

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.5 - phase: 0
(266 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
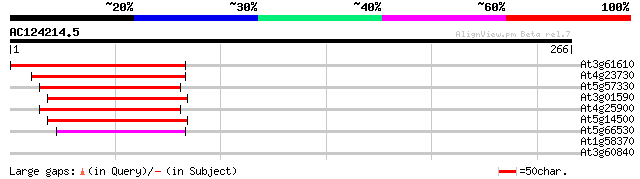
Score E
Sequences producing significant alignments: (bits) Value
At3g61610 unknown protein 121 4e-28
At4g23730 unknown protein 92 3e-19
At5g57330 apospory-associated protein C 87 1e-17
At3g01590 unknown protein 80 1e-15
At4g25900 possible apospory-associated like protein 79 3e-15
At5g14500 unknown protein 74 1e-13
At5g66530 apospory-associated protein C-like 54 1e-07
At1g58370 unknown protein 30 1.7
At3g60840 putative protein 29 2.3
>At3g61610 unknown protein
Length = 317
Score = 121 bits (303), Expect = 4e-28
Identities = 55/83 (66%), Positives = 65/83 (78%)
Query: 1 MGHSAAVWDFRAATEFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTS 60
MG A VWD + A+E KDWNG+DQV+LR P GASA++SLHG QV SWRNE GEELLFTS
Sbjct: 1 MGQYATVWDQKEASEIIKDWNGVDQVLLRNPHGASAKISLHGGQVISWRNELGEELLFTS 60
Query: 61 CKTLSKAPKAIRGGIPICFPQAG 83
K + K PK++RGGI IC+PQ G
Sbjct: 61 NKAIFKPPKSMRGGIQICYPQFG 83
>At4g23730 unknown protein
Length = 306
Score = 92.0 bits (227), Expect = 3e-19
Identities = 40/73 (54%), Positives = 52/73 (70%)
Query: 11 RAATEFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKA 70
R + KD NG DQ++L+ P+GAS ++SLHG QV SW+ ++G+ELLF S K K P
Sbjct: 12 RPTVDLVKDRNGTDQILLQNPRGASVKISLHGGQVLSWKTDKGDELLFNSTKANLKPPHP 71
Query: 71 IRGGIPICFPQAG 83
+RGGIPICFPQ G
Sbjct: 72 VRGGIPICFPQFG 84
>At5g57330 apospory-associated protein C
Length = 312
Score = 86.7 bits (213), Expect = 1e-17
Identities = 40/67 (59%), Positives = 49/67 (72%)
Query: 15 EFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGG 74
E K NG+D++VLR +G SA V L+G+ VTSW+NE GEELL S K + K PK IRGG
Sbjct: 9 ELAKGINGLDKIVLRESRGRSAEVYLYGSHVTSWKNENGEELLHLSSKAIFKPPKPIRGG 68
Query: 75 IPICFPQ 81
IP+CFPQ
Sbjct: 69 IPLCFPQ 75
>At3g01590 unknown protein
Length = 306
Score = 79.7 bits (195), Expect = 1e-15
Identities = 36/66 (54%), Positives = 48/66 (72%)
Query: 19 DWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIPIC 78
D +G +++L P+G++A V L G QV SW+NE+ EELL+ S K K PKAIRGGIP+C
Sbjct: 8 DGDGSSRIILTEPRGSTAEVLLFGGQVISWKNERREELLYMSSKAQYKPPKAIRGGIPVC 67
Query: 79 FPQAGH 84
FPQ G+
Sbjct: 68 FPQFGN 73
>At4g25900 possible apospory-associated like protein
Length = 318
Score = 78.6 bits (192), Expect = 3e-15
Identities = 36/67 (53%), Positives = 48/67 (70%)
Query: 15 EFTKDWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGG 74
E TK NG+D++++R +G SA V L+G QV+SW+NE GEELL S K + + P IRGG
Sbjct: 32 ERTKGVNGLDKIIIRDRRGRSAEVYLYGGQVSSWKNENGEELLVMSSKAIFQPPTPIRGG 91
Query: 75 IPICFPQ 81
IP+ FPQ
Sbjct: 92 IPVLFPQ 98
>At5g14500 unknown protein
Length = 306
Score = 73.6 bits (179), Expect = 1e-13
Identities = 34/66 (51%), Positives = 47/66 (70%)
Query: 19 DWNGIDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIPIC 78
D +G +++L P G++A V L+G QV SW+NE+ E+LL+ S K K PKAIRGG+PI
Sbjct: 8 DRDGSSRILLTDPAGSTAEVLLYGGQVVSWKNERREKLLYMSTKAQLKPPKAIRGGLPIS 67
Query: 79 FPQAGH 84
FPQ G+
Sbjct: 68 FPQFGN 73
>At5g66530 apospory-associated protein C-like
Length = 307
Score = 53.5 bits (127), Expect = 1e-07
Identities = 25/61 (40%), Positives = 36/61 (58%)
Query: 23 IDQVVLRTPQGASARVSLHGAQVTSWRNEQGEELLFTSCKTLSKAPKAIRGGIPICFPQA 82
+ ++VL +PQ + A + L G +TSW+ G++LLF + K I GGIP CFPQ
Sbjct: 45 LPKLVLTSPQNSEAEIYLFGGCITSWKVASGKDLLFVRPDAVFNKIKPISGGIPHCFPQF 104
Query: 83 G 83
G
Sbjct: 105 G 105
>At1g58370 unknown protein
Length = 917
Score = 29.6 bits (65), Expect = 1.7
Identities = 17/43 (39%), Positives = 25/43 (57%), Gaps = 5/43 (11%)
Query: 10 FRAATEFTKDWNGIDQVVLRTPQ-----GASARVSLHGAQVTS 47
F +ATE T++WNGI Q + Q A+A V ++G VT+
Sbjct: 238 FASATERTQNWNGIQQEITGKVQRKRVYEATAVVRIYGNNVTT 280
>At3g60840 putative protein
Length = 648
Score = 29.3 bits (64), Expect = 2.3
Identities = 11/23 (47%), Positives = 16/23 (68%)
Query: 43 AQVTSWRNEQGEELLFTSCKTLS 65
A+VT+W NE+G E L+ + LS
Sbjct: 407 AKVTAWENERGNEFLYDGVRVLS 429
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.351 0.152 0.549
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,016,923
Number of Sequences: 26719
Number of extensions: 166872
Number of successful extensions: 562
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 554
Number of HSP's gapped (non-prelim): 9
length of query: 266
length of database: 11,318,596
effective HSP length: 98
effective length of query: 168
effective length of database: 8,700,134
effective search space: 1461622512
effective search space used: 1461622512
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC124214.5


