

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123570.7 + phase: 0
(601 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
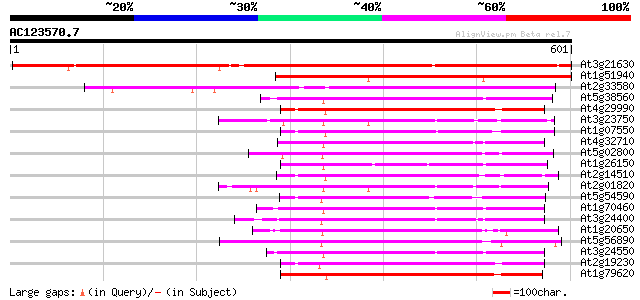
Score E
Sequences producing significant alignments: (bits) Value
At3g21630 receptor like protein kinase 642 0.0
At1g51940 protein kinase like protein 269 3e-72
At2g33580 putative protein kinase 236 3e-62
At5g38560 putative protein 224 8e-59
At4g29990 serine/threonine-specific receptor protein kinase LRRPK 224 1e-58
At3g23750 receptor-like kinase, putative 223 2e-58
At1g07550 protein kinase like 219 3e-57
At4g32710 putative protein kinase 217 1e-56
At5g02800 protein kinase - like 216 3e-56
At1g26150 Pto kinase interactor, putative 216 4e-56
At2g14510 putative receptor-like protein kinase 214 8e-56
At2g01820 putative receptor-like protein kinase 214 1e-55
At5g54590 serine/threonine-specific protein kinase-like protein 214 1e-55
At1g70460 putative protein kinase 213 2e-55
At3g24400 protein kinase, putative 212 4e-55
At1g20650 unknown protein 212 5e-55
At5g56890 unknown protein 211 7e-55
At3g24550 protein kinase, putative 211 7e-55
At2g19230 putative receptor-like protein kinase 211 1e-54
At1g79620 unknown protein 211 1e-54
>At3g21630 receptor like protein kinase
Length = 617
Score = 642 bits (1657), Expect = 0.0
Identities = 342/624 (54%), Positives = 436/624 (69%), Gaps = 35/624 (5%)
Query: 4 IKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPE-- 61
+K L +LL S F ESKC +C +ALASYYL++ T L+ ++ + S++ +
Sbjct: 3 LKISLIAPILLLFSFFFAVESKCRTSCPLALASYYLENGTTLSVINQNLNSSIAPYDQIN 62
Query: 62 --DIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNY 119
I+ YN++ I +KD +Q +RV VPFPC+C +FLGH F Y V +DTY VA +NY
Sbjct: 63 FDPILRYNSN-IKDKDRIQMGSRVLVPFPCECQPGDFLGHNFSYSVRQEDTYERVAISNY 121
Query: 120 SNLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELI 179
+NLTT E LQ N +P+ +IP + TLNV VNCSCG+ VSKD+GLF+TYPLRPEDSL I
Sbjct: 122 ANLTTMESLQARNPFPATNIPLSATLNVLVNCSCGDESVSKDFGLFVTYPLRPEDSLSSI 181
Query: 180 SNKTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTR---------------- 223
+ + + A++LQ+YNPGVNF+ G+G+VY+PG+D N + PF +
Sbjct: 182 ARSSGVSADILQRYNPGVNFNSGNGIVYVPGRDPNGAFPPFKSSKQDGVGAGVIAGIVIG 241
Query: 224 ----LILLSFCIYVTYYRKKKIRKQEFLSEESSAIFGQVKNDEVSGNATY--GTSDSASP 277
L+L+ F +Y Y RK K + F SS+I K D S + G +
Sbjct: 242 VIVALLLILFIVYYAY-RKNKSKGDSF----SSSIPLSTKADHASSTSLQSGGLGGAGVS 296
Query: 278 ANMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKAT 337
+ I V+KS EFS EELA AT+NFN++ KIGQGGFG VYYAEL GEKAAIKKMDM+A+
Sbjct: 297 PGIAAISVDKSVEFSLEELAKATDNFNLSFKIGQGGFGAVYYAELRGEKAAIKKMDMEAS 356
Query: 338 KEFLAELKVLTRVHHVNLVRLIGYCVEGSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRV 397
K+FLAELKVLTRVHHVNLVRLIGYCVEGSLFLVYEY++NGNLGQHL S EPL W+ RV
Sbjct: 357 KQFLAELKVLTRVHHVNLVRLIGYCVEGSLFLVYEYVENGNLGQHLHGSGREPLPWTKRV 416
Query: 398 KIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTV 457
+IALDSARGLEYIHEHTVP Y+HRDIKS NIL+D+ F AKVADFGLTKL + G S+ T
Sbjct: 417 QIALDSARGLEYIHEHTVPVYVHRDIKSANILIDQKFRAKVADFGLTKLTEVGGSA--TR 474
Query: 458 NMAGTFGYMPPEYAYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEE 517
GTFGYM PE YG VS+K+DVYAFGVVLYELISAK AV+ ++ + +GLV +FEE
Sbjct: 475 GAMGTFGYMAPETVYGEVSAKVDVYAFGVVLYELISAKGAVVKMTEAVGEFRGLVGVFEE 534
Query: 518 VFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLT 577
F + E L+K++DPRLGD+YP D V+KMA+L K CT + Q RP+M +VVAL+TL
Sbjct: 535 SFKETDKEEALRKIIDPRLGDSYPFDSVYKMAELGKACTQENAQLRPSMRYIVVALSTLF 594
Query: 578 STTEDWDITSIFKNPNLVNLMSGR 601
S+T +WD+ + F+N +LV+LMSGR
Sbjct: 595 SSTGNWDVGN-FQNEDLVSLMSGR 617
>At1g51940 protein kinase like protein
Length = 601
Score = 269 bits (688), Expect = 3e-72
Identities = 147/329 (44%), Positives = 205/329 (61%), Gaps = 12/329 (3%)
Query: 285 VEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKATKEFLAEL 344
+EK F+YEE+ AT+ F+ +N +G G +G VY+ L ++ A+K+M TKEF AE+
Sbjct: 273 IEKPMVFTYEEIRAATDEFSDSNLLGHGNYGSVYFGLLREQEVAVKRMTATKTKEFAAEM 332
Query: 345 KVLTRVHHVNLVRLIGYCVE-GSLFLVYEYIDNGNLGQHL---RSSDGEPLSWSIRVKIA 400
KVL +VHH NLV LIGY LF+VYEY+ G L HL +S PLSW +R +IA
Sbjct: 333 KVLCKVHHSNLVELIGYAATVDELFVVYEYVRKGMLKSHLHDPQSKGNTPLSWIMRNQIA 392
Query: 401 LDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLID-AGISSVPTVNM 459
LD+ARGLEYIHEHT Y+HRDIK+ NILLD+ F AK++DFGL KL++ G + +
Sbjct: 393 LDAARGLEYIHEHTKTHYVHRDIKTSNILLDEAFRAKISDFGLAKLVEKTGEGEISVTKV 452
Query: 460 AGTFGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGA------DLKGLV 512
GT+GY+ PEY + G +SK D+YAFGVVL+E+IS + AVI E G L ++
Sbjct: 453 VGTYGYLAPEYLSDGLATSKSDIYAFGVVLFEIISGREAVIRTEAIGTKNPERRPLASIM 512
Query: 513 VLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVA 572
+ + + LK+ VDP + D YP D +FK+A LAK C + DP RPNM VV++
Sbjct: 513 LAVLKNSPDSMNMSSLKEFVDPNMMDLYPHDCLFKIATLAKQCVDDDPILRPNMKQVVIS 572
Query: 573 LTTLTSTTEDWDITSIFKNPNLVNLMSGR 601
L+ + ++ +W+ T + L+ GR
Sbjct: 573 LSQILLSSIEWEATLAGNSQVFSGLVQGR 601
>At2g33580 putative protein kinase
Length = 664
Score = 236 bits (602), Expect = 3e-62
Identities = 157/557 (28%), Positives = 281/557 (50%), Gaps = 60/557 (10%)
Query: 81 TRVNVPFPCDCIHDEFLGHIFQYQVATK-------DTYLSVASNNYSNLTTSEWLQNFNS 133
TR V P +C G +Q+ +TY SVA++ Y L+T + + + N
Sbjct: 102 TRELVVIPANCSCSSSSGGFYQHNATYNLSGNRGDETYFSVANDTYQALSTCQAMMSQNR 161
Query: 134 YPSNDIPDTGTLNVTVNCSCGNS-DVSKDYGLFITYPLRPEDSLELISNKTEIDAELLQK 192
Y + L V + C+C + + + +TY + DS+ I+ + + +
Sbjct: 162 YGERQLTPGLNLLVPLRCACPTAKQTTAGFKYLLTYLVAMGDSISGIAEMFNSTSAAITE 221
Query: 193 YN--------------------------------PGVNFSQGSGLVYIPGKDQNRNYV-- 218
N P V + V PG + ++
Sbjct: 222 GNELTSDNIFFFTPVLVPLTTEPTKIVISPSPPPPPVVATPPQTPVDPPGSSSSHKWIYI 281
Query: 219 --PFHTRLILLSFCIYVTYYRKKKIRKQ--EFLSEESSAIFGQVKNDEVSGNATYGTSDS 274
L+LL + + +Y+++ +K L EE+ K + T + D
Sbjct: 282 GIGIGAGLLLLLSILALCFYKRRSKKKSLPSSLPEENKLFDSSTKQSIPTTTTTQWSIDL 341
Query: 275 ASPANMIGIR--VEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKM 332
++ + G++ +E + + +L +AT+NF+ N+I G VY A +NG+ AA+K +
Sbjct: 342 SNSSEAFGLKSAIESLTLYRFNDLQSATSNFSDENRIK----GSVYRATINGDDAAVKVI 397
Query: 333 DMKATKEFLAELKVLTRVHHVNLVRLIGYCV-EGSLFLVYEYIDNGNLGQHLRSSDGEPL 391
+ +E+ +L +++H N++RL G+C+ EG+ +LV+EY +NG++ L SS + L
Sbjct: 398 KGDVSS---SEINLLKKLNHSNIIRLSGFCIREGTSYLVFEYSENGSISDWLHSSGKKSL 454
Query: 392 SWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAG- 450
+W RV+IA D A L+Y+H + P +IH++++S NILLD NF AK+A+FG+ +++D G
Sbjct: 455 TWKQRVEIARDVAEALDYLHNYITPPHIHKNLESTNILLDSNFRAKIANFGVARILDEGD 514
Query: 451 ISSVPTVNMAGTFGYMPPEYA-YGSVSSKIDVYAFGVVLYELISAKAAVIM--GEDSGAD 507
+ T ++ GT GY+ PEY G ++SK+DV+AFGV + EL+S + AV + ++ +
Sbjct: 515 LDLQLTRHVEGTQGYLAPEYVENGVITSKLDVFAFGVAVLELLSGREAVTIHKKKEGEEE 574
Query: 508 LKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMS 567
++ L + V + E LK+ +DP LG+ YP++ + MAQLAK C +D RP+++
Sbjct: 575 VEMLCKVINSVLGGENVREKLKEFMDPSLGNEYPLELAYTMAQLAKSCVATDLNSRPSVT 634
Query: 568 SVVVALTTLTSTTEDWD 584
V+ L+ + S++ DW+
Sbjct: 635 QVLTTLSMIVSSSIDWE 651
>At5g38560 putative protein
Length = 681
Score = 224 bits (572), Expect = 8e-59
Identities = 130/320 (40%), Positives = 189/320 (58%), Gaps = 16/320 (5%)
Query: 269 YGTSDSASPANMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKA 327
Y +SDS +N + FSY+EL+ T+ F+ N +G+GGFG VY L+ G +
Sbjct: 312 YASSDSGMVSN-------QRSWFSYDELSQVTSGFSEKNLLGEGGFGCVYKGVLSDGREV 364
Query: 328 AIKKMDM---KATKEFLAELKVLTRVHHVNLVRLIGYCV-EGSLFLVYEYIDNGNLGQHL 383
A+K++ + + +EF AE+++++RVHH +LV L+GYC+ E LVY+Y+ N L HL
Sbjct: 365 AVKQLKIGGSQGEREFKAEVEIISRVHHRHLVTLVGYCISEQHRLLVYDYVPNNTLHYHL 424
Query: 384 RSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGL 443
+ ++W RV++A +ARG+ Y+HE P IHRDIKS NILLD +F A VADFGL
Sbjct: 425 HAPGRPVMTWETRVRVAAGAARGIAYLHEDCHPRIIHRDIKSSNILLDNSFEALVADFGL 484
Query: 444 TKLI-DAGISSVPTVNMAGTFGYMPPEYA-YGSVSSKIDVYAFGVVLYELISAKAAVIMG 501
K+ + +++ + + GTFGYM PEYA G +S K DVY++GV+L ELI+ + V
Sbjct: 485 AKIAQELDLNTHVSTRVMGTFGYMAPEYATSGKLSEKADVYSYGVILLELITGRKPVDTS 544
Query: 502 EDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQ 561
+ G + LV + Q E +LVDPRLG N+ +F+M + A C
Sbjct: 545 QPLGDE--SLVEWARPLLGQAIENEEFDELVDPRLGKNFIPGEMFRMVEAAAACVRHSAA 602
Query: 562 QRPNMSSVVVALTTLTSTTE 581
+RP MS VV AL TL T+
Sbjct: 603 KRPKMSQVVRALDTLEEATD 622
>At4g29990 serine/threonine-specific receptor protein kinase LRRPK
Length = 876
Score = 224 bits (570), Expect = 1e-58
Identities = 125/288 (43%), Positives = 176/288 (60%), Gaps = 14/288 (4%)
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKAT---KEFLAELKVL 347
F Y E+ N TNNF +G+GGFG+VY+ LNG++ A+K + ++T KEF AE+++L
Sbjct: 564 FIYSEVVNITNNFERV--LGKGGFGKVYHGFLNGDQVAVKILSEESTQGYKEFRAEVELL 621
Query: 348 TRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSARG 406
RVHH NL LIGYC E + + L+YEY+ NGNLG +L LSW R++I+LD+A+G
Sbjct: 622 MRVHHTNLTSLIGYCNEDNHMALIYEYMANGNLGDYLSGKSSLILSWEERLQISLDAAQG 681
Query: 407 LEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYM 466
LEY+H P +HRD+K NILL++N AK+ADFGL++ SS + +AGT GY+
Sbjct: 682 LEYLHYGCKPPIVHRDVKPANILLNENLQAKIADFGLSRSFPVEGSSQVSTVVAGTIGYL 741
Query: 467 PPE-YAYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPI 525
PE YA ++ K DVY+FGVVL E+I+ K A+ L V D
Sbjct: 742 DPEYYATRQMNEKSDVYSFGVVLLEVITGKPAIWHSRTESVHLSDQVGSMLANGD----- 796
Query: 526 EGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVAL 573
+K +VD RLGD + + +K+ +LA C + +QRP MS VV+ L
Sbjct: 797 --IKGIVDQRLGDRFEVGSAWKITELALACASESSEQRPTMSQVVMEL 842
>At3g23750 receptor-like kinase, putative
Length = 928
Score = 223 bits (568), Expect = 2e-58
Identities = 156/387 (40%), Positives = 210/387 (53%), Gaps = 39/387 (10%)
Query: 224 LILLSFCIYVTYYRKKKIRKQEFLSEESSAIF-GQVKNDEVSGNATYGTSDSASPANMIG 282
L +L F +Y ++K R E+ I ++ SGN Y A+ N
Sbjct: 487 LAILGFVVYKFVMKRKYGRFNRTDPEKVGKILVSDAVSNGGSGNGGYANGHGAN--NFNA 544
Query: 283 IRVEKSGEFS-------------YEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAA 328
+ SG+ S E L TNNF+ N +G+GGFG VY EL+ G K A
Sbjct: 545 LNSPSSGDNSDRFLLEGGSVTIPMEVLRQVTNNFSEDNILGRGGFGVVYAGELHDGTKTA 604
Query: 329 IKKMDM-----KATKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQH 382
+K+M+ K EF AE+ VLT+V H +LV L+GYCV G+ LVYEY+ GNLGQH
Sbjct: 605 VKRMECAAMGNKGMSEFQAEIAVLTKVRHRHLVALLGYCVNGNERLLVYEYMPQGNLGQH 664
Query: 383 L---RSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVA 439
L PL+W RV IALD ARG+EY+H ++IHRD+K NILL + AKVA
Sbjct: 665 LFEWSELGYSPLTWKQRVSIALDVARGVEYLHSLAQQSFIHRDLKPSNILLGDDMRAKVA 724
Query: 440 DFGLTKLIDAGISSVPTVNMAGTFGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAV 498
DFGL K G SV T +AGTFGY+ PEY A G V++K+DVYAFGVVL E+++ + A+
Sbjct: 725 DFGLVKNAPDGKYSVET-RLAGTFGYLAPEYAATGRVTTKVDVYAFGVVLMEILTGRKAL 783
Query: 499 IMGEDSGADLKG-LVVLFEEVFDQPHPIEGLKKLVDPRL-GDNYPIDHVFKMAQLAKVCT 556
+DS D + LV F + E + K +D L D ++ ++++A+LA CT
Sbjct: 784 ---DDSLPDERSHLVTWFRRILINK---ENIPKALDQTLEADEETMESIYRVAELAGHCT 837
Query: 557 NSDPQQRPNMSSVVVALTTLTSTTEDW 583
+PQQRP+M V L L E W
Sbjct: 838 AREPQQRPDMGHAVNVLGPL---VEKW 861
>At1g07550 protein kinase like
Length = 864
Score = 219 bits (558), Expect = 3e-57
Identities = 129/299 (43%), Positives = 178/299 (59%), Gaps = 17/299 (5%)
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKAT---KEFLAELKVL 347
F+Y ++ TNNF + IG+GGFG VY LN E+AAIK + + KEF E+++L
Sbjct: 550 FTYSDVNKMTNNFQVV--IGKGGFGVVYQGCLNNEQAAIKVLSHSSAQGYKEFKTEVELL 607
Query: 348 TRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDG-EPLSWSIRVKIALDSAR 405
RVHH LV LIGYC + + L L+YE + GNL +HL G LSW IR+KIAL+SA
Sbjct: 608 LRVHHEKLVSLIGYCDDDNGLALIYELMGKGNLKEHLSGKPGCSVLSWPIRLKIALESAI 667
Query: 406 GLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGY 465
G+EY+H P +HRD+KS NILL + F AK+ADFGL++ G + PTV +AGTFGY
Sbjct: 668 GIEYLHTGCKPKIVHRDVKSTNILLSEEFEAKIADFGLSRSFLIGNEAQPTV-VAGTFGY 726
Query: 466 MPPEYAYGS-VSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHP 524
+ PEY S +S K DVY+FGVVL E+IS + + + ++ ++ + E
Sbjct: 727 LDPEYHKTSLLSMKSDVYSFGVVLLEIISGQDVIDLSRENCNIVEWTSFILEN------- 779
Query: 525 IEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTSTTEDW 583
++ +VDP L +Y +K+ +LA C N ++RPNMS VV L T E W
Sbjct: 780 -GDIESIVDPNLHQDYDTSSAWKVVELAMSCVNRTSKERPNMSQVVHVLNECLETCEKW 837
>At4g32710 putative protein kinase
Length = 731
Score = 217 bits (553), Expect = 1e-56
Identities = 131/294 (44%), Positives = 174/294 (58%), Gaps = 10/294 (3%)
Query: 288 SGEFSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDM---KATKEFLAE 343
SG FSYEEL+ AT F+ N +G+GGFG V+ L NG + A+K++ + + +EF AE
Sbjct: 374 SGMFSYEELSKATGGFSEENLLGEGGFGYVHKGVLKNGTEVAVKQLKIGSYQGEREFQAE 433
Query: 344 LKVLTRVHHVNLVRLIGYCVEGSL-FLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALD 402
+ ++RVHH +LV L+GYCV G LVYE++ L HL + G L W +R++IA+
Sbjct: 434 VDTISRVHHKHLVSLVGYCVNGDKRLLVYEFVPKDTLEFHLHENRGSVLEWEMRLRIAVG 493
Query: 403 SARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVN--MA 460
+A+GL Y+HE PT IHRDIK+ NILLD F AKV+DFGL K SS ++ +
Sbjct: 494 AAKGLAYLHEDCSPTIIHRDIKAANILLDSKFEAKVSDFGLAKFFSDTNSSFTHISTRVV 553
Query: 461 GTFGYMPPEYA-YGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVF 519
GTFGYM PEYA G V+ K DVY+FGVVL ELI+ + + I +DS + + LV +
Sbjct: 554 GTFGYMAPEYASSGKVTDKSDVYSFGVVLLELITGRPS-IFAKDSSTN-QSLVDWARPLL 611
Query: 520 DQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVAL 573
+ E LVD RL NY + MA A C RP MS VV AL
Sbjct: 612 TKAISGESFDFLVDSRLEKNYDTTQMANMAACAAACIRQSAWLRPRMSQVVRAL 665
>At5g02800 protein kinase - like
Length = 395
Score = 216 bits (550), Expect = 3e-56
Identities = 130/342 (38%), Positives = 190/342 (55%), Gaps = 20/342 (5%)
Query: 256 GQVKNDEVSGNATYGTSDSASPANMIGIRVEKSGE------FSYEELANATNNFNMANKI 309
G K D S + + TS+ + + + K + F++ ELA AT NF I
Sbjct: 20 GDHKLDRKSSDCSVSTSEKSRAKSSLSESKSKGSDHIVAQTFTFSELATATRNFRKECLI 79
Query: 310 GQGGFGEVY--YAELNGEKAAIKKMD---MKATKEFLAELKVLTRVHHVNLVRLIGYCVE 364
G+GGFG VY Y + AAIK++D ++ +EFL E+ +L+ +HH NLV LIGYC +
Sbjct: 80 GEGGFGRVYKGYLASTSQTAAIKQLDHNGLQGNREFLVEVLMLSLLHHPNLVNLIGYCAD 139
Query: 365 GSL-FLVYEYIDNGNLGQHLR--SSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHR 421
G LVYEY+ G+L HL S +PL W+ R+KIA +A+GLEY+H+ T+P I+R
Sbjct: 140 GDQRLLVYEYMPLGSLEDHLHDISPGKQPLDWNTRMKIAAGAAKGLEYLHDKTMPPVIYR 199
Query: 422 DIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSKID 480
D+K NILLD ++ K++DFGL KL G S + + GT+GY PEYA G ++ K D
Sbjct: 200 DLKCSNILLDDDYFPKLSDFGLAKLGPVGDKSHVSTRVMGTYGYCAPEYAMTGQLTLKSD 259
Query: 481 VYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNY 540
VY+FGVVL E+I+ + A+ +G + LV +F ++ DP L Y
Sbjct: 260 VYSFGVVLLEIITGRKAIDSSRSTGE--QNLVAWARPLFKDRRK---FSQMADPMLQGQY 314
Query: 541 PIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTSTTED 582
P +++ +A +C P RP ++ VV AL+ L S D
Sbjct: 315 PPRGLYQALAVAAMCVQEQPNLRPLIADVVTALSYLASQKFD 356
>At1g26150 Pto kinase interactor, putative
Length = 760
Score = 216 bits (549), Expect = 4e-56
Identities = 127/292 (43%), Positives = 172/292 (58%), Gaps = 11/292 (3%)
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKA-AIKKMDM---KATKEFLAELKV 346
FSYEEL ATN F+ N +G+GGFG VY L E+ A+K++ + + +EF AE+
Sbjct: 418 FSYEELVIATNGFSDENLLGEGGFGRVYKGVLPDERVVAVKQLKIGGGQGDREFKAEVDT 477
Query: 347 LTRVHHVNLVRLIGYCV-EGSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSAR 405
++RVHH NL+ ++GYC+ E L+Y+Y+ N NL HL + G L W+ RVKIA +AR
Sbjct: 478 ISRVHHRNLLSMVGYCISENRRLLIYDYVPNNNLYFHLHGTPG--LDWATRVKIAAGAAR 535
Query: 406 GLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGY 465
GL Y+HE P IHRDIKS NILL+ NF A V+DFGL KL ++ T + GTFGY
Sbjct: 536 GLAYLHEDCHPRIIHRDIKSSNILLENNFHALVSDFGLAKLA-LDCNTHITTRVMGTFGY 594
Query: 466 MPPEYA-YGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHP 524
M PEYA G ++ K DV++FGVVL ELI+ + V + G + LV +
Sbjct: 595 MAPEYASSGKLTEKSDVFSFGVVLLELITGRKPVDASQPLGDE--SLVEWARPLLSNATE 652
Query: 525 IEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTL 576
E L DP+LG NY +F+M + A C +RP MS +V A +L
Sbjct: 653 TEEFTALADPKLGRNYVGVEMFRMIEAAAACIRHSATKRPRMSQIVRAFDSL 704
>At2g14510 putative receptor-like protein kinase
Length = 868
Score = 214 bits (546), Expect = 8e-56
Identities = 126/308 (40%), Positives = 181/308 (57%), Gaps = 19/308 (6%)
Query: 287 KSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKAT---KEFLAE 343
K+ F Y E+ TNNF + +G+GGFG VY+ LN E+ A+K + +T KEF E
Sbjct: 549 KNRRFKYSEVKEMTNNFEVV--LGKGGFGVVYHGFLNNEQVAVKVLSQSSTQGYKEFKTE 606
Query: 344 LKVLTRVHHVNLVRLIGYCVEG-SLFLVYEYIDNGNLGQHLRSS-DGEPLSWSIRVKIAL 401
+++L RVHHVNLV L+GYC EG L L+YE+++NGNL +HL G L+WS R+KIA+
Sbjct: 607 VELLLRVHHVNLVSLVGYCDEGIDLALIYEFMENGNLKEHLSGKRGGSVLNWSSRLKIAI 666
Query: 402 DSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAG 461
+SA G+EY+H P +HRD+KS NILL F AK+ADFGL++ G + + N+AG
Sbjct: 667 ESALGIEYLHIGCQPPMVHRDVKSTNILLGLRFEAKLADFGLSRSFLVGSQAHVSTNVAG 726
Query: 462 TFGYMPPEYAYGS-VSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFD 520
T GY+ PEY + ++ K DVY+FG+VL E I+ + + D K +V + +
Sbjct: 727 TLGYLDPEYYLKNWLTEKSDVYSFGIVLLESITGQPVIEQSRD-----KSYIVEWAKSML 781
Query: 521 QPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTSTT 580
IE ++DP L +Y +K +LA +C N QRPNM+ V L
Sbjct: 782 ANGDIE---SIMDPNLHQDYDSSSSWKALELAMLCINPSSTQRPNMTRVA---HELNECL 835
Query: 581 EDWDITSI 588
E +++T I
Sbjct: 836 EIYNLTKI 843
>At2g01820 putative receptor-like protein kinase
Length = 943
Score = 214 bits (545), Expect = 1e-55
Identities = 143/379 (37%), Positives = 210/379 (54%), Gaps = 33/379 (8%)
Query: 224 LILLSFCIYVTYYRKKKIRKQEFLSEESSAIFG---QVKNDEV----------SGNATYG 270
L+ L C+Y KK+ R S S+ + ND++ SG +
Sbjct: 497 LVGLGVCLYA----KKRKRPARVQSPSSNMVIHPHHSGDNDDIKLTVAASSLNSGGGSDS 552
Query: 271 TSDSASPANMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAI 329
S S S A+ I + + S + L N TNNF+ N +G+GGFG VY EL+ G K A+
Sbjct: 553 YSHSGSAASDIHVVEAGNLVISIQVLRNVTNNFSEENILGRGGFGTVYKGELHDGTKIAV 612
Query: 330 KKMDM-----KATKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHL 383
K+M+ K EF +E+ VLT++ H +LV L+GYC++G+ LVYEY+ G L QHL
Sbjct: 613 KRMESSVVSDKGLTEFKSEITVLTKMRHRHLVALLGYCLDGNERLLVYEYMPQGTLSQHL 672
Query: 384 ---RSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVAD 440
+ +PL W+ R+ IALD ARG+EY+H ++IHRD+K NILL + AKV+D
Sbjct: 673 FHWKEEGRKPLDWTRRLAIALDVARGVEYLHTLAHQSFIHRDLKPSNILLGDDMRAKVSD 732
Query: 441 FGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYELISAKAAVI 499
FGL +L G S+ T +AGTFGY+ PEYA G V++K+D+++ GV+L ELI+ + A
Sbjct: 733 FGLVRLAPDGKYSIET-RVAGTFGYLAPEYAVTGRVTTKVDIFSLGVILMELITGRKA-- 789
Query: 500 MGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLG-DNYPIDHVFKMAQLAKVCTNS 558
+ E D LV F V K +DP + D+ + + K+ +LA C
Sbjct: 790 LDETQPEDSVHLVTWFRRVAASKDE-NAFKNAIDPNISLDDDTVASIEKVWELAGHCCAR 848
Query: 559 DPQQRPNMSSVVVALTTLT 577
+P QRP+M+ +V L++LT
Sbjct: 849 EPYQRPDMAHIVNVLSSLT 867
>At5g54590 serine/threonine-specific protein kinase-like protein
Length = 440
Score = 214 bits (544), Expect = 1e-55
Identities = 129/293 (44%), Positives = 174/293 (59%), Gaps = 25/293 (8%)
Query: 290 EFSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAIKKM---DMKATKEFLAELK 345
E+SY +L AT NF IGQG FG VY A+++ GE A+K + + KEF E+
Sbjct: 102 EYSYRDLQKATCNFTTL--IGQGAFGPVYKAQMSTGEIVAVKVLATDSKQGEKEFQTEVM 159
Query: 346 VLTRVHHVNLVRLIGYCVE-GSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSA 404
+L R+HH NLV LIGYC E G L+Y Y+ G+L HL S EPLSW +RV IALD A
Sbjct: 160 LLGRLHHRNLVNLIGYCAEKGQHMLIYVYMSKGSLASHLYSEKHEPLSWDLRVYIALDVA 219
Query: 405 RGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTK--LIDAGISSVPTVNMAGT 462
RGLEY+H+ VP IHRDIKS NILLD++ A+VADFGL++ ++D N+ GT
Sbjct: 220 RGLEYLHDGAVPPVIHRDIKSSNILLDQSMRARVADFGLSREEMVDK-----HAANIRGT 274
Query: 463 FGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQ 521
FGY+ PEY + + + K DVY FGV+L+ELI+ + +GL+ L E
Sbjct: 275 FGYLDPEYISTRTFTKKSDVYGFGVLLFELIAGR----------NPQQGLMELVELAAMN 324
Query: 522 PHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALT 574
G +++VD RL Y + V ++A A C + P++RPNM +V LT
Sbjct: 325 AEEKVGWEEIVDSRLDGRYDLQEVNEVAAFAYKCISRAPRKRPNMRDIVQVLT 377
>At1g70460 putative protein kinase
Length = 710
Score = 213 bits (543), Expect = 2e-55
Identities = 132/317 (41%), Positives = 186/317 (58%), Gaps = 16/317 (5%)
Query: 265 GNATYGTSDSASPANMIGIRVEKSGE--FSYEELANATNNFNMANKIGQGGFGEVYYAEL 322
G Y S SA + ++G SG+ F+YEEL + T F+ N +G+GGFG VY +L
Sbjct: 318 GGGGYTRSGSAPDSAVMG-----SGQTHFTYEELTDITEGFSKHNILGEGGFGCVYKGKL 372
Query: 323 N-GEKAAIKKMDM---KATKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNG 377
N G+ A+K++ + + +EF AE+++++RVHH +LV L+GYC+ S L+YEY+ N
Sbjct: 373 NDGKLVAVKQLKVGSGQGDREFKAEVEIISRVHHRHLVSLVGYCIADSERLLIYEYVPNQ 432
Query: 378 NLGQHLRSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAK 437
L HL L W+ RV+IA+ SA+GL Y+HE P IHRDIKS NILLD F A+
Sbjct: 433 TLEHHLHGKGRPVLEWARRVRIAIGSAKGLAYLHEDCHPKIIHRDIKSANILLDDEFEAQ 492
Query: 438 VADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYELISAKA 496
VADFGL KL D+ + V T + GTFGY+ PEYA G ++ + DV++FGVVL ELI+ +
Sbjct: 493 VADFGLAKLNDSTQTHVST-RVMGTFGYLAPEYAQSGKLTDRSDVFSFGVVLLELITGRK 551
Query: 497 AVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCT 556
V + G + LV + + +LVD RL +Y + VF+M + A C
Sbjct: 552 PVDQYQPLGEE--SLVEWARPLLHKAIETGDFSELVDRRLEKHYVENEVFRMIETAAACV 609
Query: 557 NSDPQQRPNMSSVVVAL 573
+RP M VV AL
Sbjct: 610 RHSGPKRPRMVQVVRAL 626
>At3g24400 protein kinase, putative
Length = 694
Score = 212 bits (540), Expect = 4e-55
Identities = 130/338 (38%), Positives = 192/338 (56%), Gaps = 17/338 (5%)
Query: 242 RKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPANMIGIRVEKSGEFSYEELANATN 301
R F+S SS + +D+ SP +G+ + + G F+YEEL+ ATN
Sbjct: 278 RPPHFMSSGSSGDYDSNYSDQ-------SVLPPPSPGLALGLGIYQ-GTFNYEELSRATN 329
Query: 302 NFNMANKIGQGGFGEVYYAEL-NGEKAAIKKM---DMKATKEFLAELKVLTRVHHVNLVR 357
F+ AN +GQGGFG V+ L NG++ A+K++ + +EF AE+ +++RVHH +LV
Sbjct: 330 GFSEANLLGQGGFGYVFKGMLRNGKEVAVKQLKEGSSQGEREFQAEVGIISRVHHRHLVA 389
Query: 358 LIGYCV-EGSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSARGLEYIHEHTVP 416
L+GYC+ + LVYE++ N L HL + WS R+KIA+ SA+GL Y+HE+ P
Sbjct: 390 LVGYCIADAQRLLVYEFVPNNTLEFHLHGKGRPTMEWSSRLKIAVGSAKGLSYLHENCNP 449
Query: 417 TYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYA-YGSV 475
IHRDIK+ NIL+D F AKVADFGL K+ + V T + GTFGY+ PEYA G +
Sbjct: 450 KIIHRDIKASNILIDFKFEAKVADFGLAKIASDTNTHVST-RVMGTFGYLAPEYASSGKL 508
Query: 476 SSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPR 535
+ K DV++FGVVL ELI+ + + + + AD LV + +Q + + +VD +
Sbjct: 509 TEKSDVFSFGVVLLELITGRRPIDV-NNVHAD-NSLVDWARPLLNQVSELGNFEVVVDKK 566
Query: 536 LGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVAL 573
L + Y + + +M A C S +RP M V L
Sbjct: 567 LNNEYDKEEMARMVACAAACVRSTAPRRPRMDQVARVL 604
>At1g20650 unknown protein
Length = 381
Score = 212 bits (539), Expect = 5e-55
Identities = 133/339 (39%), Positives = 196/339 (57%), Gaps = 26/339 (7%)
Query: 261 DEVSGNATYGTSDSASPANMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYA 320
+ +SG G +S P G R F+++ELA AT NF N +G+GGFG VY
Sbjct: 43 ESISGILVNGKVNSPIPGG--GAR-----SFTFKELAAATRNFREVNLLGEGGFGRVYKG 95
Query: 321 ELN-GEKAAIKKMD---MKATKEFLAELKVLTRVHHVNLVRLIGYCVEGSL-FLVYEYID 375
L+ G+ AIK+++ ++ +EF+ E+ +L+ +HH NLV LIGYC G LVYEY+
Sbjct: 96 RLDSGQVVAIKQLNPDGLQGNREFIVEVLMLSLLHHPNLVTLIGYCTSGDQRLLVYEYMP 155
Query: 376 NGNLGQHL--RSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKN 433
G+L HL S+ EPLSW+ R+KIA+ +ARG+EY+H P I+RD+KS NILLDK
Sbjct: 156 MGSLEDHLFDLESNQEPLSWNTRMKIAVGAARGIEYLHCTANPPVIYRDLKSANILLDKE 215
Query: 434 FCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYELI 492
F K++DFGL KL G + + + GT+GY PEYA G ++ K D+Y FGVVL ELI
Sbjct: 216 FSPKLSDFGLAKLGPVGDRTHVSTRVMGTYGYCAPEYAMSGKLTVKSDIYCFGVVLLELI 275
Query: 493 SAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKK---LVDPRLGDNYPIDHVFKMA 549
+ + A+ +G+ G + LV + +P+ ++ KK LVDP L YP +
Sbjct: 276 TGRKAIDLGQKQGE--QNLV-----TWSRPY-LKDQKKFGHLVDPSLRGKYPRRCLNYAI 327
Query: 550 QLAKVCTNSDPQQRPNMSSVVVALTTLTSTTEDWDITSI 588
+ +C N + RP + +VVAL L + + + ++
Sbjct: 328 AIIAMCLNEEAHYRPFIGDIVVALEYLAAQSRSHEARNV 366
>At5g56890 unknown protein
Length = 1113
Score = 211 bits (538), Expect = 7e-55
Identities = 136/384 (35%), Positives = 208/384 (53%), Gaps = 26/384 (6%)
Query: 225 ILLSFCIYVTYYRKKKIRKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPANMIGIR 284
+++ F ++ ++++ K+ L+ S + S + +S S S + I
Sbjct: 645 VIVWFLVFRRQRDRRRLSKRTPLARPSLPSLSKPSGSARSLTGSRFSSTSLSFESSIAPF 704
Query: 285 VEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAIKKM---DMKATKEF 340
+ F+ E+ ATNNF+ + +G+GGFG VY + G K A+K + D + ++EF
Sbjct: 705 TLSAKTFTASEIMKATNNFDESRVLGEGGFGRVYEGVFDDGTKVAVKVLKRDDQQGSREF 764
Query: 341 LAELKVLTRVHHVNLVRLIGYCVEG-SLFLVYEYIDNGNLGQHLRSSD--GEPLSWSIRV 397
LAE+++L+R+HH NLV LIG C+E + LVYE I NG++ HL D PL W R+
Sbjct: 765 LAEVEMLSRLHHRNLVNLIGICIEDRNRSLVYELIPNGSVESHLHGIDKASSPLDWDARL 824
Query: 398 KIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTK-LIDAGISSVPT 456
KIAL +ARGL Y+HE + P IHRD KS NILL+ +F KV+DFGL + +D + +
Sbjct: 825 KIALGAARGLAYLHEDSSPRVIHRDFKSSNILLENDFTPKVSDFGLARNALDDEDNRHIS 884
Query: 457 VNMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLF 515
+ GTFGY+ PEYA G + K DVY++GVVL EL++ + V M + G
Sbjct: 885 TRVMGTFGYVAPEYAMTGHLLVKSDVYSYGVVLLELLTGRKPVDMSQPPGQ--------- 935
Query: 516 EEVFDQPHPI----EGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVV 571
E + P EGL ++D LG D + K+A +A +C + RP M VV
Sbjct: 936 ENLVSWTRPFLTSAEGLAAIIDQSLGPEISFDSIAKVAAIASMCVQPEVSHRPFMGEVVQ 995
Query: 572 ALTTLTSTTEDW----DITSIFKN 591
AL +++ ++ +TSI K+
Sbjct: 996 ALKLVSNECDEAKELNSLTSISKD 1019
>At3g24550 protein kinase, putative
Length = 652
Score = 211 bits (538), Expect = 7e-55
Identities = 126/304 (41%), Positives = 179/304 (58%), Gaps = 12/304 (3%)
Query: 276 SPANMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDM 334
SP ++G F+YEEL+ ATN F+ AN +GQGGFG V+ L +G++ A+K++
Sbjct: 256 SPGLVLGF---SKSTFTYEELSRATNGFSEANLLGQGGFGYVHKGILPSGKEVAVKQLKA 312
Query: 335 ---KATKEFLAELKVLTRVHHVNLVRLIGYCVEG-SLFLVYEYIDNGNLGQHLRSSDGEP 390
+ +EF AE+++++RVHH +LV LIGYC+ G LVYE++ N NL HL
Sbjct: 313 GSGQGEREFQAEVEIISRVHHRHLVSLIGYCMAGVQRLLVYEFVPNNNLEFHLHGKGRPT 372
Query: 391 LSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAG 450
+ WS R+KIAL SA+GL Y+HE P IHRDIK+ NIL+D F AKVADFGL K+
Sbjct: 373 MEWSTRLKIALGSAKGLSYLHEDCNPKIIHRDIKASNILIDFKFEAKVADFGLAKIASDT 432
Query: 451 ISSVPTVNMAGTFGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLK 509
+ V T + GTFGY+ PEY A G ++ K DV++FGVVL ELI+ + V D
Sbjct: 433 NTHVST-RVMGTFGYLAPEYAASGKLTEKSDVFSFGVVLLELITGRRPVDANNVYVDD-- 489
Query: 510 GLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSV 569
LV + ++ + L D ++G+ Y + + +M A C ++RP MS +
Sbjct: 490 SLVDWARPLLNRASEEGDFEGLADSKMGNEYDREEMARMVACAAACVRHSARRRPRMSQI 549
Query: 570 VVAL 573
V AL
Sbjct: 550 VRAL 553
>At2g19230 putative receptor-like protein kinase
Length = 877
Score = 211 bits (536), Expect = 1e-54
Identities = 118/289 (40%), Positives = 168/289 (57%), Gaps = 15/289 (5%)
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIK---KMDMKATKEFLAELKVL 347
+ Y E+ TNNF +GQGGFG+VYY L GE+ AIK K + KEF AE+++L
Sbjct: 559 YKYSEIVEITNNFERV--LGQGGFGKVYYGVLRGEQVAIKMLSKSSAQGYKEFRAEVELL 616
Query: 348 TRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSARG 406
RVHH NL+ LIGYC EG + L+YEYI NG LG +L + LSW R++I+LD+A+G
Sbjct: 617 LRVHHKNLIALIGYCHEGDQMALIYEYIGNGTLGDYLSGKNSSILSWEERLQISLDAAQG 676
Query: 407 LEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYM 466
LEY+H P +HRD+K NIL+++ AK+ADFGL++ S + +AGT GY+
Sbjct: 677 LEYLHNGCKPPIVHRDVKPTNILINEKLQAKIADFGLSRSFTLEGDSQVSTEVAGTIGYL 736
Query: 467 PPE-YAYGSVSSKIDVYAFGVVLYELISAKAAVIMGE-DSGADLKGLVVLFEEVFDQPHP 524
PE Y+ S K DVY+FGVVL E+I+ + + + + V L D
Sbjct: 737 DPEHYSMQQFSEKSDVYSFGVVLLEVITGQPVISRSRTEENRHISDRVSLMLSKGD---- 792
Query: 525 IEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVAL 573
+K +VDP+LG+ + +K+ ++A C + + R MS VV L
Sbjct: 793 ---IKSIVDPKLGERFNAGLAWKITEVALACASESTKTRLTMSQVVAEL 838
>At1g79620 unknown protein
Length = 971
Score = 211 bits (536), Expect = 1e-54
Identities = 121/286 (42%), Positives = 173/286 (60%), Gaps = 12/286 (4%)
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDMKATK---EFLAELKV 346
FSYEEL TNNF++++++G GG+G+VY L +G AIK+ +T+ EF E+++
Sbjct: 626 FSYEELKKITNNFSVSSELGYGGYGKVYKGMLQDGHMVAIKRAQQGSTQGGLEFKTEIEL 685
Query: 347 LTRVHHVNLVRLIGYCVE-GSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSAR 405
L+RVHH NLV L+G+C E G LVYEY+ NG+L L G L W R+++AL SAR
Sbjct: 686 LSRVHHKNLVGLVGFCFEQGEQILVYEYMSNGSLKDSLTGRSGITLDWKRRLRVALGSAR 745
Query: 406 GLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGY 465
GL Y+HE P IHRD+KS NILLD+N AKVADFGL+KL+ + + GT GY
Sbjct: 746 GLAYLHELADPPIIHRDVKSTNILLDENLTAKVADFGLSKLVSDCTKGHVSTQVKGTLGY 805
Query: 466 MPPE-YAYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHP 524
+ PE Y ++ K DVY+FGVV+ ELI+AK + G+ ++K ++ ++ F
Sbjct: 806 LDPEYYTTQKLTEKSDVYSFGVVMMELITAKQPIEKGKYIVREIKLVMNKSDDDF----- 860
Query: 525 IEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVV 570
GL+ +D L D + + + +LA C + +RP MS VV
Sbjct: 861 -YGLRDKMDRSLRDVGTLPELGRYMELALKCVDETADERPTMSEVV 905
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,438,293
Number of Sequences: 26719
Number of extensions: 592730
Number of successful extensions: 5753
Number of sequences better than 10.0: 1007
Number of HSP's better than 10.0 without gapping: 829
Number of HSP's successfully gapped in prelim test: 178
Number of HSP's that attempted gapping in prelim test: 1561
Number of HSP's gapped (non-prelim): 1157
length of query: 601
length of database: 11,318,596
effective HSP length: 105
effective length of query: 496
effective length of database: 8,513,101
effective search space: 4222498096
effective search space used: 4222498096
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC123570.7


