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVPSTMH*RLALWREDG 819
S G +++ ++ G + + F+V A YN++LGT WI + VPST H L +G
Sbjct: 245 MSLGTIKLPVLAGKMSKIVDFVVFDKPATYNIILGTPWIFQMKAVPSTYHQWLKFPTSNG 304
Query: 820 V 820
V
Sbjct: 305 V 305
Score = 32.3 bits (72), Expect = 1.8
Identities = 15/39 (38%), Positives = 22/39 (55%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKI 1303
LG+++ K+GIE N N+ A T KE+Q G+I
Sbjct: 526 LGYIVTKRGIEANPNQINAFLKTPSLRNFKEVQRLTGRI 564
>At2g07420 putative retroelement pol polyprotein
Length = 1664
Score = 45.8 bits (107), Expect = 2e-04
Identities = 22/50 (44%), Positives = 33/50 (66%)
Query: 1263 ILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
I+LG I+KKGIE+++ K K I +P T K++++FLG F RFI +
Sbjct: 1524 IVLGHKISKKGIEVDKGKIKVIMQLQPPKTVKDIRSFLGHAGFYRRFIKD 1573
>At2g10670 pseudogene
Length = 929
Score = 43.1 bits (100), Expect = 0.001
Identities = 31/96 (32%), Positives = 55/96 (57%), Gaps = 5/96 (5%)
Query: 712 QIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSGSSFGVVE-VDLV 770
Q+ N L D G V+L+P S +KK+G + KP ++ L ++R S + FG++E V ++
Sbjct: 585 QLTFNNCLCDLGASVSLMPLSVVKKLGF--VHYKPCDLTLILADRTSKTPFGLLEDVPVM 642
Query: 771 VGTVKRSTLFLVVA--SKANYNLLLGTEWIHDVGDV 804
+ V+ T F+V+ ++ L+LG ++ VG V
Sbjct: 643 INGVEVPTDFVVLEMDGESKDPLILGRPFLASVGAV 678
>At2g11680 putative retroelement pol polyprotein
Length = 301
Score = 42.4 bits (98), Expect = 0.002
Identities = 19/50 (38%), Positives = 32/50 (64%)
Query: 1263 ILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
I+LG I+KKG+++N+ K + +P T K++++FLG F RFI +
Sbjct: 6 IVLGHKISKKGVQVNKAKVDVMMQLQPPKTGKDIRSFLGHAGFYRRFIKD 55
>At2g10690 putative retroelement pol polyprotein
Length = 622
Score = 38.5 bits (88), Expect = 0.026
Identities = 19/50 (38%), Positives = 31/50 (62%)
Query: 1263 ILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
I+LG I+ KGIE+++ K + +P T K++++FLG F RFI +
Sbjct: 251 IVLGHKISGKGIEVDKAKIDVMIQLQPPKTVKDIRSFLGHAEFYRRFIKD 300
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 37.7 bits (86), Expect = 0.044
Identities = 18/53 (33%), Positives = 29/53 (53%)
Query: 757 KSGSSFGVVEVDLVVGTVKRSTLFLVVASKANYNLLLGTEWIHDVGDVPSTMH 809
++ S G V + + V + F V A YN++LGT W++++ VPST H
Sbjct: 348 ETSMSLGTVVLPVTAQGVVKMVEFTVFDRPAAYNVILGTPWLYEMKVVPSTYH 400
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 37.7 bits (86), Expect = 0.044
Identities = 23/72 (31%), Positives = 43/72 (58%), Gaps = 3/72 (4%)
Query: 713 IGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSGSSFGVVE-VDLVV 771
+ +K L D G V+L+P KK+G ++ KP N++L ++R S G++E + +++
Sbjct: 611 LAFSKCLCDLGASVSLMPLPVAKKLGFNK--YKPCNISLILADRSVRISHGLLEDLPVMI 668
Query: 772 GTVKRSTLFLVV 783
G V+ T F+V+
Sbjct: 669 GVVEVPTDFVVL 680
Score = 34.7 bits (78), Expect = 0.37
Identities = 17/50 (34%), Positives = 31/50 (62%)
Query: 1263 ILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
I+LG I+++GIE+++ K + +P T K++++FLG F FI +
Sbjct: 1115 IVLGRKISEEGIEVDKAKIDVMMQLQPPKTVKDIRSFLGHAGFYRIFIKD 1164
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 37.0 bits (84), Expect = 0.075
Identities = 19/47 (40%), Positives = 26/47 (54%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFIS 1311
LG+++ K+GIE N N+ A KE+Q G+I L RFIS
Sbjct: 664 LGYIVTKRGIEANPNQINAFLNMPSPKNFKEVQRLTGRIAALNRFIS 710
>At1g37200 hypothetical protein
Length = 1564
Score = 36.2 bits (82), Expect = 0.13
Identities = 31/95 (32%), Positives = 43/95 (44%), Gaps = 21/95 (22%)
Query: 736 KIGLSESNL----KPHNVALSYSERKSGSSFGVVEVDLVVGTVKRSTLFLVVAS---KAN 788
KI L ES KPH+ A+ V+ +D VG K S + + S KA
Sbjct: 486 KITLWESETTDLDKPHDDAI------------VIRID--VGNYKLSRIMIDTGSSVDKAP 531
Query: 789 YNLLLGTEWIHDVGDVPSTMH*RLALWREDGVLEI 823
+N +LG W+H + VPST H + E G+ I
Sbjct: 532 FNAILGRPWLHAMKAVPSTYHQCIKFPSEKGIAVI 566
>At2g10620 putative Athila retroelement ORF1 protein
Length = 451
Score = 35.4 bits (80), Expect = 0.22
Identities = 34/151 (22%), Positives = 71/151 (46%), Gaps = 13/151 (8%)
Query: 659 DEEDFIKEMVVHKTM---CYYVMNNGCVEGQHAMFERPDYHMKIHLKPLF-IKAKVNQIG 714
D F+K+++V + V+++ C A+ ++ H K+ F + + +
Sbjct: 252 DSHKFLKDLIVERIQELQGMVVLSDEC----SAIIQKKIIHKKLSDPGSFTLPCSLGPLA 307
Query: 715 LNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSGSSFGVVE-VDLVVGT 773
NK L D G +L+P S K++G ++ K N++L ++R +E + + +
Sbjct: 308 FNKCLCDLGASASLMPLSVTKRLGFTQ--YKSCNISLILADRSVRIPHDFLENIPIRIRA 365
Query: 774 VKRSTLFLVVA--SKANYNLLLGTEWIHDVG 802
V+ T F+V+ + +L+LG ++ VG
Sbjct: 366 VEIPTDFVVLEMDEEPKDHLILGRPFLTTVG 396
>At2g06890 putative retroelement integrase
Length = 1215
Score = 35.0 bits (79), Expect = 0.28
Identities = 16/51 (31%), Positives = 29/51 (56%)
Query: 1262 VILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
++ LGFV++ G+++++ K KAI T E+++F G F RF +
Sbjct: 615 LVFLGFVVSADGVKVDEEKVKAIRDWPSPKTVGEVRSFHGLAGFYRRFFKD 665
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 34.7 bits (78), Expect = 0.37
Identities = 17/47 (36%), Positives = 27/47 (57%)
Query: 1265 LGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFIS 1311
LG+++ ++GIE N + AI +E+Q +G+I L RFIS
Sbjct: 982 LGYLVTRRGIEANPKQISAIIDLPSPRNTREVQRLIGRIAALNRFIS 1028
>At3g42270 putative protein
Length = 747
Score = 33.9 bits (76), Expect = 0.63
Identities = 18/59 (30%), Positives = 37/59 (62%), Gaps = 3/59 (5%)
Query: 715 LNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSGSSFGVVE-VDLVVG 772
L++ L D G ++NL+P+S +++G+ +N P + L +++R + G++E V + VG
Sbjct: 479 LSRSLCDLGSIINLMPKSVAERLGM--TNYGPTRITLLFADRSNRIPEGILEDVPIKVG 535
>At3g30655 hypothetical protein, 3' partial
Length = 660
Score = 33.5 bits (75), Expect = 0.83
Identities = 17/61 (27%), Positives = 33/61 (53%), Gaps = 2/61 (3%)
Query: 706 IKAKVNQIGLNKVLVDGGVVVNLLPQSFLKKIGLSESNLKPHNVALSYSERKSGSSFGVV 765
+ + Q+ + L D G V+++P S +K+G + KP ++ L ++R S FG++
Sbjct: 599 LPCSIRQLTFSNCLCDLGASVSIMPLSVARKLGFVQ--YKPSDLTLILADRTSRRPFGLL 656
Query: 766 E 766
E
Sbjct: 657 E 657
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.358 0.160 0.568
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,460,388
Number of Sequences: 26719
Number of extensions: 1006808
Number of successful extensions: 5385
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 21
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 5345
Number of HSP's gapped (non-prelim): 49
length of query: 1312
length of database: 11,318,596
effective HSP length: 111
effective length of query: 1201
effective length of database: 8,352,787
effective search space: 10031697187
effective search space used: 10031697187
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC124217.1


