

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.11 + phase: 0 /pseudo
(240 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
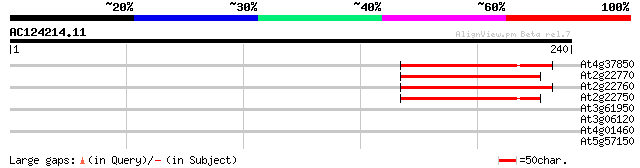
At4g37850 putative bHLH transcription factor (bHLH025) 66 2e-11
At2g22770 putative bHLH transcription factor (bHLH020) 61 5e-10
At2g22760 putative bHLH transcription factor (bHLH019) 60 1e-09
At2g22750 putative bHLH transcription factor (bHLH018) 59 2e-09
At3g61950 bHLH transcription factor (bHLH067) like protein 36 0.021
At3g06120 putative bHLH transcription factor (bHLH045) 30 1.1
At4g01460 putative bHLH transcription factor (bHLH057) 28 4.4
At5g57150 unknown protein 27 7.4
>At4g37850 putative bHLH transcription factor (bHLH025)
Length = 304
Score = 65.9 bits (159), Expect = 2e-11
Identities = 35/65 (53%), Positives = 46/65 (69%), Gaps = 1/65 (1%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIE RF D +VLI++ CEK KG + K + +IEKL++ +TNSS + FG LDITIIA+
Sbjct: 222 LPEIEVRFSDEDVLIKILCEKQKGHLAKIMAEIEKLHILITNSSVLNFGP-TLDITIIAK 280
Query: 228 PTLSF 232
F
Sbjct: 281 KESDF 285
>At2g22770 putative bHLH transcription factor (bHLH020)
Length = 320
Score = 61.2 bits (147), Expect = 5e-10
Identities = 30/60 (50%), Positives = 41/60 (68%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
+P IEAR DR++LIRVHCEK KG + K + +EK L+V NS + FG+ L ITI+ +
Sbjct: 239 MPMIEARVSDRDLLIRVHCEKNKGCMIKILSSLEKFRLEVVNSFTLPFGNSTLVITILTK 298
>At2g22760 putative bHLH transcription factor (bHLH019)
Length = 295
Score = 59.7 bits (143), Expect = 1e-09
Identities = 28/65 (43%), Positives = 40/65 (61%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIEA+ ++LIR+ CEK KG + ++ IE L++ NS + FG LDIT++AQ
Sbjct: 214 LPEIEAKISQNDILIRILCEKSKGCMINILNTIENFQLRIENSIVLPFGDSTLDITVLAQ 273
Query: 228 PTLSF 232
F
Sbjct: 274 MDKDF 278
>At2g22750 putative bHLH transcription factor (bHLH018)
Length = 304
Score = 58.9 bits (141), Expect = 2e-09
Identities = 32/60 (53%), Positives = 42/60 (69%), Gaps = 1/60 (1%)
Query: 168 LPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQ 227
LPEIE R ++VLI++ CEK KG V K + +IEKL L +TNS+ + FG DI+IIAQ
Sbjct: 223 LPEIEVRVSGKDVLIKILCEKQKGNVIKIMGEIEKLGLSITNSNVLPFGP-TFDISIIAQ 281
>At3g61950 bHLH transcription factor (bHLH067) like protein
Length = 358
Score = 35.8 bits (81), Expect = 0.021
Identities = 19/51 (37%), Positives = 29/51 (56%)
Query: 164 DYKRLPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMT 214
D +P+IEA +V ++V CEK +G + K I +EKL L V + + T
Sbjct: 269 DQTCIPKIEATVIQNHVSLKVQCEKKQGQLLKGIISLEKLKLTVLHLNITT 319
>At3g06120 putative bHLH transcription factor (bHLH045)
Length = 202
Score = 30.0 bits (66), Expect = 1.1
Identities = 19/64 (29%), Positives = 34/64 (52%), Gaps = 10/64 (15%)
Query: 171 IEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQPTL 230
+EA+ NV++RV +I G + K I +EKL+ +V + L+I+ + + L
Sbjct: 117 VEAKISGSNVVLRVVSRRIVGQLVKIISVLEKLSFQVLH----------LNISSMEETVL 166
Query: 231 SFYV 234
F+V
Sbjct: 167 YFFV 170
>At4g01460 putative bHLH transcription factor (bHLH057)
Length = 315
Score = 28.1 bits (61), Expect = 4.4
Identities = 19/69 (27%), Positives = 32/69 (45%), Gaps = 12/69 (17%)
Query: 170 EIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFGSCALDITIIAQPT 229
E+EA +V ++V C++ K + K I IE+L L + L +TI +
Sbjct: 223 EVEATVIQNHVSLKVRCKRGKRQILKAIVSIEELKLAI------------LHLTISSSFD 270
Query: 230 LSFYVFHLR 238
Y F+L+
Sbjct: 271 FVIYSFNLK 279
>At5g57150 unknown protein
Length = 226
Score = 27.3 bits (59), Expect = 7.4
Identities = 12/39 (30%), Positives = 20/39 (50%)
Query: 177 DRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTF 215
+R +++ V C K + K E LNLK+ S+ +F
Sbjct: 165 ERTMVVSVTCNKRTDTMVKLCEVFESLNLKILTSNLTSF 203
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.364 0.163 0.551
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,883,929
Number of Sequences: 26719
Number of extensions: 125124
Number of successful extensions: 889
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 881
Number of HSP's gapped (non-prelim): 8
length of query: 240
length of database: 11,318,596
effective HSP length: 96
effective length of query: 144
effective length of database: 8,753,572
effective search space: 1260514368
effective search space used: 1260514368
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 58 (26.9 bits)
Medicago: description of AC124214.11


