

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123575.7 - phase: 0 /pseudo
(378 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
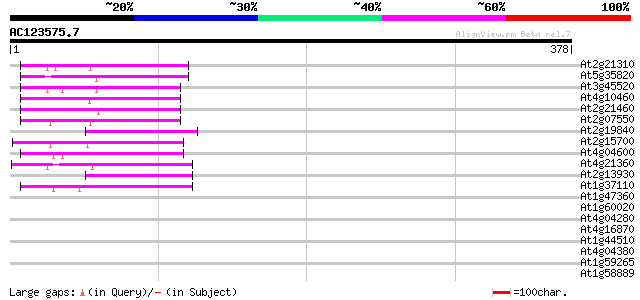
Score E
Sequences producing significant alignments: (bits) Value
At2g21310 putative retroelement pol polyprotein 67 2e-11
At5g35820 copia-like retrotransposable element 64 1e-10
At3g45520 copia-like polyprotein 62 5e-10
At4g10460 putative retrotransposon 62 7e-10
At2g21460 putative retroelement pol polyprotein 60 2e-09
At2g07550 putative retroelement pol polyprotein 57 2e-08
At2g19840 copia-like retroelement pol polyprotein 57 2e-08
At2g15700 copia-like retroelement pol polyprotein 56 4e-08
At4g04600 putative polyprotein 51 1e-06
At4g21360 putative transposable element 47 1e-05
At2g13930 putative retroelement pol polyprotein 46 4e-05
At1g37110 44 2e-04
At1g47360 polyprotein, putative 41 0.001
At1g60020 hypothetical protein 39 0.006
At4g04280 putative transposon protein 37 0.023
At4g16870 retrotransposon like protein 33 0.20
At1g44510 polyprotein, putative 33 0.26
At4g04380 putative polyprotein 32 0.44
At1g59265 polyprotein, putative 31 0.98
At1g58889 polyprotein, putative 31 0.98
>At2g21310 putative retroelement pol polyprotein
Length = 838
Score = 66.6 bits (161), Expect = 2e-11
Identities = 45/125 (36%), Positives = 65/125 (52%), Gaps = 12/125 (9%)
Query: 8 IEKFTGSNDFGLWKVKM--RIILI-------QQKCVEALKGKAQMPAHLTPAEKT---EM 55
+EK G D+ LWK K+ I L+ + + +E + A+ + LT E E
Sbjct: 8 VEKLDGEGDYVLWKEKLLAHIELLGLLEGLEEDEAIEEEESTAETDSLLTKTEDKVLKEK 67
Query: 56 NDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVES 115
KA S ++L LG+ VLR+V +E M LD L+M KS +R LKQ+LY Y+M +S
Sbjct: 68 RGKARSTVILSLGNHVLRKVIKEKTAAGMIRVLDKLFMAKSLPNRIYLKQRLYGYKMSDS 127
Query: 116 KTNNE 120
T E
Sbjct: 128 MTIEE 132
>At5g35820 copia-like retrotransposable element
Length = 1342
Score = 63.9 bits (154), Expect = 1e-10
Identities = 44/136 (32%), Positives = 64/136 (46%), Gaps = 26/136 (19%)
Query: 8 IEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMND---------- 57
+EKF G D+ LWK K+ L + + L+G + + TE++D
Sbjct: 8 VEKFDGDGDYILWKEKL---LAHMEMLGLLEGLGEEEEAVVEDSTTEISDGGNQDPETAT 64
Query: 58 -------------KAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLK 104
KA S I+L LG+ VLR+V ++ M LD L+M KS +R LK
Sbjct: 65 SKLEDKILKEKRGKARSTIILSLGNNVLRKVIKQKTAAGMIKVLDQLFMAKSLPNRIYLK 124
Query: 105 QQLYFYRMVESKTNNE 120
Q+LY Y+M E+ T E
Sbjct: 125 QRLYGYKMSENMTMEE 140
>At3g45520 copia-like polyprotein
Length = 1363
Score = 62.0 bits (149), Expect = 5e-10
Identities = 43/123 (34%), Positives = 63/123 (50%), Gaps = 15/123 (12%)
Query: 8 IEKFTGSNDFGLWKVKM--RIILIQQKCV----EALKGKAQMPAHLTPAEKTEMND---- 57
+EKF G D+ +WK K+ I ++ V E GK + EK E
Sbjct: 8 VEKFDGRGDYTMWKEKLLAHIDMLGLSAVLRESETPMGKERDSEKSDEDEKEEREKMEAF 67
Query: 58 -----KAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRM 112
KA S IVL + D+VLR++ +ET +M LD LYM+K+ +R LKQ+LY ++M
Sbjct: 68 EEKKRKARSTIVLSVSDRVLRKIKKETSAAAMLEALDRLYMSKALPNRIYLKQKLYSFKM 127
Query: 113 VES 115
E+
Sbjct: 128 SEN 130
>At4g10460 putative retrotransposon
Length = 1230
Score = 61.6 bits (148), Expect = 7e-10
Identities = 42/121 (34%), Positives = 59/121 (48%), Gaps = 13/121 (10%)
Query: 8 IEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEK-------------TE 54
+EKF G D+ LWK K+ + AL+ + L E+ E
Sbjct: 8 MEKFDGHGDYTLWKEKLMAHMDLLGLTVALRETQSVSDPLESEEEGKESEKGDKEALMEE 67
Query: 55 MNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVE 114
KA S IVL + D+VLR+ +E SM LD LYM+K+ +R LKQ+LY Y+M E
Sbjct: 68 KRQKARSTIVLSVSDQVLRKSKKEKTAPSMLEALDKLYMSKALPNRIYLKQKLYSYKMQE 127
Query: 115 S 115
+
Sbjct: 128 N 128
>At2g21460 putative retroelement pol polyprotein
Length = 1333
Score = 60.5 bits (145), Expect = 2e-09
Identities = 40/123 (32%), Positives = 62/123 (49%), Gaps = 15/123 (12%)
Query: 8 IEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDK--------- 58
+EKF G D+ +WK K+ L ALK + + + + TE +K
Sbjct: 8 VEKFDGRGDYTMWKEKLMAHLDILGLSVALKEEDDLVEKVAEMQLTEEEEKEEVLRRELL 67
Query: 59 ------AISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRM 112
A SAIVL + D+VLR++ +E +M LD LYM+K+ +R KQ+LY ++M
Sbjct: 68 EEKRRKARSAIVLSVTDRVLRKIKKEQSAAAMLGVLDKLYMSKALPNRIYQKQKLYSFKM 127
Query: 113 VES 115
E+
Sbjct: 128 SEN 130
>At2g07550 putative retroelement pol polyprotein
Length = 1356
Score = 57.0 bits (136), Expect = 2e-08
Identities = 38/123 (30%), Positives = 64/123 (51%), Gaps = 15/123 (12%)
Query: 8 IEKFTGSNDFGLWKVKMRI---ILIQQKCVEALKGKAQMPAHLTPAEKT----------- 53
+EKF G D+ +WK K+ IL ++ + + + L +++
Sbjct: 8 VEKFDGRGDYTMWKEKLLAHMDILGLNTALKESESTGEKKSVLDESDEDYEEKLEKFEAL 67
Query: 54 -EMNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRM 112
E KA SAIVL + D+VLR++ +E+ +M LD LYM+K+ +R KQ+LY ++M
Sbjct: 68 EEKKKKARSAIVLSVTDRVLRKIKKESTAAAMLLALDKLYMSKALPNRIYPKQKLYSFKM 127
Query: 113 VES 115
E+
Sbjct: 128 SEN 130
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 56.6 bits (135), Expect = 2e-08
Identities = 28/75 (37%), Positives = 42/75 (55%)
Query: 52 KTEMNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYR 111
+ + ++KA+ I + +GDKVLR + W LD LY+ KS +R L+ ++Y YR
Sbjct: 37 RIDQDEKAMDMIFINVGDKVLRNIENSKTAAEAWATLDKLYLVKSLPNRVYLQLKVYNYR 96
Query: 112 MVESKTNNEATDGIQ 126
M +SKT E D Q
Sbjct: 97 MQDSKTLEENVDEFQ 111
>At2g15700 copia-like retroelement pol polyprotein
Length = 1166
Score = 55.8 bits (133), Expect = 4e-08
Identities = 38/138 (27%), Positives = 65/138 (46%), Gaps = 23/138 (16%)
Query: 3 DSKWNIEKFTGSNDFGLWKVKMRI---------------ILIQQKCVEALKGKAQMPAHL 47
+++ +++F G+ DF LWKV+M +L +E A H+
Sbjct: 4 NARVEVDRFDGTGDFSLWKVRMLAHYGVLGLKGILNDEQLLRDPPVIEEEAAVAGRDYHV 63
Query: 48 TPAE--------KTEMNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTH 99
E K E ++KA IVL +G++VLR++ +MW+ L LYM S +
Sbjct: 64 GDFELPSNVDLIKFEKSEKAKDLIVLNVGNQVLRKIKNCETAAAMWSTLKRLYMETSLPN 123
Query: 100 RQCLKQQLYFYRMVESKT 117
R L+ + Y Y+M +S++
Sbjct: 124 RIYLQLKFYTYKMTDSRS 141
>At4g04600 putative polyprotein
Length = 922
Score = 50.8 bits (120), Expect = 1e-06
Identities = 37/136 (27%), Positives = 61/136 (44%), Gaps = 26/136 (19%)
Query: 8 IEKFTGSNDFGLWKVKMRIIL----------------------IQQKCV----EALKGKA 41
I+ F G NDF LWK+++ L ++K V E+ A
Sbjct: 9 IKAFDGDNDFSLWKMRIMAQLGVLGLKGTLTDFALTKTEPLTKSEEKQVASGDESSDSSA 68
Query: 42 QMPAHLTPAEKTEMNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQ 101
+ + K E +++AI+ I+ + D VLR+V ++W L+ LYM S +R
Sbjct: 69 VLTKEVPDPIKIEKSEQAINIIINHISDTVLRKVNHYKTAATLWELLNELYMETSLPNRI 128
Query: 102 CLKQQLYFYRMVESKT 117
+ + Y +RM+ SKT
Sbjct: 129 YAQLKFYSFRMMTSKT 144
>At4g21360 putative transposable element
Length = 1308
Score = 47.4 bits (111), Expect = 1e-05
Identities = 37/150 (24%), Positives = 70/150 (46%), Gaps = 32/150 (21%)
Query: 2 MDSKWNIEKFTGSNDFGLWKVKM-------------------RIILIQQKCVEALKGKAQ 42
M +K I+ F G DF LWK+++ + IL+ V++ K +++
Sbjct: 3 MSTKVEIKTFNGDRDFSLWKIRIEAQLGVLGLKPALSDFTLTKTILV----VKSEKKESE 58
Query: 43 MPAHLTPAEKTE---------MNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYM 93
T ++KTE +D+A + I+ + D VL +V +W L+ L+M
Sbjct: 59 SEDDETDSKKTEEVPDPIKFEQSDQAKNFIINHITDTVLLKVQHCVTAAELWATLNKLFM 118
Query: 94 TKSWTHRQCLKQQLYFYRMVESKTNNEATD 123
S +R + +LY ++MV++ + ++ TD
Sbjct: 119 ETSLPNRIYTQLRLYSFKMVDNLSIDQNTD 148
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 45.8 bits (107), Expect = 4e-05
Identities = 25/72 (34%), Positives = 37/72 (50%)
Query: 52 KTEMNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYR 111
+ E DKA + I L + DKVLR++ W LD L+M +S HR + Y ++
Sbjct: 42 RLERCDKAKNVIFLNVADKVLRKIELCKTAAEAWETLDRLFMIRSLPHRVYTQLSFYTFK 101
Query: 112 MVESKTNNEATD 123
M E+K +E D
Sbjct: 102 MQENKKIDENID 113
>At1g37110
Length = 1356
Score = 43.5 bits (101), Expect = 2e-04
Identities = 33/137 (24%), Positives = 59/137 (42%), Gaps = 21/137 (15%)
Query: 8 IEKFTGSNDFGLWKVKMRIIL------------IQQKCVEALKGKAQMPA---------H 46
I+ F G DF LWK++++ L K V K +A+ +
Sbjct: 10 IKVFNGDRDFSLWKIRIQAQLGVLGLKDTLTDFSLTKTVPLTKSEAKQESGDGESSGTKE 69
Query: 47 LTPAEKTEMNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQ 106
+ K E +++A + I+ + D VL +V +W L+ YM S +R + +
Sbjct: 70 VPDPVKIEQSEQAKNIIINHISDVVLLKVNHYATTADLWATLNKKYMETSLPNRIYTQLK 129
Query: 107 LYFYRMVESKTNNEATD 123
LY ++MV + T ++ D
Sbjct: 130 LYSFKMVSTMTIDQNVD 146
>At1g47360 polyprotein, putative
Length = 1182
Score = 40.8 bits (94), Expect = 0.001
Identities = 35/116 (30%), Positives = 50/116 (42%), Gaps = 18/116 (15%)
Query: 8 IEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAIVLCL 67
I F S DF LWK ++ L + + GK P LT E+ + K I LC
Sbjct: 6 IAVFDESGDFSLWKTRIMAHLSVIGLKDVVIGKTITP--LTAEEEEDPEKK----IELC- 58
Query: 68 GDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVESKTNNEATD 123
+ A+E W LD L+M +S HR + Y ++M E+K +E D
Sbjct: 59 ------QTAQEA-----WETLDRLFMIRSLPHRVYTQLSFYTFKMQENKKIDENID 103
>At1g60020 hypothetical protein
Length = 1194
Score = 38.5 bits (88), Expect = 0.006
Identities = 30/112 (26%), Positives = 50/112 (43%), Gaps = 12/112 (10%)
Query: 7 NIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHL---------TPAEKT-EMN 56
NI K T +N + +W ++ +L + G P+ + PA K +
Sbjct: 27 NITKLTPTN-YIMWNRQVHALLDGYDLAGYIDGSVTAPSEMITTAGVSAANPAYKFWKRQ 85
Query: 57 DKAI-SAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQL 107
DK I SAI+ + + ++R +W KL S+Y T SW H Q ++Q +
Sbjct: 86 DKLIYSAILGTITTTIQPLLSRSNTAAEIWEKLKSIYATPSWGHIQQMRQHI 137
>At4g04280 putative transposon protein
Length = 1104
Score = 36.6 bits (83), Expect = 0.023
Identities = 32/143 (22%), Positives = 59/143 (40%), Gaps = 24/143 (16%)
Query: 5 KWNIEKFTGSNDFGLWKVKMRIIL------------IQQKCVEALKGKAQMPAHLTPAE- 51
K I+ F G DF WK+++ L K V K + + + A
Sbjct: 2 KVEIKTFNGDRDFSFWKIRIEAQLGVLGLKNSLTDFKLTKTVPVAKKEEKESEYEDDASD 61
Query: 52 -----------KTEMNDKAISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHR 100
K E +++A + I+ + D VL +V +W L+ L+M S +R
Sbjct: 62 IKQATEEPDPIKFEQSEQAKNFIINHITDTVLLKVQHCKTAAEIWATLNKLFMETSLLNR 121
Query: 101 QCLKQQLYFYRMVESKTNNEATD 123
+ +LY ++MV++ + ++ D
Sbjct: 122 IYTQLKLYSFKMVDTLSIDQNVD 144
>At4g16870 retrotransposon like protein
Length = 1474
Score = 33.5 bits (75), Expect = 0.20
Identities = 29/112 (25%), Positives = 50/112 (43%), Gaps = 12/112 (10%)
Query: 7 NIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPA-HLTPAEKTEMN--------- 56
N+ K T SN++ +W +++ +L + L G + PA LT N
Sbjct: 30 NVTKLT-SNNYLMWSLQIHALLDGYELAGHLDGSIETPAPTLTTNNVVSANPQYTLWKRQ 88
Query: 57 DKAI-SAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQL 107
D+ I SA++ + V V+R T +W L + Y S+ H + L+ Q+
Sbjct: 89 DRLIFSALIGAISPPVQPLVSRATKASQIWKTLTNTYAKSSYDHIKQLRTQI 140
>At1g44510 polyprotein, putative
Length = 1459
Score = 33.1 bits (74), Expect = 0.26
Identities = 26/112 (23%), Positives = 48/112 (42%), Gaps = 12/112 (10%)
Query: 7 NIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKT----------EMN 56
N+ K T +N + +W +++ +L L +P T +
Sbjct: 29 NVTKLTSTN-YLMWSIQIHALLDGYDLAGYLDNSVVIPPETTTINSVVSANPSFTLWKRQ 87
Query: 57 DKAI-SAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQL 107
DK I SA++ + V V+R T +W+ L++ Y S+ H + L+QQ+
Sbjct: 88 DKLIFSALIGAISPAVQSLVSRATNSSQIWSTLNNTYAKPSYGHIKQLRQQI 139
>At4g04380 putative polyprotein
Length = 778
Score = 32.3 bits (72), Expect = 0.44
Identities = 22/70 (31%), Positives = 41/70 (58%), Gaps = 6/70 (8%)
Query: 50 AEKTEMNDKAISAIVLCLGDKVLREVARETIVVSMWNKL----DSLYMTKSWTHRQCLKQ 105
+EK++ +K V +K + R TIV+S+ +++ D LYM+K+ ++ LKQ
Sbjct: 32 SEKSDEEEKEECEKVEAFEEK--KRKTRSTIVLSVSDRVLEASDRLYMSKALPNQIYLKQ 89
Query: 106 QLYFYRMVES 115
+LY ++M E+
Sbjct: 90 KLYRFKMSEN 99
>At1g59265 polyprotein, putative
Length = 1466
Score = 31.2 bits (69), Expect = 0.98
Identities = 26/112 (23%), Positives = 45/112 (39%), Gaps = 12/112 (10%)
Query: 7 NIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAE----------KTEMN 56
N+ K T +N + +W ++ + + L G MP + + +
Sbjct: 22 NVTKLTSTN-YLMWSRQVHALFDGYELAGFLDGSTTMPPATIGTDAAPRVNPDYTRWKRQ 80
Query: 57 DKAI-SAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQL 107
DK I SA++ + V V+R T +W L +Y S+ H L+ QL
Sbjct: 81 DKLIYSAVLGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQLRTQL 132
>At1g58889 polyprotein, putative
Length = 1466
Score = 31.2 bits (69), Expect = 0.98
Identities = 26/112 (23%), Positives = 45/112 (39%), Gaps = 12/112 (10%)
Query: 7 NIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAE----------KTEMN 56
N+ K T +N + +W ++ + + L G MP + + +
Sbjct: 22 NVTKLTSTN-YLMWSRQVHALFDGYELAGFLDGSTTMPPATIGTDAAPRVNPDYTRWKRQ 80
Query: 57 DKAI-SAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQL 107
DK I SA++ + V V+R T +W L +Y S+ H L+ QL
Sbjct: 81 DKLIYSAVLGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQLRTQL 132
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.356 0.157 0.537
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,673,274
Number of Sequences: 26719
Number of extensions: 224217
Number of successful extensions: 981
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 950
Number of HSP's gapped (non-prelim): 27
length of query: 378
length of database: 11,318,596
effective HSP length: 101
effective length of query: 277
effective length of database: 8,619,977
effective search space: 2387733629
effective search space used: 2387733629
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC123575.7


