

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123571.7 - phase: 0
(247 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
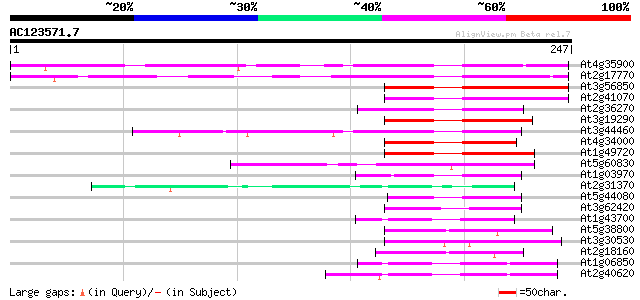
Score E
Sequences producing significant alignments: (bits) Value
At4g35900 bZIP transcription factor AtbZIP14 116 1e-26
At2g17770 bZIP transcription factor atbzip27 100 7e-22
At3g56850 ABA-responsive element binding bZip transcription fact... 67 7e-12
At2g41070 bZIP transcription factor AtbZIP12 / DPBF4 63 1e-10
At2g36270 bZip transcription factor AtbZip39 59 2e-09
At3g19290 abscisic acid responsive elements-binding factor (ABRE... 57 9e-09
At3g44460 bZIP transcription factor DPBF2 / AtbZip67 56 2e-08
At4g34000 abscisic acid responsive elements-binding factor (ABRE... 55 5e-08
At1g49720 abscisic acid responsive elements-binding factor (ABRE... 55 5e-08
At5g60830 bZip transcription factor AtbZip70 49 3e-06
At1g03970 G-box binding bZip transcription factor GBF4 / AtbZip40 48 6e-06
At2g31370 bZIP transcription factor PosF21 / AtbZip59 45 4e-05
At5g44080 bZIP protein AtbZIP13 44 8e-05
At3g62420 bZip transcription factor AtbZip53 44 8e-05
At1g43700 VirE2-interacting protein VIP1 / bZip factor AtbZip51 43 2e-04
At5g38800 bZIP transcription factor AtbZip43 41 5e-04
At3g30530 bZip transcription factor AtbZip42 41 5e-04
At2g18160 G-box binding bZIP transcription factor GBF5 / AtbZip2 41 5e-04
At1g06850 bZip transcription factor AtbZip52 41 5e-04
At2g40620 bZIP transcription factor AtbZip18 41 7e-04
>At4g35900 bZIP transcription factor AtbZIP14
Length = 285
Score = 116 bits (290), Expect = 1e-26
Identities = 96/259 (37%), Positives = 124/259 (47%), Gaps = 52/259 (20%)
Query: 1 MEEVWKDINLSSLN---------DQNTRPMIMSTRNSTFGGGVILQDFLTRPLTLDPTKS 51
MEEVW DINL+S++ N P + I QDFL L +P +
Sbjct: 63 MEEVWNDINLASIHHLNRHSPHPQHNHEPRFRGQNHHNQNPNSIFQDFLKGSLNQEPAPT 122
Query: 52 LDYSSNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLY----DPLIRHNK 107
S + S N ++ + S+ PP T LSLNS F + DPL+
Sbjct: 123 --------SQTTGSAPNGDSTTVTVLYSSPFPPPATVLSLNSGAGFEFLDNQDPLVT--- 171
Query: 108 HNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISPGER 167
+NS L N N S VP+SS FGK+ G+ + G R
Sbjct: 172 -SNSNLHTHHHLSNAHAFNTS----------FEALVPSSS----FGKKRGQDSNEGSGNR 216
Query: 168 RNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQELW 227
R+KRMIKNRESAARSRARKQ AYTNELE +V LQ ENARL+RQQ +L
Sbjct: 217 RHKRMIKNRESAARSRARKQ------------AYTNELELEVAHLQAENARLKRQQDQL- 263
Query: 228 EAESGGQQKKKSSLYRTSS 246
+ + QQ KK++L R+S+
Sbjct: 264 KMAAAIQQPKKNTLQRSST 282
>At2g17770 bZIP transcription factor atbzip27
Length = 234
Score = 100 bits (249), Expect = 7e-22
Identities = 89/251 (35%), Positives = 114/251 (44%), Gaps = 64/251 (25%)
Query: 1 MEEVWKDINLSSLNDQNT-----RPMIMSTRNSTFGGGVILQDFLTRPLTLDPTKSLDYS 55
MEEVWK+INL SL+ PM+ + + I QDFL PL P S
Sbjct: 40 MEEVWKEINLGSLHYHRQLNIGHEPMLKNQNPNNS----IFQDFLNMPLNQPPPPPPPPS 95
Query: 56 SNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLYDPLIRHNKHNNSQLLL 115
S+ +++ S PP T LSLNS F +
Sbjct: 96 SSTIVTALYG-------------SLPLPPPATVLSLNSGVGFEF---------------- 126
Query: 116 QQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISPGERRNKRMIKN 175
N+ SN S+ F C GK+ G+ D + G+RR KRMIKN
Sbjct: 127 LDTTENLLASNPR-------------SFEESAKFGCLGKKRGQDSDDTRGDRRYKRMIKN 173
Query: 176 RESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQELWEAESGGQQ 235
RESAARSRARKQ AYTNELE ++ LQ ENARL+ QQ++L AE+ Q
Sbjct: 174 RESAARSRARKQ------------AYTNELELEIAHLQTENARLKIQQEQLKIAEATQNQ 221
Query: 236 KKKSSLYRTSS 246
KK +L R+S+
Sbjct: 222 VKK-TLQRSST 231
>At3g56850 ABA-responsive element binding bZip transcription factor
AREB3 / AtbZip66
Length = 297
Score = 67.4 bits (163), Expect = 7e-12
Identities = 44/81 (54%), Positives = 49/81 (60%), Gaps = 12/81 (14%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT+ELE KV L+EEN RLR+Q++
Sbjct: 226 ERRQKRMIKNRESAARSRARKQ------------AYTHELEIKVSRLEEENERLRKQKEV 273
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
S K L RTSS
Sbjct: 274 EKILPSVPPPDPKRQLRRTSS 294
>At2g41070 bZIP transcription factor AtbZIP12 / DPBF4
Length = 262
Score = 63.2 bits (152), Expect = 1e-10
Identities = 42/81 (51%), Positives = 48/81 (58%), Gaps = 12/81 (14%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT+ELE KV L+EEN +LRR ++
Sbjct: 191 ERRQKRMIKNRESAARSRARKQ------------AYTHELEIKVSRLEEENEKLRRLKEV 238
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
S K L RT+S
Sbjct: 239 EKILPSEPPPDPKWKLRRTNS 259
>At2g36270 bZip transcription factor AtbZip39
Length = 442
Score = 58.9 bits (141), Expect = 2e-09
Identities = 37/73 (50%), Positives = 44/73 (59%), Gaps = 12/73 (16%)
Query: 154 KRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQ 213
KR + P ERR +RMIKNRESAARSRARKQ AYT ELE ++ L+
Sbjct: 344 KRVVDGPVEKVVERRQRRMIKNRESAARSRARKQ------------AYTVELEAELNQLK 391
Query: 214 EENARLRRQQQEL 226
EENA+L+ EL
Sbjct: 392 EENAQLKHALAEL 404
>At3g19290 abscisic acid responsive elements-binding factor
(ABRE/ABF4) / bZip transcription factor AtbZip38
Length = 431
Score = 57.0 bits (136), Expect = 9e-09
Identities = 32/65 (49%), Positives = 43/65 (65%), Gaps = 12/65 (18%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR +RMIKNRESAARSRARKQ AYT ELE +++ L++ N L+++Q E
Sbjct: 352 ERRQRRMIKNRESAARSRARKQ------------AYTLELEAEIEKLKKTNQELQKKQAE 399
Query: 226 LWEAE 230
+ E +
Sbjct: 400 MVEMQ 404
>At3g44460 bZIP transcription factor DPBF2 / AtbZip67
Length = 331
Score = 55.8 bits (133), Expect = 2e-08
Identities = 54/183 (29%), Positives = 80/183 (43%), Gaps = 29/183 (15%)
Query: 55 SSNNSSSSVASDQNNNNAS---FYCPISTTPPPLVTALSLNSRPDFLYDPLI-------- 103
S NS+ ++ Q + F PL T + ++S DF Y+P
Sbjct: 130 SGQNSAENIRRQQTLGEITLEDFLVKAGVVQEPLKTTMRMSSS-DFGYNPEFGVGLHCQN 188
Query: 104 RHNKHNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNV-GVPASSSFTCFGKRFGEAPDI 162
++N +N + + + + S C +Q + G+ A KR + P
Sbjct: 189 QNNYGDNRSVYSENRPFYSVLGESSSCMTGNGRSNQYLTGLDAFR----IKKRIIDGPPE 244
Query: 163 SPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQ 222
ERR +RMIKNRESAARSRAR+Q AYT ELE ++ L EEN +L+
Sbjct: 245 ILMERRQRRMIKNRESAARSRARRQ------------AYTVELELELNNLTEENTKLKEI 292
Query: 223 QQE 225
+E
Sbjct: 293 VEE 295
>At4g34000 abscisic acid responsive elements-binding factor
(ABRE/ABF3) / bZip transcription factor AtbZip37
Length = 449
Score = 54.7 bits (130), Expect = 5e-08
Identities = 32/58 (55%), Positives = 38/58 (65%), Gaps = 12/58 (20%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQ 223
ERR KRMIKNRESAARSRARKQ AYT ELE ++ L+E N L+++Q
Sbjct: 373 ERRQKRMIKNRESAARSRARKQ------------AYTMELEAEIAQLKELNEELQKKQ 418
>At1g49720 abscisic acid responsive elements-binding factor
(ABRE/ABF1)/ bZip transcription factor AtbZip35
Length = 392
Score = 54.7 bits (130), Expect = 5e-08
Identities = 32/66 (48%), Positives = 42/66 (63%), Gaps = 12/66 (18%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT ELE +++ L+ N L+++Q E
Sbjct: 312 ERRQKRMIKNRESAARSRARKQ------------AYTLELEAEIESLKLVNQDLQKKQAE 359
Query: 226 LWEAES 231
+ + +
Sbjct: 360 IMKTHN 365
>At5g60830 bZip transcription factor AtbZip70
Length = 188
Score = 48.5 bits (114), Expect = 3e-06
Identities = 43/136 (31%), Positives = 63/136 (45%), Gaps = 14/136 (10%)
Query: 98 LYDPLIRHNKHNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFG 157
+ D + H+ H NS L + SNVS N S D N A + F G
Sbjct: 22 ILDGVPLHDDHFNSAFLPNTDFNVHLQSNVSTRINNQSHLDPN----AENIFHNEG---- 73
Query: 158 EAPDISPGERRNKRMIKNRESAARSRARKQEKITSF--LFSKFSAYTNELEQKVQLLQEE 215
++P ERR +RM+ NRESA RSR RK+++I + + L +KV L E
Sbjct: 74 ----LAPEERRARRMVSNRESARRSRMRKKKQIEELQQQVEQLMMLNHHLSEKVINLLES 129
Query: 216 NARLRRQQQELWEAES 231
N ++ ++ +L E S
Sbjct: 130 NHQILQENSQLKEKVS 145
>At1g03970 G-box binding bZip transcription factor GBF4 / AtbZip40
Length = 270
Score = 47.8 bits (112), Expect = 6e-06
Identities = 33/73 (45%), Positives = 42/73 (57%), Gaps = 13/73 (17%)
Query: 153 GKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLL 212
G+ EA D + +R+ KRMIKNRESAARSR RKQ AY ELE L
Sbjct: 176 GRVMMEAMDKAAAQRQ-KRMIKNRESAARSRERKQ------------AYQVELETLAAKL 222
Query: 213 QEENARLRRQQQE 225
+EEN +L ++ +E
Sbjct: 223 EEENEQLLKEIEE 235
>At2g31370 bZIP transcription factor PosF21 / AtbZip59
Length = 398
Score = 45.1 bits (105), Expect = 4e-05
Identities = 56/189 (29%), Positives = 75/189 (39%), Gaps = 41/189 (21%)
Query: 37 QDFLTRPLTLDPTKSLDYSSNNSSSSVASDQNN---NNASFYCPISTTPPPLVTALSLNS 93
+D L+ L +D S S SS+ V N +T P T SL
Sbjct: 96 EDLLSMYLDMDKFNS----SATSSAQVGEPSGTAWKNETMMQTGTGSTSNPQNTVNSLGE 151
Query: 94 RPDFLYDPLIRHNKHNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFG 153
RP IRH Q Q G N++ ++ + D + S S T
Sbjct: 152 RPR------IRH----------QHSQSMDGSMNINEMLMSGNEDDSAIDAKKSMSAT--- 192
Query: 154 KRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQ 213
+ E I P +R KR+ NR+SAARS+ RK T ++F ELE+KVQ LQ
Sbjct: 193 -KLAELALIDP--KRAKRIWANRQSAARSKERK----TRYIF--------ELERKVQTLQ 237
Query: 214 EENARLRRQ 222
E L Q
Sbjct: 238 TEATTLSAQ 246
>At5g44080 bZIP protein AtbZIP13
Length = 315
Score = 43.9 bits (102), Expect = 8e-05
Identities = 27/59 (45%), Positives = 34/59 (56%), Gaps = 12/59 (20%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
+R +RMIKNRESAARSR RKQ AY ELE L+EEN L ++ ++
Sbjct: 233 QRQRRMIKNRESAARSRERKQ------------AYQVELEALAAKLEEENELLSKEIED 279
>At3g62420 bZip transcription factor AtbZip53
Length = 146
Score = 43.9 bits (102), Expect = 8e-05
Identities = 25/60 (41%), Positives = 35/60 (57%), Gaps = 12/60 (20%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ER+ KRMI NRESA RSR RKQ+++ +L +V LL+ +NA++ Q E
Sbjct: 24 ERKRKRMISNRESARRSRMRKQKQL------------GDLINEVTLLKNDNAKITEQVDE 71
>At1g43700 VirE2-interacting protein VIP1 / bZip factor AtbZip51
Length = 341
Score = 42.7 bits (99), Expect = 2e-04
Identities = 29/70 (41%), Positives = 36/70 (51%), Gaps = 14/70 (20%)
Query: 153 GKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLL 212
G R E + P +R KR++ NR+SAARS+ RK YT ELE+KVQ L
Sbjct: 184 GDRLAELALLDP--KRAKRILANRQSAARSKERK------------IRYTGELERKVQTL 229
Query: 213 QEENARLRRQ 222
Q E L Q
Sbjct: 230 QNEATTLSAQ 239
>At5g38800 bZIP transcription factor AtbZip43
Length = 165
Score = 41.2 bits (95), Expect = 5e-04
Identities = 30/84 (35%), Positives = 44/84 (51%), Gaps = 11/84 (13%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQ----------EE 215
ER+ KR I NRESA RSR RKQ ++ L+S+ +E Q ++ L EE
Sbjct: 71 ERKQKRKISNRESARRSRMRKQRQVDE-LWSQVMWLRDENHQLLRKLNCVLESQEKVIEE 129
Query: 216 NARLRRQQQELWEAESGGQQKKKS 239
N +L+ + EL + S Q + +S
Sbjct: 130 NVQLKEETTELKQMISDMQLQNQS 153
>At3g30530 bZip transcription factor AtbZip42
Length = 173
Score = 41.2 bits (95), Expect = 5e-04
Identities = 28/87 (32%), Positives = 45/87 (51%), Gaps = 9/87 (10%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKI----TSFLFSKFSAY-----TNELEQKVQLLQEEN 216
ER+ +RMI NRESA RSR RKQ + + ++ + + N L + + +EN
Sbjct: 80 ERKQRRMISNRESARRSRMRKQRHLDELWSQVMWLRIENHQLLDKLNNLSESHDKVLQEN 139
Query: 217 ARLRRQQQELWEAESGGQQKKKSSLYR 243
A+L+ + EL + S Q + S +R
Sbjct: 140 AQLKEETFELKQVISDMQIQSPFSCFR 166
>At2g18160 G-box binding bZIP transcription factor GBF5 / AtbZip2
Length = 171
Score = 41.2 bits (95), Expect = 5e-04
Identities = 27/75 (36%), Positives = 41/75 (54%), Gaps = 11/75 (14%)
Query: 162 ISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLL--------- 212
++ ER+ KRM+ NRESA RSR RKQ+ + L ++ + +N+ Q + L
Sbjct: 26 VTVDERKRKRMLSNRESARRSRMRKQKHVDD-LTAQINQLSNDNRQILNSLTVTSQLYMK 84
Query: 213 -QEENARLRRQQQEL 226
Q EN+ L Q +EL
Sbjct: 85 IQAENSVLTAQMEEL 99
>At1g06850 bZip transcription factor AtbZip52
Length = 337
Score = 41.2 bits (95), Expect = 5e-04
Identities = 30/88 (34%), Positives = 46/88 (52%), Gaps = 15/88 (17%)
Query: 154 KRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQ 213
++ E +I P +R KR++ NR+SAARS+ RK + Y ELE+KVQ LQ
Sbjct: 139 EKLSELWNIDP--KRAKRILANRQSAARSKERK------------ARYIQELERKVQSLQ 184
Query: 214 EENARLRRQQQELWEAESGGQQKKKSSL 241
E L Q L++ ++ G + + L
Sbjct: 185 TEATTL-SAQLTLYQRDTNGLANENTEL 211
>At2g40620 bZIP transcription factor AtbZip18
Length = 367
Score = 40.8 bits (94), Expect = 7e-04
Identities = 35/109 (32%), Positives = 52/109 (47%), Gaps = 22/109 (20%)
Query: 140 NVGVPASSSFTCFGKRFGEAPD-------ISPGERRNKRMIKNRESAARSRARKQEKITS 192
++ V SS+ + APD + P +R KR+I NR+SAARS+ RK
Sbjct: 118 SLSVDGSSTLESIEAKKAMAPDKLAELWVVDP--KRAKRIIANRQSAARSKERK------ 169
Query: 193 FLFSKFSAYTNELEQKVQLLQEENARLRRQQQELWEAESGGQQKKKSSL 241
+ Y ELE+KVQ LQ E L Q L++ ++ G + + L
Sbjct: 170 ------ARYILELERKVQTLQTEATTL-SAQLSLFQRDTTGLSSENTEL 211
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.127 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,642,715
Number of Sequences: 26719
Number of extensions: 238791
Number of successful extensions: 1200
Number of sequences better than 10.0: 125
Number of HSP's better than 10.0 without gapping: 60
Number of HSP's successfully gapped in prelim test: 67
Number of HSP's that attempted gapping in prelim test: 1074
Number of HSP's gapped (non-prelim): 156
length of query: 247
length of database: 11,318,596
effective HSP length: 97
effective length of query: 150
effective length of database: 8,726,853
effective search space: 1309027950
effective search space used: 1309027950
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC123571.7


