

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.3 - phase: 0
(529 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
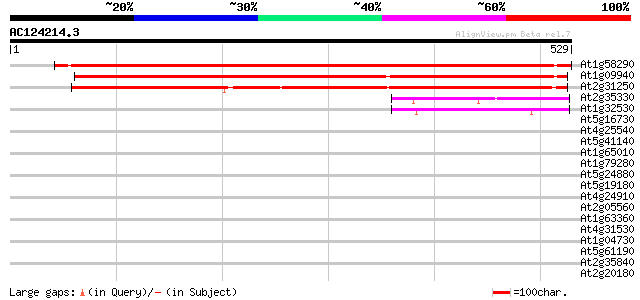
Score E
Sequences producing significant alignments: (bits) Value
At1g58290 putative protein 643 0.0
At1g09940 putative glutamyl-tRNA reductase 2 precursor 635 0.0
At2g31250 putative glutamyl tRNA reductase 536 e-152
At2g35330 unknown protein 53 5e-07
At1g32530 unknown protein 42 6e-04
At5g16730 putative protein 38 0.012
At4g25540 putative DNA mismatch repair protein 37 0.027
At5g41140 putative protein 34 0.23
At1g65010 hypothetical protein 34 0.23
At1g79280 hypothetical protein 33 0.30
At5g24880 glutamic acid-rich protein 33 0.39
At5g19180 putative ubiquitin activating enzyme E1 (ECR1) 32 0.87
At4g24910 putative protein 32 1.1
At2g05560 hypothetical protein 32 1.1
At1g63360 disease resistance protein, putative 31 1.5
At4g31530 unknown protein 31 1.9
At1g04730 hypothetical protein 31 1.9
At5g61190 putative protein 30 2.5
At2g35840 putative sucrose-6F-phosphate phosphohydrolase 30 2.5
At2g20180 putative bHLH transcription factor (bHLH015) 30 3.3
>At1g58290 putative protein
Length = 543
Score = 643 bits (1658), Expect = 0.0
Identities = 335/488 (68%), Positives = 399/488 (81%), Gaps = 5/488 (1%)
Query: 43 KCTLRSDNPLPQNFIVSKPSPLEILKTSSADRYTKEKSSIIVIGLNVHTAPVEMREKLAI 102
+C L + + N + S LE LK S+ADRYTKE+SSI+VIGL++HTAPVEMREKLAI
Sbjct: 60 RCELSASSDSASN--AASISALEQLKNSAADRYTKERSSIVVIGLSIHTAPVEMREKLAI 117
Query: 103 PEAQWPQVIQELCALNHIEEAAVLSTCNRIEIYLVALSQHRGVREVTDWISKKSGVSVPE 162
PEA+WP+ I ELC LNHIEEAAVLSTCNR+EIY++ALSQHRGV+EVT+W+SK SG+ V E
Sbjct: 118 PEAEWPRAIAELCGLNHIEEAAVLSTCNRMEIYVLALSQHRGVKEVTEWMSKTSGIPVSE 177
Query: 163 ISKHQILLYNKDATQHLFEVAAGLDSLVLGEGQILSQVKQVVKSGQGVPGFDRKISGLFK 222
I +H+ LLYNKDATQH+FEV+AGLDSLVLGEGQIL+QVKQVVK GQGV GF R ISGLFK
Sbjct: 178 ICQHRFLLYNKDATQHIFEVSAGLDSLVLGEGQILAQVKQVVKVGQGVNGFGRNISGLFK 237
Query: 223 QAISVGKRVRTETNISSGSVSVSSAAVELALMKRLESSFGDAKVLVIGAGKMGKLVIKHL 282
AI+VGKRVRTETNI+SG+VSVSSAAVELALMK +SS A++ VIGAGKMGKLVIKHL
Sbjct: 238 HAITVGKRVRTETNIASGAVSVSSAAVELALMKLPQSSNVSARMCVIGAGKMGKLVIKHL 297
Query: 283 VAKGCRKMVVVNRSEEKVNAIRKELKDVDIVYRPLSEMMECAAEADVIFTGTASESLLFS 342
+AKGC K+VVVNRSEE+V+AIR+E+ ++I+YRPL EM+ CA+EADV+FT TASE+ LF
Sbjct: 298 MAKGCTKVVVVNRSEERVSAIREEMPGIEIIYRPLDEMLACASEADVVFTSTASETPLFL 357
Query: 343 KENVEILPPVGQGV-RRRLFVDISIPRNVDPGVSELENALVYNVDDLREVVDANKEDRQR 401
KE+VE LP V R FVDIS+PRNV V E+E A VYNVDDL+EVV ANKEDR R
Sbjct: 358 KEHVENLPQASPEVGGLRHFVDISVPRNVGSCVGEVETARVYNVDDLKEVVAANKEDRMR 417
Query: 402 KAMEARGIIKEELNTFEAWKDSLETVPTIKKFRAYVERIRASELEKCLSKMRGDVSKEQK 461
KAMEA+ II EE FEAW+DSLETVPTIKK RAY ERIR +ELEKC+SKM D++K+
Sbjct: 418 KAMEAQTIITEESTQFEAWRDSLETVPTIKKLRAYAERIRVAELEKCMSKMGDDINKKTT 477
Query: 462 EAMYALSMGIVNKLLHGPMQHLRCDGNDTKCLDEVLENMRALNRMYDLETEISLMEEKIR 521
A+ LS GIVN+ LHGPMQHLRCDG+D++ L E LENM ALNRMY LE +I +EEK++
Sbjct: 478 RAVDDLSRGIVNRFLHGPMQHLRCDGSDSRTLSETLENMHALNRMYGLEKDI--LEEKLK 535
Query: 522 VKMEKAKK 529
E+ +K
Sbjct: 536 AMAEQQQK 543
>At1g09940 putative glutamyl-tRNA reductase 2 precursor
Length = 530
Score = 635 bits (1637), Expect = 0.0
Identities = 331/467 (70%), Positives = 390/467 (82%), Gaps = 7/467 (1%)
Query: 62 SPLEILKTSSADRYTKEKSSIIVIGLNVHTAPVEMREKLAIPEAQWPQVIQELCALNHIE 121
S LE LKTS+ DRYTKE+SSI+VIGL++HTAPVEMREKLAIPEA+WP+ I ELC LNHIE
Sbjct: 68 SALEQLKTSAIDRYTKERSSIVVIGLSIHTAPVEMREKLAIPEAEWPRAIAELCGLNHIE 127
Query: 122 EAAVLSTCNRIEIYLVALSQHRGVREVTDWISKKSGVSVPEISKHQILLYNKDATQHLFE 181
EAAVLSTCNR+EIY++ALSQHRGV+EVT+W+SK SG+ V EI +H+ LLYNKD TQH+FE
Sbjct: 128 EAAVLSTCNRMEIYVLALSQHRGVKEVTEWMSKTSGIPVSEICQHRFLLYNKDVTQHIFE 187
Query: 182 VAAGLDSLVLGEGQILSQVKQVVKSGQGVPGFDRKISGLFKQAISVGKRVRTETNISSGS 241
V+AGLDSLVLGEGQIL+QVKQVVK GQGV GF R ISGLFK AI+VGKRVRTETNI++G+
Sbjct: 188 VSAGLDSLVLGEGQILAQVKQVVKVGQGVNGFGRNISGLFKHAITVGKRVRTETNIAAGA 247
Query: 242 VSVSSAAVELALMKRLESSF-GDAKVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSEEKV 300
VSVSSAAVELALMK ESS A++LV+GAGKMGKLVIKHLVAKGC KMVVVNRSEEKV
Sbjct: 248 VSVSSAAVELALMKLPESSHASSARMLVVGAGKMGKLVIKHLVAKGCTKMVVVNRSEEKV 307
Query: 301 NAIRKEL-KDVDIVYRPLSEMMECAAEADVIFTGTASESLLFSKENVEILPPVGQGVRRR 359
A+R E+ V+I+Y+PL EM+ CAAEADV+FT TASE+ LF KE VE LPPV R
Sbjct: 308 AAVRNEMPPGVEIIYKPLDEMLSCAAEADVVFTSTASETPLFLKEQVETLPPVRDA---R 364
Query: 360 LFVDISIPRNVDPGVSELENALVYNVDDLREVVDANKEDRQRKAMEARGIIKEELNTFEA 419
LFVDIS+PRNV V+E++ V+NVDDL+EVV ANKEDR RKAM+A+ II +E FEA
Sbjct: 365 LFVDISVPRNVGSCVAEIDGTRVFNVDDLKEVVAANKEDRVRKAMDAQAIITDESKHFEA 424
Query: 420 WKDSLETVPTIKKFRAYVERIRASELEKCLSKMRGDVSKEQKEAMYALSMGIVNKLLHGP 479
W+DSLETVPTIKK R Y ERI A+E+EK L KM D++K+ ++ + L GIVNKLLHGP
Sbjct: 425 WRDSLETVPTIKKLRGYTERIIAAEIEKSLPKMGIDMNKKMRKTVDDLIRGIVNKLLHGP 484
Query: 480 MQHLRCDGNDTKCLDEVLENMRALNRMYDLETEISLMEEKIRVKMEK 526
MQHLRCDGND++ L E L+NM+ALNRMY L+ EI +EEKIR K+EK
Sbjct: 485 MQHLRCDGNDSRTLSETLDNMQALNRMYGLDAEI--LEEKIRAKVEK 529
>At2g31250 putative glutamyl tRNA reductase
Length = 524
Score = 536 bits (1380), Expect = e-152
Identities = 294/473 (62%), Positives = 370/473 (78%), Gaps = 13/473 (2%)
Query: 59 SKPSPLEILKTSSA-DRYTKEKSSIIVIGLNVHTAPVEMREKLAIPEAQWPQVIQELCAL 117
S SPL+ LK S A DRYT+E+SSI+VIGL+ HT+P+EMRE+LAIPEA+WP I +LCAL
Sbjct: 59 SDSSPLDKLKNSPAIDRYTRERSSIVVIGLSFHTSPLEMRERLAIPEAEWPLAITQLCAL 118
Query: 118 NHIEEAAVLSTCNRIEIYLVALSQHRGVREVTDWISKKSGVSVPEISKHQILLYNKDATQ 177
NHIEEAAVLSTCNRIEIY+ ALS++RGV+EVT+W+SK+SG+ V +I +H+ LLYNKDATQ
Sbjct: 119 NHIEEAAVLSTCNRIEIYVSALSRYRGVKEVTEWMSKRSGIPVSDICQHRFLLYNKDATQ 178
Query: 178 HLFEVAAGLDSLVLGEGQILSQVK---QVVKSGQGVPGFDRKISGLFKQAISVGKRVRTE 234
HLF+V+AGL+SLV+GE QI SQV+ QVVK GF R IS LF++A GKRVR +
Sbjct: 179 HLFQVSAGLESLVIGENQIQSQVRKAEQVVKQ----EGFGRIISTLFEKANKAGKRVRAQ 234
Query: 235 TNISSGSVSVSSAAVELALMKRLESSFGDAKVLVIGAGKMGKLVIKHLVAKGCRKMVVVN 294
TNI+SG+VSVSSAAVELAL K L S A +LVIGAG+MGK +I+HLVAKGC KMVV+N
Sbjct: 235 TNIASGAVSVSSAAVELALTK-LPGSVSSAMMLVIGAGEMGKRIIEHLVAKGCTKMVVMN 293
Query: 295 RSEEKVNAIRKELKD-VDIVYRPLSEMMECAAEADVIFTGTASESLLFSKENVEILPPVG 353
RSE+KV AIRKE++ V+I+Y+PL E++ CAAEA+VIFT T+SE+ LF KE+VEILPP
Sbjct: 294 RSEDKVAAIRKEMQSGVEIIYKPLDEILACAAEANVIFTSTSSETPLFLKEHVEILPPCP 353
Query: 354 QGVRRRLFVDISIPRNVDPGVSELENALVYNVDDLREVVDANKEDRQRKAMEARGIIKEE 413
RLFVDIS+PRNV V+EL++A VYNVDDL+EVV ANKEDR RK+MEA II+EE
Sbjct: 354 ADY-ARLFVDISVPRNVGSCVAELDSARVYNVDDLKEVVAANKEDRARKSMEALPIIREE 412
Query: 414 LNTFEAWKDSLETVPTIKKFRAYVERIRASELEKCLSKMRGDVSKEQKEAMYALSMGIVN 473
FE W+DSL+T PTI+K R+ ERIRA +EK +SK + K+ +EA+ + IVN
Sbjct: 413 TIEFEGWRDSLQTFPTIRKLRSKTERIRAECVEKLISKHGNGMDKKTREAVEKQTRIIVN 472
Query: 474 KLLHGPMQHLRCDGNDTKCLDEVLENMRALNRMYDLETEISLMEEKIRVKMEK 526
+L PM+HLR DG + L E LENM+A+NR+Y+L+ E L+EEKIR K +K
Sbjct: 473 NILDYPMKHLRYDGTGSSKLRETLENMQAVNRIYELDGE--LLEEKIREKKDK 523
>At2g35330 unknown protein
Length = 738
Score = 52.8 bits (125), Expect = 5e-07
Identities = 42/175 (24%), Positives = 79/175 (45%), Gaps = 8/175 (4%)
Query: 361 FVDISIPRNVDPGVSELEN----ALVYNVDDLREVVDANKEDRQRKAMEARGIIKEELNT 416
F D+++ NVD EL++ L+ V DL++ + K+ Q+KAM+A + +EL+
Sbjct: 377 FRDLNLDDNVDSAPEELKDDALIGLLQQVQDLKKQLKERKDWAQKKAMQAAQKVSDELSE 436
Query: 417 FEAWKDSLETVPTIKKFRAYVERI---RASELEKCLSKMRGDVSKEQKEAMYALSMGIVN 473
++ + E + +KK + E + SE+E L K G V K + AL
Sbjct: 437 LKSLRSEREEIQRVKKGKQTREDSTLKKLSEMENALRKASGQVDK-ANAVVRALENESAE 495
Query: 474 KLLHGPMQHLRCDGNDTKCLDEVLENMRALNRMYDLETEISLMEEKIRVKMEKAK 528
L + T C++ + + L ++ E + ++++I + EK K
Sbjct: 496 IRAEMEASKLSASESLTACMEASKKEKKCLKKLLAWEKQKMKLQDEITAEKEKIK 550
>At1g32530 unknown protein
Length = 711
Score = 42.4 bits (98), Expect = 6e-04
Identities = 35/174 (20%), Positives = 76/174 (43%), Gaps = 6/174 (3%)
Query: 361 FVDISIPRNVDP-GVSELENALV---YNVDDLREVVDANKEDRQRKAMEARGIIKEELNT 416
F D+++ N++ GV + + +V + V D + V KE Q+ AM+A + EEL
Sbjct: 350 FRDLNLDDNLESVGVDDKDCVIVDLLHQVKDFEKKVKERKEWAQKNAMQAAQKVSEELAE 409
Query: 417 FEAWKDSLETVPTIKKFRAYVERIRASELEKCLSKMRGDVSKEQKEAMYALSMGIVNKLL 476
+ E + +KK + VE A ++R S+ + + + N +
Sbjct: 410 LKTLSSEREGIQLLKKGKQAVEESTAKRFTDKEIELRKACSQNDRANVIVRKLENQNAEI 469
Query: 477 HGPMQHLRCDGNDT--KCLDEVLENMRALNRMYDLETEISLMEEKIRVKMEKAK 528
+ + +++ C++ + + L ++ E +I ++++I + EK K
Sbjct: 470 RAEREGSKLSASESLKACMEASKKEKKCLKKLVAWEKQILKLQDEITAEKEKIK 523
>At5g16730 putative protein
Length = 853
Score = 38.1 bits (87), Expect = 0.012
Identities = 95/488 (19%), Positives = 191/488 (38%), Gaps = 44/488 (9%)
Query: 13 LRSSVKFSPAQFPKSTFAPQRLSFTTPKIPKCTLRSDNPLPQNFIVSKPSPLEILKTSSA 72
+ S K S ++ +T + S TP+I K T+ N PS S
Sbjct: 1 MASKTKTSLSETTTTTTPTGKSSPATPRIAKRTVNKSETSNNN----SPSTTTPHSRLSL 56
Query: 73 DRYT-KEKSSIIVIGLNVHTAPVEMREKLA-IPEAQWPQVIQELCALNHIEEAAVLSTCN 130
DR + KSS+ + T P + + ++A + + PQ L + + A +
Sbjct: 57 DRSSPNSKSSVERRSPKLPTPPEKSQARVAAVKGTESPQTTTRLSQIKEDLKKANERISS 116
Query: 131 RIEIYLVALSQHRGVREVTDWISKKSGVSVPEISKHQILLYNKDATQHLFEVAAGLDSLV 190
+ AL + + ++ + ++ K + + K Q + + V AG++++
Sbjct: 117 LEKDKAKALDELKQAKKEAEQVTLK----LDDALKAQKHVEENSEIEKFQAVEAGIEAVQ 172
Query: 191 LGEGQILSQVKQVVKSGQG-------VPGFDRKISGLFKQAISVGKRVRTETNISSGSVS 243
E ++ +++ V V KI+ A + ++ +S +
Sbjct: 173 NNEEELKKELETVKNQHASDSAALVAVRQELEKINEELAAAFDAKSKALSQAEDASKTAE 232
Query: 244 VSSAAVELALMKRLESSFGDAKVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSEEKVNAI 303
+ + V++ L S K L+ + + +VAK ++VV+ R E
Sbjct: 233 IHAEKVDI-----LSSELTRLKALLDSTREKTAISDNEMVAKLEDEIVVLKRDLESARGF 287
Query: 304 RKELKDVDIVYRPLSEMMECAAEADVIFTGTASESLLFSKENVEILPPVGQGVRRRLFVD 363
E+K+ +++ L+ +E A A+ ++E +KE E L + R
Sbjct: 288 EAEVKEKEMIVEKLNVDLEAAKMAESNAHSLSNEWQSKAKELEEQLEEANKLERSASVSL 347
Query: 364 ISIPRNVDPGVSELENALVYNVDDLRE-------VVDANKEDRQRKAMEARGIIKEELN- 415
S+ + ++ +L + + DL+E V KED + + + G ++EE++
Sbjct: 348 ESVMKQLEGSNDKLHDTET-EITDLKERIVTLETTVAKQKEDLE-VSEQRLGSVEEEVSK 405
Query: 416 ---TFEAWKDSLETVP-----TIKKFRAYVERIR--ASELEKCLSKMRGDVSKEQ--KEA 463
E K LETV +KK + R++ + E K LS + +E+ K+A
Sbjct: 406 NEKEVEKLKSELETVKEEKNRALKKEQDATSRVQRLSEEKSKLLSDLESSKEEEEKSKKA 465
Query: 464 MYALSMGI 471
M +L+ +
Sbjct: 466 MESLASAL 473
>At4g25540 putative DNA mismatch repair protein
Length = 1076
Score = 37.0 bits (84), Expect = 0.027
Identities = 27/89 (30%), Positives = 47/89 (52%), Gaps = 11/89 (12%)
Query: 3 MAASSSTDHTLRSSVKFSPAQFPKSTFAPQRLSFTTPKIPKCTLRSDNPLP-----QNFI 57
+A SS+ + ++V FSP+ K L+ +PK PK + + NP+P Q F+
Sbjct: 27 VAESSTPPPKISATVSFSPS---KRKLLSDHLAAASPKKPKLSPHTQNPVPDPNLHQRFL 83
Query: 58 --VSKPSPLE-ILKTSSADRYTKEKSSII 83
+PSP E + +TSS+ +YT + ++
Sbjct: 84 QRFLEPSPEEYVPETSSSRKYTPLEQQVV 112
>At5g41140 putative protein
Length = 983
Score = 33.9 bits (76), Expect = 0.23
Identities = 43/196 (21%), Positives = 83/196 (41%), Gaps = 17/196 (8%)
Query: 332 TGTASESLLFSKENVEILPPVGQGVRRRLFVDISIPRNVDPGVSELEN-ALVYNVDD--- 387
T ++ L+ + +++E + +G R + VD+ PR + E + DD
Sbjct: 402 TQESNTELILAVQDLEAM----EGQRTKKTVDLPGPRTCERNTEESRRMSCTSETDDDED 457
Query: 388 ---LREVVDANKEDRQRKAMEARGIIKEELNTFEAWKDSLETVPTIKKFRAYVERIRASE 444
L E+V + + ++ +E R I + N E +K E + + + I E
Sbjct: 458 QKALDELVKGHMDAKEAHVLERR--ITDLYNEIEIYKRDKEDLEIQVEQLSLDYEILKQE 515
Query: 445 LEKCLSKMRGDVSKEQKEAMYALSMGIVNKLLHGPMQHLRCDGNDTKCLDEVLENMRALN 504
K+ +EQ + Y S +VN ++ H+ + + K + E +L
Sbjct: 516 NHDISYKLEQSQVQEQLKMQYECSSSLVN--VNELENHV--ESLEAKLKKQYKECSESLY 571
Query: 505 RMYDLETEISLMEEKI 520
R+ +LET+I MEE++
Sbjct: 572 RIKELETQIKGMEEEL 587
>At1g65010 hypothetical protein
Length = 1318
Score = 33.9 bits (76), Expect = 0.23
Identities = 75/377 (19%), Positives = 160/377 (41%), Gaps = 38/377 (10%)
Query: 174 DATQHLFEVAAGLDSLVLGEGQILSQVKQVVKSGQGVPGFDRKISGLFKQAISV-GKRVR 232
D + L V A + SL EG +L +++++ K + + + K+ + ++A + G+
Sbjct: 593 DKEEDLKNVTAEISSLREWEGSVLEKIEELSKVKESLVDKETKLQSITQEAEELKGREAA 652
Query: 233 TETNISSGSVSVSSAAVELALMKRLESSFGDAKVLVIG-AGKMGKLVIKH--LVAKGCRK 289
I S + +S E ++ + D K G K+ +L + + L
Sbjct: 653 HMKQIEELSTANASLVDEATKLQSIVQESEDLKEKEAGYLKKIEELSVANESLADNVTDL 712
Query: 290 MVVVNRSE---EKVNAIRKELKDVDIVYRPL----SEMMECAAEADVIFTGTAS------ 336
+V S+ E+ A K+++++ + L +++ EA+ + AS
Sbjct: 713 QSIVQESKDLKEREVAYLKKIEELSVANESLVDKETKLQHIDQEAEELRGREASHLKKIE 772
Query: 337 ------ESLLFSKENVEILPPVGQGVRRRLFVDISIPRNVDPGVSELENALVYNVDDLRE 390
E+L+ + N++ + + +R R +++ + +D +S L NV +L+
Sbjct: 773 ELSKENENLVDNVANMQNIAEESKDLRER---EVAYLKKIDE-LSTANGTLADNVTNLQN 828
Query: 391 VVDANKEDRQRKAMEARGIIKEELNTF-EAWKDSLETVPTIKKFRAYVERIRASELEKC- 448
+ + NKE R+R+ + EEL+ E+ D + T+ + + + L+K
Sbjct: 829 ISEENKELRERETTLLKK--AEELSELNESLVDKASKLQTVVQENEELRERETAYLKKIE 886
Query: 449 -LSKMRGDVSKEQKEAMYALSMGIVNKLLHGPMQHLRCDGNDTKCLDEVLENMRALNRMY 507
LSK+ +S +E +S +L +L+ +K +++L L+ M
Sbjct: 887 ELSKLHEILS--DQETKLQISNHEKEELKERETAYLKKIEELSKVQEDLLNKENELHGMV 944
Query: 508 ----DLETEISLMEEKI 520
DL ++ SL ++KI
Sbjct: 945 VEIEDLRSKDSLAQKKI 961
Score = 32.7 bits (73), Expect = 0.51
Identities = 70/326 (21%), Positives = 130/326 (39%), Gaps = 51/326 (15%)
Query: 251 LALMKRLESSFGDAKVLVIGAGKMG-----------KLVIKHLVAKGCRKMVVVNRSEEK 299
LA KR E SF K + + G K ++ + ++ + + + E+
Sbjct: 106 LAAQKRAEESFEVEKFRAVELEQAGLEAVQKKDVTSKNELESIRSQHALDISALLSTTEE 165
Query: 300 VNAIRKELK-DVDIVYRPLSEMMEC-------AAEADVIFTGTAS-ESLLFSKENVEILP 350
+ ++ EL D + LS E A +A+++ + ++LL SKE E +
Sbjct: 166 LQRVKHELSMTADAKNKALSHAEEATKIAEIHAEKAEILASELGRLKALLGSKEEKEAIE 225
Query: 351 PVGQGVRRRLFVDISIPRNVDPGVSELENALVYN---VDDLREVVDANK----------E 397
G + +L +I + R VS LE++L V+ L+ ++A K E
Sbjct: 226 --GNEIVSKLKSEIELLRGELEKVSILESSLKEQEGLVEQLKVDLEAAKMAESCTNSSVE 283
Query: 398 DRQRKAMEARGIIKEELNTFEAWKDSLETV------------PTIKKFRAYVERIRASEL 445
+ + K E ++E + + +S+E+V T A E+I L
Sbjct: 284 EWKNKVHELEKEVEESNRSKSSASESMESVMKQLAELNHVLHETKSDNAAQKEKIEL--L 341
Query: 446 EKCLSKMRGDVSKEQKEAMYALSMGI-VNKLLHGPMQHLRCDGND-TKCLDEVLENMRAL 503
EK + R D+ + ++ A + L+ L + T+ LD +
Sbjct: 342 EKTIEAQRTDLEEYGRQVCIAKEEASKLENLVESIKSELEISQEEKTRALDNEKAATSNI 401
Query: 504 NRMYDLETEISLMEEKIRVKMEKAKK 529
+ D TE+S+ E+ +V+ EK+KK
Sbjct: 402 QNLLDQRTELSIELERCKVEEEKSKK 427
>At1g79280 hypothetical protein
Length = 2111
Score = 33.5 bits (75), Expect = 0.30
Identities = 73/355 (20%), Positives = 145/355 (40%), Gaps = 51/355 (14%)
Query: 109 QVIQELCALNHIEEAAVLS---TCNRIEIYLVALSQH------RGVREVTDWISKKSGVS 159
++ EL ++ +AA ++ TC+ +E ++LSQ + + +D+ + + ++
Sbjct: 31 KIYAELDSVRAKADAASITAEQTCSLLEQKYLSLSQDFSSLESQNAKLQSDFDDRLAELA 90
Query: 160 VPEISKHQILLYNKDATQHLFEVAAGLDSLVLGEGQILSQVKQVVKSGQGVPGFDRKISG 219
+ KHQ+ L + + + ++ + L + Q++ ++Q D +IS
Sbjct: 91 QSQAQKHQLHLQSIEKDGEVERMSTEMSELHKSKRQLMELLEQK----------DAEISE 140
Query: 220 LFKQAIS-VGKRVRTETNISSGSVSVSSAAVELALMKRLESSFGDAKVLVIGAGKMGKLV 278
S + K V+ S ++ A ELA + + S K L + K +
Sbjct: 141 KNSTIKSYLDKIVKLTDTSSEKEARLAEATAELARSQAMCSRLSQEKELT---ERHAKWL 197
Query: 279 IKHLVAKGCRKMVVVNRSEEKVNAIRKELKDVDIVYRPLSEMMECAAEADVIFTGTASES 338
+ L AK + R + + + +L DV+ Y +EC S S
Sbjct: 198 DEELTAKVDSYAELRRRHSDLESEMSAKLVDVEKNY------IEC------------SSS 239
Query: 339 LLFSKENV-EILPPVGQGVRRRLFVDISIPRNVDPGVSELENALVYNVDDLREVVDANKE 397
L + KE + E+ +G L D+S ++ E A ++ + L ++ + E
Sbjct: 240 LNWHKERLRELETKIGS-----LQEDLSSCKDAATTTEEQYTAELFTANKLVDLYKESSE 294
Query: 398 DRQRKAMEARGIIK---EELNTFE-AWKDSLETVPTIKKFRAYVERIRASELEKC 448
+ RKA E G+IK L+ E ++K+ L+ + K+ +LEKC
Sbjct: 295 EWSRKAGELEGVIKALEARLSQVESSYKERLDKEVSTKQLLEKENGDLKQKLEKC 349
>At5g24880 glutamic acid-rich protein
Length = 443
Score = 33.1 bits (74), Expect = 0.39
Identities = 39/166 (23%), Positives = 67/166 (39%), Gaps = 21/166 (12%)
Query: 299 KVNAIRKELKDVDIVYRPLSEMMECAAEADVIFTGTASESLLFSKENVEILPPVGQGVRR 358
KV + ++ D DI P E E + V+ + E S + PVG
Sbjct: 207 KVESDHLQVSDHDIE-EPKYEKEEKEVQEKVVQANESVEEKAESSGPTPVASPVG----- 260
Query: 359 RLFVDISIPRNVDPGVSELENALVYNVDDLREVVDANKE--DRQRKAMEARGIIKEELNT 416
++ + V+ELE L+ N DD+ E + KE + Q E +K++++
Sbjct: 261 ---------KDCNAVVAELEEKLIKNEDDIEEKTEEMKEQDNNQANKSEEEEDVKKKIDE 311
Query: 417 FEAWKDSLETVPTIKKFRAYVERIRASELEKCLSKMRGDVSKEQKE 462
++ E V T K VE + E+ + + V +E+KE
Sbjct: 312 ----NETPEKVDTESKEVESVEETTQEKEEEVKEEGKERVEEEEKE 353
>At5g19180 putative ubiquitin activating enzyme E1 (ECR1)
Length = 454
Score = 32.0 bits (71), Expect = 0.87
Identities = 19/57 (33%), Positives = 33/57 (57%), Gaps = 1/57 (1%)
Query: 265 KVLVIGAGKMGKLVIKHLVAKGCRKMVVVNRSE-EKVNAIRKELKDVDIVYRPLSEM 320
++LVIGAG +G ++K L G R + V++ E N R+ L ++ V +P +E+
Sbjct: 48 RILVIGAGGLGCELLKDLALSGFRNLEVIDMDRIEVTNLNRQFLFRIEDVGKPKAEV 104
>At4g24910 putative protein
Length = 315
Score = 31.6 bits (70), Expect = 1.1
Identities = 38/166 (22%), Positives = 71/166 (41%), Gaps = 16/166 (9%)
Query: 13 LRSSVKFSPAQFPKSTFAPQRLSFTTPKIPKCTLRSDNPLPQNFIVSKPSPLEILKTSSA 72
L ++FSP P S P+ +T K+PK T+R PL F++S S L +L+ S
Sbjct: 13 LNPLLRFSP---PSSPDNPKHQRLSTIKMPKFTVRKLIPL-LIFVLSSLSVLRLLRISYK 68
Query: 73 DRYTKEKSSIIVIGLNVHTAPVEMREKLAIPEAQWPQVIQELCALNHIEEAAVLSTCNRI 132
+ SS L +P + ++L E + +EL L+ + S CN +
Sbjct: 69 SSSKSQSSSSTTFRL----SPADSSQQLRANEGPTALMEKELKLLS--DTVTRRSPCNIL 122
Query: 133 ------EIYLVALSQHRGVREVTDWISKKSGVSVPEISKHQILLYN 172
+ +++ RG+ + + K + E++ + +Y+
Sbjct: 123 VFGFAPQYLMLSSINTRGITVILEDEPAKIMIPKAEVNPNNTRIYS 168
>At2g05560 hypothetical protein
Length = 1471
Score = 31.6 bits (70), Expect = 1.1
Identities = 49/189 (25%), Positives = 76/189 (39%), Gaps = 40/189 (21%)
Query: 154 KKSGVSVPEISKHQILLYNKDATQHLFEVAAGLDSLVLGEGQILSQVKQVVKSGQ----- 208
KKS VP+ K Q++ DA H F+ VL Q++ V+++G
Sbjct: 719 KKSSPKVPKKVKKQLVYEQDDAHPHGFKAKT-----VLVPDVPTQQIEVVIRAGNVSYNP 773
Query: 209 -------GVPGFDRKISG------LFKQAISVGKRVRTETNI---SSGSVSVSSAAVELA 252
V F I G LF+ + S + + + +G +++ + L
Sbjct: 774 LDMVDPSKVEEFSNIIKGPSMIHTLFRVSQSAQRHLEMMAMLMCHKNGEQMIANRCIVLD 833
Query: 253 LMKR--LESSFGDAKVLVIGAG-KMGKLVIKHLVAKGCRKMVVVNRSEEKVNAIRKELKD 309
+M L GD K + G K GKL+ +A G V +NR K LKD
Sbjct: 834 MMLTHLLTKRVGDFKKCINKNGFKWGKLLSD--IANG----VHINREPNM-----KWLKD 882
Query: 310 VDIVYRPLS 318
VD+VY P++
Sbjct: 883 VDVVYAPMN 891
>At1g63360 disease resistance protein, putative
Length = 884
Score = 31.2 bits (69), Expect = 1.5
Identities = 23/82 (28%), Positives = 41/82 (49%), Gaps = 12/82 (14%)
Query: 351 PVGQGVRRRLFVDISIPRNVDPGVSELENALVYNVDDLREVVDANKEDRQR--KAMEARG 408
P V + L + +S N++ ++ LE + + + A ++D +R K EARG
Sbjct: 11 PCVNKVSQWLDMKVSYTHNLEKNLAALEKTM--------KELKAKRDDLERRLKREEARG 62
Query: 409 IIKEELNTFEAWKDSLETVPTI 430
+ + L+ F+ W DS+ TV I
Sbjct: 63 L--QRLSEFQVWLDSVATVEDI 82
>At4g31530 unknown protein
Length = 324
Score = 30.8 bits (68), Expect = 1.9
Identities = 19/76 (25%), Positives = 35/76 (46%)
Query: 231 VRTETNISSGSVSVSSAAVELALMKRLESSFGDAKVLVIGAGKMGKLVIKHLVAKGCRKM 290
VR E ++SS V + E L SS ++V G G +G+LV+ L+ + R
Sbjct: 42 VRNEASLSSSITPVQAFTEEAVDTSDLASSSSKLVLVVGGTGGVGQLVVASLLKRNIRSR 101
Query: 291 VVVNRSEEKVNAIRKE 306
+++ ++ K+
Sbjct: 102 LLLRDLDKATKLFGKQ 117
>At1g04730 hypothetical protein
Length = 954
Score = 30.8 bits (68), Expect = 1.9
Identities = 17/47 (36%), Positives = 29/47 (61%), Gaps = 2/47 (4%)
Query: 266 VLVIGAGKMGKLVIKHLVAKGC-RKMVVVNRSEEK-VNAIRKELKDV 310
+L+ GA +GK + H+ AK C ++V +N S+E+ +AI + DV
Sbjct: 348 LLLCGAPGLGKTTLAHIAAKHCGYRVVEINASDERSASAIETRILDV 394
>At5g61190 putative protein
Length = 976
Score = 30.4 bits (67), Expect = 2.5
Identities = 17/61 (27%), Positives = 32/61 (51%), Gaps = 4/61 (6%)
Query: 288 RKMVVVNRSEEKVNAIRKELKDVDIVYRPLSEMMECAAEADVIFTGTASESLLFSKENVE 347
+ +V+VN+SE +NA + E KD +P+ + E +++ T + F K+N E
Sbjct: 527 KHVVMVNQSEVLINASKLEEKDAQEKAQPIESIAEPQSQSQ----NTQEHTKFFEKQNEE 582
Query: 348 I 348
+
Sbjct: 583 L 583
>At2g35840 putative sucrose-6F-phosphate phosphohydrolase
Length = 422
Score = 30.4 bits (67), Expect = 2.5
Identities = 17/44 (38%), Positives = 24/44 (53%)
Query: 408 GIIKEELNTFEAWKDSLETVPTIKKFRAYVERIRASELEKCLSK 451
GI+K+E + F K ET K YVE+ +A E+ K LS+
Sbjct: 105 GIVKQEASNFPELKLQAETEQRPHKVSFYVEKSKAQEVTKELSQ 148
>At2g20180 putative bHLH transcription factor (bHLH015)
Length = 478
Score = 30.0 bits (66), Expect = 3.3
Identities = 29/110 (26%), Positives = 47/110 (42%), Gaps = 23/110 (20%)
Query: 395 NKEDRQRKAMEARGIIKEELNTFEAWKDSLETVPTIKKFRAYVERIRASELEKCLSKMRG 454
N +DR+RK EA + E + E + + T T +R RA+E+ + R
Sbjct: 246 NVDDRKRKEREATTTDETESRSEETKQARVSTTST--------KRSRAAEVHNLSERKRR 297
Query: 455 DVSKEQKEAMYALSMGIVNKLLHGPMQHLRCDGND-TKCLDEVLENMRAL 503
D E+ +A+ L RC+ +D LDE +E M++L
Sbjct: 298 DRINERMKALQELIP--------------RCNKSDKASMLDEAIEYMKSL 333
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.132 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,779,563
Number of Sequences: 26719
Number of extensions: 457616
Number of successful extensions: 1639
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 36
Number of HSP's that attempted gapping in prelim test: 1596
Number of HSP's gapped (non-prelim): 69
length of query: 529
length of database: 11,318,596
effective HSP length: 104
effective length of query: 425
effective length of database: 8,539,820
effective search space: 3629423500
effective search space used: 3629423500
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC124214.3


