

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124216.5 - phase: 0
(340 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
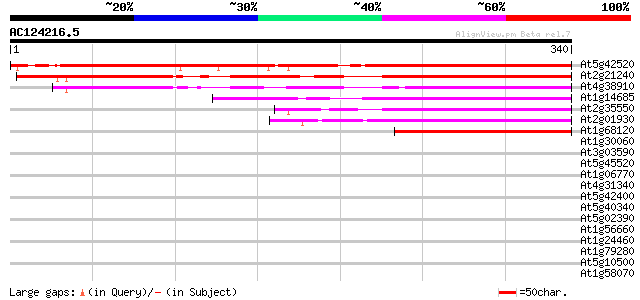
Score E
Sequences producing significant alignments: (bits) Value
At5g42520 unknown protein 434 e-122
At2g21240 unknown protein 294 5e-80
At4g38910 unknown protein 255 2e-68
At1g14685 unknown protein 156 1e-38
At2g35550 unknown protein 153 1e-37
At2g01930 unknown protein 151 6e-37
At1g68120 hypothetical protein 148 4e-36
At1g30060 hypothetical protein 39 0.005
At3g03590 unknown protein 35 0.077
At5g45520 putative protein 34 0.13
At1g06770 unknown protein 33 0.17
At4g31340 unknown protein 33 0.29
At5g42400 trithorax-related protein 7 (ATXR7) 31 0.85
At5g40340 unknown protein 31 0.85
At5g02390 putative protein 31 0.85
At1g56660 hypothetical protein 31 0.85
At1g24460 unknown protein 31 0.85
At1g79280 hypothetical protein 31 1.1
At5g10500 unknown protein 30 1.4
At1g58070 unknown protein 30 1.4
>At5g42520 unknown protein
Length = 342
Score = 434 bits (1115), Expect = e-122
Identities = 237/360 (65%), Positives = 279/360 (76%), Gaps = 38/360 (10%)
Query: 1 MDD---RENGRHKADQYKSAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLALSEK 57
MDD RENGRHKA + QGQWLMQ HQ PSMKQ+MSI+AERDAAIQERNLA+SEK
Sbjct: 1 MDDGGHRENGRHKA----AVQGQWLMQ---HQ-PSMKQVMSIIAERDAAIQERNLAISEK 52
Query: 58 KAALAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANG---GMSSCPPGCQIS 114
KAA+AERDMAFLQRDTAIAERNNA+MERD+A+ LQ+REN++ MS+CPPGCQIS
Sbjct: 53 KAAVAERDMAFLQRDTAIAERNNAIMERDSALTALQYRENSMVTAPAANMSACPPGCQIS 112
Query: 115 RGVKHIHHLPQ----QVNHLPNMGDSSYGTRELHTTDALPAAPVS---LEVGKPPRRAKR 167
RGVKH+HH Q +H+P + +++Y TRE+ D LP +P + LE KP +R KR
Sbjct: 113 RGVKHLHHPHMHHHHQQHHIPQLTENAYETREMEPNDGLPTSPPAGSTLESAKP-KRGKR 171
Query: 168 --PKESKSDSPNKKTPKS-RKVKKEG-DDLNKTMFANNKELEWKSSEEIINGDDDLNKQL 223
PK + + NK+ PK+ RKVKKE DDLNK MF K++ + D+D +K +
Sbjct: 172 VNPKATTQTAANKRGPKNQRKVKKESEDDLNKIMFV-------KTTHDYT--DEDSSKHI 222
Query: 224 AI-SKADWKPQDLA-LNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVY 281
I SK+DWK Q++ LNQV YD++TMP P CSCTGVLRQCYKWGNGGWQS+CCTTTLS+Y
Sbjct: 223 LIGSKSDWKSQEMVGLNQVVYDETTMPPPVCSCTGVLRQCYKWGNGGWQSSCCTTTLSMY 282
Query: 282 PLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEG-HDLSHPVDLKDHWAKHGTNRYITIK 340
PLPA+PNKRHARVGGRKMSGSAFNKLLSRLAAEG HDLS+PVDLKDHWAKHGTNRYITIK
Sbjct: 283 PLPALPNKRHARVGGRKMSGSAFNKLLSRLAAEGHHDLSNPVDLKDHWAKHGTNRYITIK 342
>At2g21240 unknown protein
Length = 296
Score = 294 bits (752), Expect = 5e-80
Identities = 167/342 (48%), Positives = 210/342 (60%), Gaps = 59/342 (17%)
Query: 5 ENGRHKADQYKSAQGQW-LMQQHQ--HQHPSM---KQIMSIMAERDAAIQERNLALSEKK 58
+N R K D +K AQ W ++ QHQ QH ++ K+IMSI+AERDAA+ ERN A+S KK
Sbjct: 8 DNARFKPDYFKGAQSMWNMIPQHQIKEQHNALVMNKKIMSILAERDAAVHERNQAVSAKK 67
Query: 59 AALAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANGGMSSCPPGCQISRGVK 118
A+A RD A QRD A++ER+ AL+ERDNA A LQ EN+L N +S G K
Sbjct: 68 EAVAARDEALQQRDKALSERDKALIERDNAYAALQHHENSL-NFALS----------GGK 116
Query: 119 HIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNK 178
+ GD +G E H + P + + EV KR KE+K
Sbjct: 117 CVD------------GDDCFGIGEPHKLEVFPLSTIPPEVTNTKVVNKRKKENKQGLS-- 162
Query: 179 KTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKPQDLALN 238
KVKK G+DLN+ + A K+ S+ DW QD+ LN
Sbjct: 163 ------KVKKVGEDLNRRVPAPGKK----------------------SRTDWDSQDVGLN 194
Query: 239 QVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRK 298
V +D++TMP P CSCTG RQCYKWGNGGWQS+CCTTTLS YPLP +PNKRH+R+GGRK
Sbjct: 195 LVTFDETTMPVPMCSCTGSTRQCYKWGNGGWQSSCCTTTLSQYPLPQMPNKRHSRMGGRK 254
Query: 299 MSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
MSG+ F++LLSRL+AEG+DLS PVDLKD+WA+HGTNRYITIK
Sbjct: 255 MSGNVFSRLLSRLSAEGYDLSCPVDLKDYWARHGTNRYITIK 296
>At4g38910 unknown protein
Length = 258
Score = 255 bits (652), Expect = 2e-68
Identities = 142/317 (44%), Positives = 191/317 (59%), Gaps = 65/317 (20%)
Query: 27 QHQHPSM---KQIMSIMAERDAAIQERNLALSEKKAALAERDMAFLQRDTAIAERNNALM 83
+ QH ++ K+I+SI+AERDAA++ERN A++ K ALA RD A QRD A++ER+NA+M
Sbjct: 4 KEQHNALVMNKKIISILAERDAAVKERNEAVAATKEALASRDEALEQRDKALSERDNAIM 63
Query: 84 ERDNAIATLQFRENALANGGMSSCPPGCQISRGVKHIHHLPQQVNHLPNMGDSSYGTREL 143
E ++A+ L++REN L + SC RG G + T E
Sbjct: 64 ETESALNALRYRENNL--NYILSCA-----KRG-----------------GSQRFITEES 99
Query: 144 HTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKE 203
H + P + + P E+ + P K+ +S++ KK G+DLN+ + + K+
Sbjct: 100 HLPNPSPISTI-------------PPEAANTRPTKRKKESKQGKKMGEDLNRPVASPGKK 146
Query: 204 LEWKSSEEIINGDDDLNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYK 263
S+ DW D+ V +D+ TMP P C+CTG RQCYK
Sbjct: 147 ----------------------SRKDWDSNDVL---VTFDEMTMPVPMCTCTGTARQCYK 181
Query: 264 WGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVD 323
WGNGGWQS+CCTTTLS YPLP +PNKRH+RVGGRKMSGS F++LLSRLA EGH+LS PVD
Sbjct: 182 WGNGGWQSSCCTTTLSEYPLPQMPNKRHSRVGGRKMSGSVFSRLLSRLAGEGHELSSPVD 241
Query: 324 LKDHWAKHGTNRYITIK 340
LK++WA+HGTNRYITIK
Sbjct: 242 LKNYWARHGTNRYITIK 258
>At1g14685 unknown protein
Length = 279
Score = 156 bits (395), Expect = 1e-38
Identities = 83/217 (38%), Positives = 119/217 (54%), Gaps = 23/217 (10%)
Query: 124 PQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNKKTPKS 183
P N LP + LH PV LE + KR +K+ S TPK+
Sbjct: 86 PNYGNVLPETSSAPSMQMNLHHHLQTEENPVKLEEEIVVQTKKRKTNAKAGS----TPKA 141
Query: 184 RKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKPQDLALNQVAYD 243
+K +K D+ + N +++ N + K K DL +N V+ D
Sbjct: 142 KKPRKPKDENS-------------------NNNNNNNTNVTRVKPAKKSVDLVINGVSMD 182
Query: 244 DSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHARVGGRKMSGSA 303
S +P P C+CTG +QCY+WG GGWQSACCTT +S++PLP +R AR+ GRKMS A
Sbjct: 183 ISGLPVPICTCTGAPQQCYRWGCGGWQSACCTTNISMHPLPMSTKRRGARISGRKMSQGA 242
Query: 304 FNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
F K+L +LA++G + +P+DLK HWA+HGTN+++TI+
Sbjct: 243 FKKVLEKLASDGFNFGNPIDLKSHWARHGTNKFVTIR 279
>At2g35550 unknown protein
Length = 271
Score = 153 bits (386), Expect = 1e-37
Identities = 81/182 (44%), Positives = 109/182 (59%), Gaps = 20/182 (10%)
Query: 161 PPRRAKR--PKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDD 218
PP R R + K S N+ K+ K K + K +NK + S E
Sbjct: 108 PPIRDSRNDTETVKQKSVNQSPSKALKPKPQ----RKKRSVSNKSKKTPSIPE------- 156
Query: 219 LNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTL 278
+K + K D+ ++ ++D S +P P CSCTGV R CYKWG GGWQS+CCT ++
Sbjct: 157 -------TKREKKNLDINIDISSFDTSGVPPPVCSCTGVSRVCYKWGMGGWQSSCCTISI 209
Query: 279 SVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYIT 338
S YPLP + AR+ GRKMS A+ KLL+RLA EG+DLSHP+DLK+HWA+HGTN+++T
Sbjct: 210 STYPLPMSTTRPGARLAGRKMSNGAYVKLLARLADEGYDLSHPLDLKNHWARHGTNKFVT 269
Query: 339 IK 340
IK
Sbjct: 270 IK 271
>At2g01930 unknown protein
Length = 283
Score = 151 bits (381), Expect = 6e-37
Identities = 76/186 (40%), Positives = 113/186 (59%), Gaps = 7/186 (3%)
Query: 158 VGKPPRRAKRPKESKSDSP---NKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIIN 214
+ +P + R +E+ P ++T K RK++ G T+ K + K ++ N
Sbjct: 102 IHQPVLNSSRFEENPIPPPAPCEEQTGKKRKMR--GSIATPTVPKAKKMRKPKEERDVTN 159
Query: 215 GDDDLNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACC 274
+++ +Q K K DL +N V+ D S +P P C+CTG +QCY+WG GGWQSACC
Sbjct: 160 --NNVQQQQQRVKPVKKSVDLVINGVSMDISGLPVPVCTCTGTPQQCYRWGCGGWQSACC 217
Query: 275 TTTLSVYPLPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTN 334
TT +SVYPLP +R AR+ GRKMS AF K+L +L+ EG+ + +DLK HWA+HGTN
Sbjct: 218 TTNISVYPLPMSTKRRGARISGRKMSQGAFKKVLEKLSTEGYSFGNAIDLKSHWARHGTN 277
Query: 335 RYITIK 340
+++TI+
Sbjct: 278 KFVTIR 283
>At1g68120 hypothetical protein
Length = 269
Score = 148 bits (374), Expect = 4e-36
Identities = 60/107 (56%), Positives = 83/107 (77%)
Query: 234 DLALNQVAYDDSTMPAPACSCTGVLRQCYKWGNGGWQSACCTTTLSVYPLPAVPNKRHAR 293
++ +N V+ D +P P CSCTG+ +QCY+WG GGWQSACCTT +S+YPLP +R AR
Sbjct: 163 EMVINGVSMDIGGLPVPVCSCTGMPQQCYRWGCGGWQSACCTTNVSMYPLPVNTKRRGAR 222
Query: 294 VGGRKMSGSAFNKLLSRLAAEGHDLSHPVDLKDHWAKHGTNRYITIK 340
+ GRKMS AF K+L +L+++G D S+P+DLK HWAKHGTN+++TI+
Sbjct: 223 IAGRKMSQGAFRKVLEKLSSDGFDFSNPIDLKSHWAKHGTNKFVTIR 269
>At1g30060 hypothetical protein
Length = 176
Score = 38.5 bits (88), Expect = 0.005
Identities = 21/77 (27%), Positives = 42/77 (54%), Gaps = 1/77 (1%)
Query: 154 VSLEVGKPPRRAKRPKESKSDSPNKKTPKSRKVK-KEGDDLNKTMFANNKELEWKSSEEI 212
+S+ + + + R +S+SD+ ++K P S K K E + N T N+++ EE
Sbjct: 87 LSVHLVRVEEKEMRKSKSESDNKSRKKPSSSKAKGLESGEGNVTRMGENQQICDADQEEP 146
Query: 213 INGDDDLNKQLAISKAD 229
++GD+++ K +A + D
Sbjct: 147 VDGDEEMKKTIAKAWTD 163
>At3g03590 unknown protein
Length = 143
Score = 34.7 bits (78), Expect = 0.077
Identities = 17/44 (38%), Positives = 24/44 (53%)
Query: 145 TTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNKKTPKSRKVKK 188
T+ + A KP +AK ++KSDSP KKTP+S + K
Sbjct: 23 TSSLVSAGSTKKPAAKPKAKAKPKPKAKSDSPAKKTPRSTGIFK 66
>At5g45520 putative protein
Length = 1167
Score = 33.9 bits (76), Expect = 0.13
Identities = 24/81 (29%), Positives = 41/81 (49%), Gaps = 16/81 (19%)
Query: 161 PPRRAKRPKESKSDSPNKKTPKSRKVKKEGD-------DLNKTMFANNKELEWKSSEEII 213
PP+ +K ESK D + + KV+++GD DL++ N+ + E S+++I
Sbjct: 880 PPKESKDTMESKRDDQKENS----KVQEKGDVDKGKAADLDEGKKENDVKAESSKSDKVI 935
Query: 214 NGDDDLN-----KQLAISKAD 229
GD++ N K + SK D
Sbjct: 936 EGDEEKNPPQKSKDIIQSKPD 956
>At1g06770 unknown protein
Length = 421
Score = 33.5 bits (75), Expect = 0.17
Identities = 35/114 (30%), Positives = 52/114 (44%), Gaps = 14/114 (12%)
Query: 154 VSLEVGKPPRRAKRP--KESKSDSPNKKTPKSRKVKKEGDDLNKT--------MFANNKE 203
VS + G RR K P KE ++ S ++T K K + GD+L ++ F NK
Sbjct: 116 VSAQAGTTRRRTKAPTRKELRNGSLAERTVK--KEESSGDELLESTSSPDTLNKFTQNKR 173
Query: 204 LEWKS-SEEIINGDDDLNKQLAISKADWKPQDLALNQVAYDDSTMPAPACSCTG 256
KS E I N ++ + SK DWKP + L +VA + + A +G
Sbjct: 174 QSKKSCKESISNKENKDGDEPWDSKMDWKPLNF-LVEVANGTKPLKSSASQGSG 226
>At4g31340 unknown protein
Length = 437
Score = 32.7 bits (73), Expect = 0.29
Identities = 24/102 (23%), Positives = 46/102 (44%), Gaps = 4/102 (3%)
Query: 47 IQERNLALSEKKAALAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANGGMSS 106
+ + N + ++ + E+ +D +AE+ L ER++ IA+LQ ++L G S
Sbjct: 43 LDQLNAKIRALESQIDEKTREVQGKDEVVAEKEKLLKEREDKIASLQTEVSSLQKKGSSD 102
Query: 107 CPPGCQISRGVKHIHHLPQQVNHLPNMGDSSYGTRELHTTDA 148
Q+ + L +QV L N + +E +T+A
Sbjct: 103 SAK--QLGKAQARADELEKQVEVLKNFLEQK--NKEKDSTEA 140
>At5g42400 trithorax-related protein 7 (ATXR7)
Length = 1421
Score = 31.2 bits (69), Expect = 0.85
Identities = 14/36 (38%), Positives = 21/36 (57%)
Query: 144 HTTDALPAAPVSLEVGKPPRRAKRPKESKSDSPNKK 179
HTT+ P +S++ G+P A +P E S P+KK
Sbjct: 1138 HTTERSPIKDLSVDDGRPKPIALKPLEKLSSKPSKK 1173
>At5g40340 unknown protein
Length = 1008
Score = 31.2 bits (69), Expect = 0.85
Identities = 16/43 (37%), Positives = 24/43 (55%), Gaps = 2/43 (4%)
Query: 155 SLEVGKPPRRAKRPKESKSDSPN--KKTPKSRKVKKEGDDLNK 195
S+E K R+ K+PK + + PN +K K +K K+EG K
Sbjct: 813 SVESTKKERKRKKPKHDEEEVPNETEKPEKKKKKKREGKSKKK 855
>At5g02390 putative protein
Length = 835
Score = 31.2 bits (69), Expect = 0.85
Identities = 23/80 (28%), Positives = 36/80 (44%), Gaps = 1/80 (1%)
Query: 147 DALPAAPVSLEVGKPPRRAKRPKESKSDSPNKKTPKS-RKVKKEGDDLNKTMFANNKELE 205
D +P + LE +RPK+ S +K++ S K KK+ + K+ N++E
Sbjct: 67 DGIPVPKIKLEDKTDVESVQRPKKPSSAVKSKESSNSGEKTKKKHNPEEKSKKLNSEERS 126
Query: 206 WKSSEEIINGDDDLNKQLAI 225
K+ EI L K L I
Sbjct: 127 RKTHSEIKRSVKALIKALVI 146
>At1g56660 hypothetical protein
Length = 522
Score = 31.2 bits (69), Expect = 0.85
Identities = 21/66 (31%), Positives = 33/66 (49%), Gaps = 2/66 (3%)
Query: 157 EVGKPPRRAKRPKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGD 216
E K P++ K+ KE + +KK K +K K E DL K KE + ++ +E+ D
Sbjct: 189 EKKKKPKKEKKQKEESKSNEDKKV-KGKKEKGEKGDLEKEDEEKKKEHD-ETDQEMKEKD 246
Query: 217 DDLNKQ 222
NK+
Sbjct: 247 SKKNKK 252
>At1g24460 unknown protein
Length = 1791
Score = 31.2 bits (69), Expect = 0.85
Identities = 43/191 (22%), Positives = 79/191 (40%), Gaps = 29/191 (15%)
Query: 56 EKKAALAERDMAFLQRDTAIAERNNALMERDNAIATLQFRENALANGG-MSSCPPGCQIS 114
E + ++D++ L A R L ++N + +QF+EN NGG M S ++
Sbjct: 1269 ENTISALQKDLSSLISACGAAARELQLEVKNNLLELVQFQEN--ENGGEMESTEDPQEL- 1325
Query: 115 RGVKHIHHLPQQVNHLPNMGDSSYGTRELHTTDALPAAPVSLEVGKPPRRAKRPKESKSD 174
H+ Q++ L + + + T +L T AA V D
Sbjct: 1326 ----HVSECAQRIKELSSAAEKACATLKLFETTNNAAATVI-----------------RD 1364
Query: 175 SPNKKTPKSRKVKKE--GDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISKADWKP 232
N+ T S ++K DLN+T ++ E + +S EE+ + L + + W
Sbjct: 1365 MENRLTEASVALEKAVLERDLNQTK-VSSSEAKVESLEELCQDLKLQLENLRVKEEKWHE 1423
Query: 233 QDLALNQVAYD 243
+++ L+ + YD
Sbjct: 1424 KEVELSTL-YD 1433
>At1g79280 hypothetical protein
Length = 2111
Score = 30.8 bits (68), Expect = 1.1
Identities = 22/78 (28%), Positives = 36/78 (45%), Gaps = 15/78 (19%)
Query: 15 KSAQGQWLMQQHQHQHPSMKQIMSIMAERDAAIQERNLAL--------------SEKKAA 60
K + + + + H S +Q+M ++ ++DA I E+N + SEK+A
Sbjct: 106 KDGEVERMSTEMSELHKSKRQLMELLEQKDAEISEKNSTIKSYLDKIVKLTDTSSEKEAR 165
Query: 61 LAERDMAFLQRDTAIAER 78
LAE A L R A+ R
Sbjct: 166 LAEA-TAELARSQAMCSR 182
>At5g10500 unknown protein
Length = 848
Score = 30.4 bits (67), Expect = 1.4
Identities = 16/72 (22%), Positives = 37/72 (51%), Gaps = 1/72 (1%)
Query: 168 PKESKSDSPNKKTPKSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQLAISK 227
P+ K+ + NK PK + + + K MF + K ++ +++ ++N L+K A+ +
Sbjct: 122 PRHHKNKTSNKNVPKVPDLPIKDPEAAKKMFMSRKAIQEQNASSVVN-KSGLSKTEAVEE 180
Query: 228 ADWKPQDLALNQ 239
D +++ + Q
Sbjct: 181 IDKLQKEILVLQ 192
>At1g58070 unknown protein
Length = 284
Score = 30.4 bits (67), Expect = 1.4
Identities = 15/42 (35%), Positives = 23/42 (54%)
Query: 182 KSRKVKKEGDDLNKTMFANNKELEWKSSEEIINGDDDLNKQL 223
KS+ K E D K+ A K + +K ++E NG+D K+L
Sbjct: 172 KSKIAKAEADAAGKSAAAMTKSVSFKDTKEKENGEDQRRKEL 213
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.128 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,077,403
Number of Sequences: 26719
Number of extensions: 353713
Number of successful extensions: 1256
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 1158
Number of HSP's gapped (non-prelim): 90
length of query: 340
length of database: 11,318,596
effective HSP length: 100
effective length of query: 240
effective length of database: 8,646,696
effective search space: 2075207040
effective search space used: 2075207040
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC124216.5


