

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123898.1 - phase: 0 /pseudo
(177 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
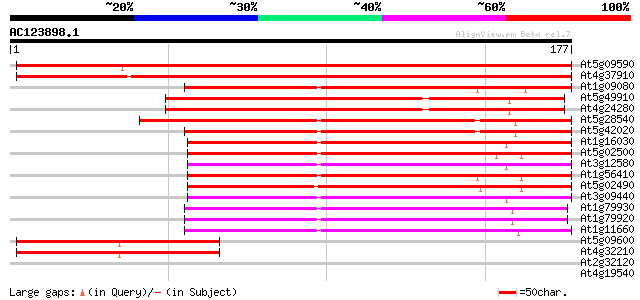
Score E
Sequences producing significant alignments: (bits) Value
At5g09590 heat shock protein 70 (Hsc70-5) 277 2e-75
At4g37910 heat shock protein 70 like protein 259 5e-70
At1g09080 putative luminal binding protein 121 2e-28
At5g49910 heat shock protein 70 (Hsc70-7) 120 5e-28
At4g24280 hsp 70-like protein 118 2e-27
At5g28540 luminal binding protein 114 2e-26
At5g42020 luminal binding protein (dbj|BAA13948.1) 114 3e-26
At1g16030 unknown protein 110 4e-25
At5g02500 dnaK-type molecular chaperone hsc70.1 108 2e-24
At3g12580 putative protein 106 7e-24
At1g56410 hypothetical protein 105 1e-23
At5g02490 dnaK-type molecular chaperone hsc70.1 - like 101 2e-22
At3g09440 heat-shock protein (At-hsc70-3) 100 3e-22
At1g79930 putative heat-shock protein 86 8e-18
At1g79920 putative heat-shock protein (At1g79920) 86 1e-17
At1g11660 heat-shock protein like 74 5e-14
At5g09600 putative protein 69 2e-12
At4g32210 putative protein 69 2e-12
At2g32120 70kD heat shock protein 37 0.007
At4g19540 ATP binding protein - like 31 0.40
>At5g09590 heat shock protein 70 (Hsc70-5)
Length = 682
Score = 277 bits (708), Expect = 2e-75
Identities = 138/179 (77%), Positives = 155/179 (86%), Gaps = 4/179 (2%)
Query: 3 AATALLRSLRRRDAASSTVTAFRSLTGNTKPA----YVGHNWSSLSRPFSSRAAGNDVIG 58
A ALLRS+RRR+ SS +A+R L+ + K + Y+G N+ S SR FSS+ AGNDVIG
Sbjct: 2 ATAALLRSIRRREVVSSPFSAYRCLSSSGKASLNSSYLGQNFRSFSRAFSSKPAGNDVIG 61
Query: 59 IDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTI 118
IDLGTTNSCV+VMEGKNPKV+EN+EGARTTPSVVAF KGELLVGTPAKRQAVTNP NT+
Sbjct: 62 IDLGTTNSCVAVMEGKNPKVIENAEGARTTPSVVAFNTKGELLVGTPAKRQAVTNPTNTV 121
Query: 119 SGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAKGQQYSPSQIGAFVLTKMKE 177
SG KRLIGR+FDDPQTQKEMKMVPYKIV+APNGDAWVEA GQQYSPSQIGAF+LTKMKE
Sbjct: 122 SGTKRLIGRKFDDPQTQKEMKMVPYKIVRAPNGDAWVEANGQQYSPSQIGAFILTKMKE 180
>At4g37910 heat shock protein 70 like protein
Length = 682
Score = 259 bits (662), Expect = 5e-70
Identities = 132/175 (75%), Positives = 149/175 (84%), Gaps = 1/175 (0%)
Query: 3 AATALLRSLRRRDAASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSSRAAGNDVIGIDLG 62
A+ ALLRS RRR+ ++V+AF+S++ N K + G L+RPF SR GNDVIGIDLG
Sbjct: 2 ASVALLRSFRRREVQMASVSAFKSVSANGKNSMFG-KLGYLARPFCSRPVGNDVIGIDLG 60
Query: 63 TTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAK 122
TTNSCVSVMEGK +V+EN+EG+RTTPSVVA QKGELLVGTPAKRQAVTNP NTI G+K
Sbjct: 61 TTNSCVSVMEGKTARVIENAEGSRTTPSVVAMNQKGELLVGTPAKRQAVTNPTNTIFGSK 120
Query: 123 RLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAKGQQYSPSQIGAFVLTKMKE 177
RLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEA GQ++SPSQIGA VLTKMKE
Sbjct: 121 RLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEANGQKFSPSQIGANVLTKMKE 175
>At1g09080 putative luminal binding protein
Length = 678
Score = 121 bits (303), Expect = 2e-28
Identities = 66/125 (52%), Positives = 84/125 (66%), Gaps = 4/125 (3%)
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
VIGIDLGTT SCV V K+ +++ N +G R TPS VAFT E L+G AK QA NPE
Sbjct: 52 VIGIDLGTTYSCVGVYHNKHVEIIANDQGNRITPSWVAFTDT-ERLIGEAAKNQAAKNPE 110
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIV-KAPNGDAWVEAKGQQ--YSPSQIGAFVL 172
TI KRLIGR+FDDP Q+++K +PYK+V K V+ KG++ +SP +I A +L
Sbjct: 111 RTIFDPKRLIGRKFDDPDVQRDIKFLPYKVVNKDGKPYIQVKVKGEEKLFSPEEISAMIL 170
Query: 173 TKMKE 177
TKMKE
Sbjct: 171 TKMKE 175
>At5g49910 heat shock protein 70 (Hsc70-7)
Length = 718
Score = 120 bits (300), Expect = 5e-28
Identities = 64/128 (50%), Positives = 85/128 (66%), Gaps = 4/128 (3%)
Query: 50 RAAGNDVIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQ 109
R V+GIDLGTTNS V+ MEG P +V N+EG RTTPSVVA+T+ + LVG AKRQ
Sbjct: 74 RVVNEKVVGIDLGTTNSAVAAMEGGKPTIVTNAEGQRTTPSVVAYTKSKDRLVGQIAKRQ 133
Query: 110 AVTNPENTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVE--AKGQQYSPSQI 167
AV NPENT KR IGRR + + +E K V Y+++K NG+ ++ A G+Q++ +I
Sbjct: 134 AVVNPENTFFSVKRFIGRRMN--EVAEESKQVSYRVIKDENGNVKLDCPAIGKQFAAEEI 191
Query: 168 GAFVLTKM 175
A VL K+
Sbjct: 192 SAQVLRKL 199
>At4g24280 hsp 70-like protein
Length = 718
Score = 118 bits (295), Expect = 2e-27
Identities = 63/128 (49%), Positives = 84/128 (65%), Gaps = 4/128 (3%)
Query: 50 RAAGNDVIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQ 109
R V+GIDLGTTNS V+ MEG P +V N+EG RTTPSVVA+T+ G+ LVG AKRQ
Sbjct: 74 RVVNEKVVGIDLGTTNSAVAAMEGGKPTIVTNAEGQRTTPSVVAYTKSGDRLVGQIAKRQ 133
Query: 110 AVTNPENTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVE--AKGQQYSPSQI 167
AV NPENT KR IGR+ + + +E K V Y++V+ N + +E A +Q++ +I
Sbjct: 134 AVVNPENTFFSVKRFIGRKMN--EVDEESKQVSYRVVRDENNNVKLECPAINKQFAAEEI 191
Query: 168 GAFVLTKM 175
A VL K+
Sbjct: 192 SAQVLRKL 199
>At5g28540 luminal binding protein
Length = 669
Score = 114 bits (286), Expect = 2e-26
Identities = 63/141 (44%), Positives = 86/141 (60%), Gaps = 7/141 (4%)
Query: 42 SLSRPFSSRAAGNDVIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELL 101
+LS VIGIDLGTT SCV V + + +++ N +G R TPS V FT E L
Sbjct: 23 ALSSAIEEATKLGSVIGIDLGTTYSCVGVYKNGHVEIIANDQGNRITPSWVGFTDS-ERL 81
Query: 102 VGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK--- 158
+G AK QA NPE T+ KRLIGR+F+D + QK+ K+VPY+IV +G +++ K
Sbjct: 82 IGEAAKNQAAVNPERTVFDVKRLIGRKFEDKEVQKDRKLVPYQIVN-KDGKPYIQVKIKD 140
Query: 159 --GQQYSPSQIGAFVLTKMKE 177
+ +SP +I A +LTKMKE
Sbjct: 141 GETKVFSPEEISAMILTKMKE 161
>At5g42020 luminal binding protein (dbj|BAA13948.1)
Length = 668
Score = 114 bits (285), Expect = 3e-26
Identities = 61/127 (48%), Positives = 83/127 (65%), Gaps = 7/127 (5%)
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
VIGIDLGTT SCV V + + +++ N +G R TPS V FT E L+G AK QA NPE
Sbjct: 37 VIGIDLGTTYSCVGVYKNGHVEIIANDQGNRITPSWVGFTDS-ERLIGEAAKNQAAVNPE 95
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK-----GQQYSPSQIGAF 170
T+ KRLIGR+F+D + QK+ K+VPY+IV +G +++ K + +SP +I A
Sbjct: 96 RTVFDVKRLIGRKFEDKEVQKDRKLVPYQIVN-KDGKPYIQVKIKDGETKVFSPEEISAM 154
Query: 171 VLTKMKE 177
+LTKMKE
Sbjct: 155 ILTKMKE 161
>At1g16030 unknown protein
Length = 646
Score = 110 bits (275), Expect = 4e-25
Identities = 58/125 (46%), Positives = 77/125 (61%), Gaps = 5/125 (4%)
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V +++ N +G RTTPS VAFT E L+G AK Q NP+N
Sbjct: 9 IGIDLGTTYSCVGVWMNDRVEIIPNDQGNRTTPSYVAFTDT-ERLIGDAAKNQVALNPQN 67
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV----EAKGQQYSPSQIGAFVL 172
T+ AKRLIGR+F DP Q ++ P+K+V P + + + +Q+SP +I + VL
Sbjct: 68 TVFDAKRLIGRKFSDPSVQSDILHWPFKVVSGPGEKPMIVVSYKNEEKQFSPEEISSMVL 127
Query: 173 TKMKE 177
KMKE
Sbjct: 128 VKMKE 132
>At5g02500 dnaK-type molecular chaperone hsc70.1
Length = 651
Score = 108 bits (269), Expect = 2e-24
Identities = 60/125 (48%), Positives = 78/125 (62%), Gaps = 5/125 (4%)
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 10 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDS-ERLIGDAAKNQVAMNPVN 68
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGD--AWVEAKGQ--QYSPSQIGAFVL 172
T+ AKRLIGRRF D Q +MK+ P+KI P +VE KG+ +++ +I + VL
Sbjct: 69 TVFDAKRLIGRRFSDSSVQSDMKLWPFKIQAGPADKPMIYVEYKGEEKEFAAEEISSMVL 128
Query: 173 TKMKE 177
KM+E
Sbjct: 129 IKMRE 133
>At3g12580 putative protein
Length = 650
Score = 106 bits (264), Expect = 7e-24
Identities = 56/125 (44%), Positives = 75/125 (59%), Gaps = 5/125 (4%)
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 10 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDS-ERLIGDAAKNQVAMNPTN 68
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV----EAKGQQYSPSQIGAFVL 172
T+ AKRLIGRR+ DP Q + P+K+V P + + + +Q+S +I + VL
Sbjct: 69 TVFDAKRLIGRRYSDPSVQADKSHWPFKVVSGPGEKPMIVVNHKGEEKQFSAEEISSMVL 128
Query: 173 TKMKE 177
KM+E
Sbjct: 129 IKMRE 133
>At1g56410 hypothetical protein
Length = 617
Score = 105 bits (262), Expect = 1e-23
Identities = 59/125 (47%), Positives = 77/125 (61%), Gaps = 5/125 (4%)
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 10 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDS-ERLIGDAAKNQVAMNPVN 68
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIV--KAPNGDAWVEAKGQ--QYSPSQIGAFVL 172
T+ AKRLIGRRF D Q +MK P+K+ +A +V KG+ Q++ +I + VL
Sbjct: 69 TVFDAKRLIGRRFSDASVQSDMKFWPFKVTPGQADKPMIFVNYKGEEKQFAAEEISSMVL 128
Query: 173 TKMKE 177
KM+E
Sbjct: 129 IKMRE 133
>At5g02490 dnaK-type molecular chaperone hsc70.1 - like
Length = 653
Score = 101 bits (252), Expect = 2e-22
Identities = 58/125 (46%), Positives = 76/125 (60%), Gaps = 5/125 (4%)
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 10 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTD-SERLIGDAAKNQVAMNPVN 68
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVK--APNGDAWVEAKGQ--QYSPSQIGAFVL 172
T+ AKRLIGRRF D Q + ++ P+ I+ A VE KG+ Q++ +I + VL
Sbjct: 69 TVFDAKRLIGRRFSDASVQSDRQLWPFTIISGTAEKPMIVVEYKGEEKQFAAEEISSMVL 128
Query: 173 TKMKE 177
KM+E
Sbjct: 129 IKMRE 133
>At3g09440 heat-shock protein (At-hsc70-3)
Length = 649
Score = 100 bits (250), Expect = 3e-22
Identities = 53/125 (42%), Positives = 75/125 (59%), Gaps = 5/125 (4%)
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 10 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDS-ERLIGDAAKNQVAMNPIN 68
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV----EAKGQQYSPSQIGAFVL 172
T+ AKRLIGRRF D Q ++K+ P+ + P + + + +++S +I + +L
Sbjct: 69 TVFDAKRLIGRRFTDSSVQSDIKLWPFTLKSGPAEKPMIVVNYKGEDKEFSAEEISSMIL 128
Query: 173 TKMKE 177
KM+E
Sbjct: 129 IKMRE 133
>At1g79930 putative heat-shock protein
Length = 831
Score = 86.3 bits (212), Expect = 8e-18
Identities = 43/125 (34%), Positives = 73/125 (58%), Gaps = 5/125 (4%)
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
V+G D G N V+V + VV N E R TP++V F K + +GT + NP+
Sbjct: 3 VVGFDFGNENCLVAVARQRGIDVVLNDESNRETPAIVCFGDK-QRFIGTAGAASTMMNPK 61
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEA----KGQQYSPSQIGAFV 171
N+IS KRLIGR+F DP+ Q+++K +P+ + + P+G + A + + ++P+Q+ +
Sbjct: 62 NSISQIKRLIGRQFSDPELQRDIKSLPFSVTEGPDGYPLIHANYLGEKRAFTPTQVMGMM 121
Query: 172 LTKMK 176
L+ +K
Sbjct: 122 LSNLK 126
>At1g79920 putative heat-shock protein (At1g79920)
Length = 736
Score = 85.5 bits (210), Expect = 1e-17
Identities = 43/125 (34%), Positives = 73/125 (58%), Gaps = 5/125 (4%)
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
V+G D G N V+V + VV N E R TP++V F K + +GT + NP+
Sbjct: 3 VVGFDFGNENCLVAVARQRGIDVVLNDESNRETPAIVCFGDK-QRFIGTAGAASTMMNPK 61
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEA----KGQQYSPSQIGAFV 171
N+IS KRLIGR+F DP+ Q+++K +P+ + + P+G + A + + ++P+Q+ +
Sbjct: 62 NSISQIKRLIGRQFSDPELQRDIKSLPFSVTEGPDGYPLIHANYLGEIRAFTPTQVMGMM 121
Query: 172 LTKMK 176
L+ +K
Sbjct: 122 LSNLK 126
>At1g11660 heat-shock protein like
Length = 773
Score = 73.6 bits (179), Expect = 5e-14
Identities = 39/126 (30%), Positives = 72/126 (56%), Gaps = 5/126 (3%)
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
V+G D+G N ++V + + V+ N E R P++V+F +K + +G A A +P+
Sbjct: 3 VVGFDVGNENCVIAVAKQRGIDVLLNDESNRENPAMVSFGEK-QRFMGAAAAASATMHPK 61
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAKG----QQYSPSQIGAFV 171
+TIS KRLIGR+F +P Q ++++ P++ + +G + + Q +SP QI +
Sbjct: 62 STISQLKRLIGRKFREPDVQNDLRLFPFETSEDSDGGIQIRLRYMGEIQSFSPVQILGML 121
Query: 172 LTKMKE 177
L+ +K+
Sbjct: 122 LSHLKQ 127
>At5g09600 putative protein
Length = 213
Score = 68.6 bits (166), Expect = 2e-12
Identities = 32/68 (47%), Positives = 47/68 (69%), Gaps = 4/68 (5%)
Query: 3 AATALLRSLRRRDAASSTVTAFRSLTGNTKP----AYVGHNWSSLSRPFSSRAAGNDVIG 58
AATAL RS+RRRD S+ ++ ++SL GN +P +Y+G N++S SR F S+ ND++G
Sbjct: 2 AATALFRSIRRRDVVSAPLSVYKSLAGNAQPSWGSSYIGQNYASFSRAFGSKPVVNDILG 61
Query: 59 IDLGTTNS 66
LGT N+
Sbjct: 62 TGLGTNNA 69
>At4g32210 putative protein
Length = 1086
Score = 68.6 bits (166), Expect = 2e-12
Identities = 32/68 (47%), Positives = 47/68 (69%), Gaps = 4/68 (5%)
Query: 3 AATALLRSLRRRDAASSTVTAFRSLTGNTKP----AYVGHNWSSLSRPFSSRAAGNDVIG 58
AATAL RS+RRRD S+ ++ ++SL GN +P +Y+G N++S SR F S+ ND++G
Sbjct: 875 AATALFRSIRRRDVVSAPLSVYKSLAGNAQPSWGSSYIGQNYASFSRAFGSKPVVNDILG 934
Query: 59 IDLGTTNS 66
LGT N+
Sbjct: 935 TGLGTNNA 942
>At2g32120 70kD heat shock protein
Length = 563
Score = 36.6 bits (83), Expect = 0.007
Identities = 34/139 (24%), Positives = 62/139 (44%), Gaps = 15/139 (10%)
Query: 48 SSRAAGNDVIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAK 107
SS + +GID+GT+ ++V G ++ N+ + S V F K E+ G +
Sbjct: 22 SSPSLPEIALGIDIGTSQCSIAVWNGSQVHILRNTRNQKLIKSFVTF--KDEVPAGGVSN 79
Query: 108 RQAVTNPENTISGA-----KRLIGRRFDDPQTQKEMKMVPYKIVKAPNG-----DAWVEA 157
+ A + + ++GA KRL+GR DP K +P+ + G A V
Sbjct: 80 QLA--HEQEMLTGAAIFNMKRLVGRVDTDPVVHAS-KNLPFLVQTLDIGVRPFIAALVNN 136
Query: 158 KGQQYSPSQIGAFVLTKMK 176
+ +P ++ A L +++
Sbjct: 137 AWRSTTPEEVLAIFLVELR 155
>At4g19540 ATP binding protein - like
Length = 313
Score = 30.8 bits (68), Expect = 0.40
Identities = 34/141 (24%), Positives = 54/141 (38%), Gaps = 35/141 (24%)
Query: 3 AATALLRSLRRRDAASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSSRAAGNDVIGIDLG 62
A ALLRSLRRR+ ++ ++A++ FSS +AG + L
Sbjct: 2 ATVALLRSLRRRELHAAHISAYK---------------------FSSASAGGRTTELRLH 40
Query: 63 TTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAK 122
++V GK + + VA K EL +G + + G
Sbjct: 41 GVKDIIAVASGKGGV----GKSSTAVNLAVALANKCELKIGL---------LDADVYGPS 87
Query: 123 RLIGRRFDD-PQTQKEMKMVP 142
I + PQ ++MKM+P
Sbjct: 88 VPIMMNINQKPQVNQDMKMIP 108
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.127 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,815,392
Number of Sequences: 26719
Number of extensions: 149902
Number of successful extensions: 323
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 277
Number of HSP's gapped (non-prelim): 29
length of query: 177
length of database: 11,318,596
effective HSP length: 93
effective length of query: 84
effective length of database: 8,833,729
effective search space: 742033236
effective search space used: 742033236
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC123898.1


