

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
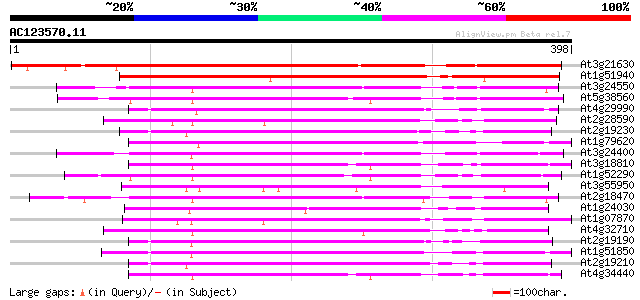
Query= AC123570.11 + phase: 0
(398 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g21630 receptor like protein kinase 386 e-107
At1g51940 protein kinase like protein 240 1e-63
At3g24550 protein kinase, putative 213 2e-55
At5g38560 putative protein 211 7e-55
At4g29990 serine/threonine-specific receptor protein kinase LRRPK 208 5e-54
At2g28590 putative protein kinase 206 1e-53
At2g19230 putative receptor-like protein kinase 205 4e-53
At1g79620 unknown protein 204 5e-53
At3g24400 protein kinase, putative 204 7e-53
At3g18810 protein kinase, putative 202 3e-52
At1g52290 protein kinase, putative 202 3e-52
At3g55950 receptor kinase - like protein 201 4e-52
At2g18470 putative protein kinase 201 4e-52
At1g24030 protein kinase like protein 201 6e-52
At1g07870 putative protein kinase (At1g07870) 201 8e-52
At4g32710 putative protein kinase 200 1e-51
At2g19190 putative receptor-like protein kinase 200 1e-51
At1g51850 Putative protein kinase 200 1e-51
At2g19210 putative receptor-like protein kinase 199 2e-51
At4g34440 serine/threonine protein kinase - like 199 2e-51
>At3g21630 receptor like protein kinase
Length = 617
Score = 386 bits (991), Expect = e-107
Identities = 219/432 (50%), Positives = 277/432 (63%), Gaps = 61/432 (14%)
Query: 2 IEIELLKRYN--VNFSQGSGLVYIRGKDGNGLYVPLPPK--------------------- 38
+ ++L+RYN VNF+ G+G+VY+ G+D NG + P
Sbjct: 187 VSADILQRYNPGVNFNSGNGIVYVPGRDPNGAFPPFKSSKQDGVGAGVIAGIVIGVIVAL 246
Query: 39 --------YFRKKNGEEESKFPPEDSMTPSTK-DVDKDTNDDNGS----------KYIWV 79
Y +KN + F S+ STK D T+ +G I V
Sbjct: 247 LLILFIVYYAYRKNKSKGDSF--SSSIPLSTKADHASSTSLQSGGLGGAGVSPGIAAISV 304
Query: 80 DKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELK 139
DKS EFS EELA ATDNF+L+ KIGQGGFG VYY ELRG+K AIKKM M+A+++FL+ELK
Sbjct: 305 DKSVEFSLEELAKATDNFNLSFKIGQGGFGAVYYAELRGEKAAIKKMDMEASKQFLAELK 364
Query: 140 VLTSVHHWNLVHLIGYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVAR 199
VLT VHH NLV LIGYCVEG LFLVYEY+ENGNL QHLH S +EP+ + R++IALD AR
Sbjct: 365 VLTRVHHVNLVRLIGYCVEGSLFLVYEYVENGNLGQHLHGSGREPLPWTKRVQIALDSAR 424
Query: 200 GLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGY 259
GLEYIH+H+VPVY+HRDIKS NIL+++ F KVADFGLTKLT+ SA T GTFGY
Sbjct: 425 GLEYIHEHTVPVYVHRDIKSANILIDQKFRAKVADFGLTKLTEVGGSA--TRGAMGTFGY 482
Query: 260 MPPENAYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSL 319
M PE YG +S K+DVYAFGVVLYELISAK AV+K+ E++ E++ L
Sbjct: 483 MAPETVYGEVSAKVDVYAFGVVLYELISAKGAVVKM--------------TEAVGEFRGL 528
Query: 320 VALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV 379
V +F+E +T D + LRK++DPRLG +Y DS+ KMA+L KAC + + RP+MR +V
Sbjct: 529 VGVFEESFKET-DKEEALRKIIDPRLGDSYPFDSVYKMAELGKACTQENAQLRPSMRYIV 587
Query: 380 VSLMELNSSIDN 391
V+L L SS N
Sbjct: 588 VALSTLFSSTGN 599
>At1g51940 protein kinase like protein
Length = 601
Score = 240 bits (612), Expect = 1e-63
Identities = 139/321 (43%), Positives = 200/321 (62%), Gaps = 20/321 (6%)
Query: 79 VDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSEL 138
++K F+YEE+ ATD FS + +G G +G VY+G LR Q++A+K+M T+EF +E+
Sbjct: 273 IEKPMVFTYEEIRAATDEFSDSNLLGHGNYGSVYFGLLREQEVAVKRMTATKTKEFAAEM 332
Query: 139 KVLTSVHHWNLVHLIGYCVE-GFLFLVYEYMENGNLSQHLHNSEKE---PMTLSTRMKIA 194
KVL VHH NLV LIGY LF+VYEY+ G L HLH+ + + P++ R +IA
Sbjct: 333 KVLCKVHHSNLVELIGYAATVDELFVVYEYVRKGMLKSHLHDPQSKGNTPLSWIMRNQIA 392
Query: 195 LDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTV-HV 253
LD ARGLEYIH+H+ Y+HRDIK+ NILL+E F K++DFGL KL + + +V V
Sbjct: 393 LDAARGLEYIHEHTKTHYVHRDIKTSNILLDEAFRAKISDFGLAKLVEKTGEGEISVTKV 452
Query: 254 AGTFGYMPPEN-AYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNES 312
GT+GY+ PE + G + K D+YAFGVVL+E+IS + AVI+ + I T
Sbjct: 453 VGTYGYLAPEYLSDGLATSKSDIYAFGVVLFEIISGREAVIRTE---------AIGTKN- 502
Query: 313 IDEYKSLVALFDEVMNQTGDCID--DLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPK 370
E + L ++ V+ + D ++ L++ VDP + Y D + K+A LAK C++ DP
Sbjct: 503 -PERRPLASIMLAVLKNSPDSMNMSSLKEFVDPNMMDLYPHDCLFKIATLAKQCVDDDPI 561
Query: 371 QRPTMRDVVVSLME-LNSSID 390
RP M+ VV+SL + L SSI+
Sbjct: 562 LRPNMKQVVISLSQILLSSIE 582
>At3g24550 protein kinase, putative
Length = 652
Score = 213 bits (541), Expect = 2e-55
Identities = 138/368 (37%), Positives = 200/368 (53%), Gaps = 45/368 (12%)
Query: 34 PLPPKYFRKKNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPEFSYEELANA 93
P PP + G + S P +P + KS F+YEEL+ A
Sbjct: 232 PPPPAFMSSSGGSDYSDLPVLPPPSPGL--------------VLGFSKST-FTYEELSRA 276
Query: 94 TDNFSLAKKIGQGGFGEVYYGEL-RGQKIAIKKMKM---QATREFLSELKVLTSVHHWNL 149
T+ FS A +GQGGFG V+ G L G+++A+K++K Q REF +E+++++ VHH +L
Sbjct: 277 TNGFSEANLLGQGGFGYVHKGILPSGKEVAVKQLKAGSGQGEREFQAEVEIISRVHHRHL 336
Query: 150 VHLIGYCVEGFL-FLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDHS 208
V LIGYC+ G LVYE++ N NL HLH + M STR+KIAL A+GL Y+H+
Sbjct: 337 VSLIGYCMAGVQRLLVYEFVPNNNLEFHLHGKGRPTMEWSTRLKIALGSAKGLSYLHEDC 396
Query: 209 VPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPE-NAYG 267
P IHRDIK+ NIL++ F KVADFGL K+ N+ +T V GTFGY+ PE A G
Sbjct: 397 NPKIIHRDIKASNILIDFKFEAKVADFGLAKIASDTNTHVST-RVMGTFGYLAPEYAASG 455
Query: 268 RISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVM 327
+++ K DV++FGVVL ELI+ + V N +D+ SLV ++
Sbjct: 456 KLTEKSDVFSFGVVLLELITGRRPV--------------DANNVYVDD--SLVDWARPLL 499
Query: 328 NQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV------VS 381
N+ + D L D ++G Y + +++M A AC+ ++RP M +V VS
Sbjct: 500 NRASE-EGDFEGLADSKMGNEYDREEMARMVACAAACVRHSARRRPRMSQIVRALEGNVS 558
Query: 382 LMELNSSI 389
L +LN +
Sbjct: 559 LSDLNEGM 566
>At5g38560 putative protein
Length = 681
Score = 211 bits (536), Expect = 7e-55
Identities = 136/377 (36%), Positives = 202/377 (53%), Gaps = 45/377 (11%)
Query: 35 LPPKYFRKKNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPE--------FS 86
+PP + G + F S P + +GS Y++ FS
Sbjct: 276 MPPSAYSSPQGSDVVLFNSRSSAPPKMRS-------HSGSDYMYASSDSGMVSNQRSWFS 328
Query: 87 YEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKMKM---QATREFLSELKVLT 142
Y+EL+ T FS +G+GGFG VY G L G+++A+K++K+ Q REF +E+++++
Sbjct: 329 YDELSQVTSGFSEKNLLGEGGFGCVYKGVLSDGREVAVKQLKIGGSQGEREFKAEVEIIS 388
Query: 143 SVHHWNLVHLIGYCV-EGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGL 201
VHH +LV L+GYC+ E LVY+Y+ N L HLH + MT TR+++A ARG+
Sbjct: 389 RVHHRHLVTLVGYCISEQHRLLVYDYVPNNTLHYHLHAPGRPVMTWETRVRVAAGAARGI 448
Query: 202 EYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVA----GTF 257
Y+H+ P IHRDIKS NILL+ +F VADFGL K+ A D HV+ GTF
Sbjct: 449 AYLHEDCHPRIIHRDIKSSNILLDNSFEALVADFGLAKI---AQELDLNTHVSTRVMGTF 505
Query: 258 GYMPPENA-YGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEY 316
GYM PE A G++S K DVY++GV+L ELI+ + V T++ + +
Sbjct: 506 GYMAPEYATSGKLSEKADVYSYGVILLELITGRKPV---------------DTSQPLGD- 549
Query: 317 KSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMR 376
+SLV ++ Q + ++ +LVDPRLG N+ + +M + A AC+ +RP M
Sbjct: 550 ESLVEWARPLLGQAIE-NEEFDELVDPRLGKNFIPGEMFRMVEAAAACVRHSAAKRPKMS 608
Query: 377 DVVVSLMELNSSIDNKN 393
VV +L L + D N
Sbjct: 609 QVVRALDTLEEATDITN 625
>At4g29990 serine/threonine-specific receptor protein kinase LRRPK
Length = 876
Score = 208 bits (529), Expect = 5e-54
Identities = 126/310 (40%), Positives = 182/310 (58%), Gaps = 32/310 (10%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQAT---REFLSELKVL 141
F Y E+ N T+NF + +G+GGFG+VY+G L G ++A+K + ++T +EF +E+++L
Sbjct: 564 FIYSEVVNITNNFE--RVLGKGGFGKVYHGFLNGDQVAVKILSEESTQGYKEFRAEVELL 621
Query: 142 TSVHHWNLVHLIGYCVE-GFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARG 200
VHH NL LIGYC E + L+YEYM NGNL +L ++ R++I+LD A+G
Sbjct: 622 MRVHHTNLTSLIGYCNEDNHMALIYEYMANGNLGDYLSGKSSLILSWEERLQISLDAAQG 681
Query: 201 LEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYM 260
LEY+H P +HRD+K NILLNEN K+ADFGL++ S+ + VAGT GY+
Sbjct: 682 LEYLHYGCKPPIVHRDVKPANILLNENLQAKIADFGLSRSFPVEGSSQVSTVVAGTIGYL 741
Query: 261 PPE-NAYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSL 319
PE A +++ K DVY+FGVVL E+I+ K A+ +TE
Sbjct: 742 DPEYYATRQMNEKSDVYSFGVVLLEVITGKPAIWH-SRTE-------------------S 781
Query: 320 VALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV 379
V L D+V + + D++ +VD RLG + + S K+ +LA AC + +QRPTM VV
Sbjct: 782 VHLSDQVGSMLAN--GDIKGIVDQRLGDRFEVGSAWKITELALACASESSEQRPTMSQVV 839
Query: 380 VSLMELNSSI 389
MEL SI
Sbjct: 840 ---MELKQSI 846
>At2g28590 putative protein kinase
Length = 424
Score = 206 bits (525), Expect = 1e-53
Identities = 128/331 (38%), Positives = 187/331 (55%), Gaps = 29/331 (8%)
Query: 67 DTNDDNGSKYIWVDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYG--ELRGQKIAIK 124
D D N + V K+ F++EEL+ +T NF +G+GGFG+VY G E Q +AIK
Sbjct: 68 DAKDTNVEDEVIVKKAQTFTFEELSVSTGNFKSDCFLGEGGFGKVYKGFIEKINQVVAIK 127
Query: 125 KMKM---QATREFLSELKVLTSVHHWNLVHLIGYCVEGFL-FLVYEYMENGNLSQHLHN- 179
++ Q REF+ E+ L+ H NLV LIG+C EG LVYEYM G+L HLH+
Sbjct: 128 QLDRNGAQGIREFVVEVLTLSLADHPNLVKLIGFCAEGVQRLLVYEYMPLGSLDNHLHDL 187
Query: 180 -SEKEPMTLSTRMKIALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLT 238
S K P+ +TRMKIA ARGLEY+HD P I+RD+K NIL++E + K++DFGL
Sbjct: 188 PSGKNPLAWNTRMKIAAGAARGLEYLHDTMKPPVIYRDLKCSNILIDEGYHAKLSDFGLA 247
Query: 239 KLTDAANSADNTVHVAGTFGYMPPENAY-GRISRKIDVYAFGVVLYELISAKAAVIKIDK 297
K+ + + V GT+GY P+ A G+++ K DVY+FGVVL ELI+ + A
Sbjct: 248 KVGPRGSETHVSTRVMGTYGYCAPDYALTGQLTFKSDVYSFGVVLLELITGRKA------ 301
Query: 298 TEFEFKSLEIKTNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKM 357
+ + + ++S+ E+ + LF + N +K+VDP L +Y + + +
Sbjct: 302 ----YDNTRTRNHQSLVEWAN--PLFKDRKN--------FKKMVDPLLEGDYPVRGLYQA 347
Query: 358 AKLAKACINRDPKQRPTMRDVVVSLMELNSS 388
+A C+ P RP + DVV++L L SS
Sbjct: 348 LAIAAMCVQEQPSMRPVIADVVMALDHLASS 378
>At2g19230 putative receptor-like protein kinase
Length = 877
Score = 205 bits (521), Expect = 4e-53
Identities = 122/311 (39%), Positives = 178/311 (57%), Gaps = 28/311 (9%)
Query: 79 VDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIK---KMKMQATREFL 135
+D + Y E+ T+NF + +GQGGFG+VYYG LRG+++AIK K Q +EF
Sbjct: 553 LDTKRYYKYSEIVEITNNFE--RVLGQGGFGKVYYGVLRGEQVAIKMLSKSSAQGYKEFR 610
Query: 136 SELKVLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIA 194
+E+++L VHH NL+ LIGYC EG + L+YEY+ NG L +L ++ R++I+
Sbjct: 611 AEVELLLRVHHKNLIALIGYCHEGDQMALIYEYIGNGTLGDYLSGKNSSILSWEERLQIS 670
Query: 195 LDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVA 254
LD A+GLEY+H+ P +HRD+K NIL+NE K+ADFGL++ + + VA
Sbjct: 671 LDAAQGLEYLHNGCKPPIVHRDVKPTNILINEKLQAKIADFGLSRSFTLEGDSQVSTEVA 730
Query: 255 GTFGYMPPEN-AYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESI 313
GT GY+ PE+ + + S K DVY+FGVVL E+I+ + VI +TE N I
Sbjct: 731 GTIGYLDPEHYSMQQFSEKSDVYSFGVVLLEVITGQ-PVISRSRTE---------ENRHI 780
Query: 314 DEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRP 373
+ SL M G D++ +VDP+LG ++ K+ ++A AC + K R
Sbjct: 781 SDRVSL-------MLSKG----DIKSIVDPKLGERFNAGLAWKITEVALACASESTKTRL 829
Query: 374 TMRDVVVSLME 384
TM VV L E
Sbjct: 830 TMSQVVAELKE 840
>At1g79620 unknown protein
Length = 971
Score = 204 bits (520), Expect = 5e-53
Identities = 121/320 (37%), Positives = 184/320 (56%), Gaps = 30/320 (9%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKMKMQATR---EFLSELKV 140
FSYEEL T+NFS++ ++G GG+G+VY G L+ G +AIK+ + +T+ EF +E+++
Sbjct: 626 FSYEELKKITNNFSVSSELGYGGYGKVYKGMLQDGHMVAIKRAQQGSTQGGLEFKTEIEL 685
Query: 141 LTSVHHWNLVHLIGYCVE-GFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVAR 199
L+ VHH NLV L+G+C E G LVYEYM NG+L L + R+++AL AR
Sbjct: 686 LSRVHHKNLVGLVGFCFEQGEQILVYEYMSNGSLKDSLTGRSGITLDWKRRLRVALGSAR 745
Query: 200 GLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGY 259
GL Y+H+ + P IHRD+KS NILL+EN T KVADFGL+KL + V GT GY
Sbjct: 746 GLAYLHELADPPIIHRDVKSTNILLDENLTAKVADFGLSKLVSDCTKGHVSTQVKGTLGY 805
Query: 260 MPPE-NAYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKS 318
+ PE +++ K DVY+FGVV+ ELI+AK I+K ++ + +++ N+S D++
Sbjct: 806 LDPEYYTTQKLTEKSDVYSFGVVMMELITAKQ---PIEKGKYIVREIKLVMNKSDDDFYG 862
Query: 319 LVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDV 378
L D + G ++ + + +LA C++ +RPTM +V
Sbjct: 863 LRDKMDRSLRDVG------------------TLPELGRYMELALKCVDETADERPTMSEV 904
Query: 379 VVSLMELNSSIDNKNSTGSA 398
V E+ I N ++ S+
Sbjct: 905 V---KEIEIIIQNSGASSSS 921
>At3g24400 protein kinase, putative
Length = 694
Score = 204 bits (519), Expect = 7e-53
Identities = 129/367 (35%), Positives = 202/367 (54%), Gaps = 36/367 (9%)
Query: 34 PLPPKYFRK-KNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPEFSYEELAN 92
P PP + +G+ +S + + + P + + G+ F+YEEL+
Sbjct: 277 PRPPHFMSSGSSGDYDSNYSDQSVLPPPSPGLALGLGIYQGT----------FNYEELSR 326
Query: 93 ATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKMK---MQATREFLSELKVLTSVHHWN 148
AT+ FS A +GQGGFG V+ G LR G+++A+K++K Q REF +E+ +++ VHH +
Sbjct: 327 ATNGFSEANLLGQGGFGYVFKGMLRNGKEVAVKQLKEGSSQGEREFQAEVGIISRVHHRH 386
Query: 149 LVHLIGYCV-EGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDH 207
LV L+GYC+ + LVYE++ N L HLH + M S+R+KIA+ A+GL Y+H++
Sbjct: 387 LVALVGYCIADAQRLLVYEFVPNNTLEFHLHGKGRPTMEWSSRLKIAVGSAKGLSYLHEN 446
Query: 208 SVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPENA-Y 266
P IHRDIK+ NIL++ F KVADFGL K+ N+ +T V GTFGY+ PE A
Sbjct: 447 CNPKIIHRDIKASNILIDFKFEAKVADFGLAKIASDTNTHVST-RVMGTFGYLAPEYASS 505
Query: 267 GRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEV 326
G+++ K DV++FGVVL ELI+ + + ++ SLV +
Sbjct: 506 GKLTEKSDVFSFGVVLLELITGRRPI----------------DVNNVHADNSLVDWARPL 549
Query: 327 MNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELN 386
+NQ + + + +VD +L Y + +++M A AC+ +RP M D V ++E N
Sbjct: 550 LNQVSE-LGNFEVVVDKKLNNEYDKEEMARMVACAAACVRSTAPRRPRM-DQVARVLEGN 607
Query: 387 SSIDNKN 393
S + N
Sbjct: 608 ISPSDLN 614
>At3g18810 protein kinase, putative
Length = 700
Score = 202 bits (514), Expect = 3e-52
Identities = 128/324 (39%), Positives = 189/324 (57%), Gaps = 34/324 (10%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGEL-RGQKIAIKKMKM---QATREFLSELKV 140
F+Y+ELA AT FS ++ +GQGGFG V+ G L G++IA+K +K Q REF +E+ +
Sbjct: 325 FTYDELAAATQGFSQSRLLGQGGFGYVHKGILPNGKEIAVKSLKAGSGQGEREFQAEVDI 384
Query: 141 LTSVHHWNLVHLIGYCVEGFL-FLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVAR 199
++ VHH LV L+GYC+ G LVYE++ N L HLH + + TR+KIAL A+
Sbjct: 385 ISRVHHRFLVSLVGYCIAGGQRMLVYEFLPNDTLEFHLHGKSGKVLDWPTRLKIALGSAK 444
Query: 200 GLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVA----G 255
GL Y+H+ P IHRDIK+ NILL+E+F KVADFGL KL S DN HV+ G
Sbjct: 445 GLAYLHEDCHPRIIHRDIKASNILLDESFEAKVADFGLAKL-----SQDNVTHVSTRIMG 499
Query: 256 TFGYMPPENA-YGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESID 314
TFGY+ PE A G+++ + DV++FGV+L EL++ + V L + +S+
Sbjct: 500 TFGYLAPEYASSGKLTDRSDVFSFGVMLLELVTGRRPV-----------DLTGEMEDSLV 548
Query: 315 EYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPT 374
++ + L N D D +LVDPRL Y +++M A A + ++RP
Sbjct: 549 DWARPICL-----NAAQD--GDYSELVDPRLENQYEPHEMAQMVACAAAAVRHSARRRPK 601
Query: 375 MRDVVVSLMELNSSIDNKNSTGSA 398
M +V +L E ++++D+ + G A
Sbjct: 602 MSQIVRAL-EGDATLDDLSEGGKA 624
>At1g52290 protein kinase, putative
Length = 509
Score = 202 bits (513), Expect = 3e-52
Identities = 137/365 (37%), Positives = 199/365 (53%), Gaps = 36/365 (9%)
Query: 40 FRKKNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPE---FSYEELANATDN 96
F K+ + K ED +D D DD+ + W F+YE+L+ AT N
Sbjct: 84 FYKRKKRKLKKKKKEDIEASINRD-SLDPKDDSNNLQQWSSSEIGQNLFTYEDLSKATSN 142
Query: 97 FSLAKKIGQGGFGEVYYGEL-RGQKIAIKKMKM---QATREFLSELKVLTSVHHWNLVHL 152
FS +GQGGFG V+ G L G +AIK++K Q REF +E++ ++ VHH +LV L
Sbjct: 143 FSNTNLLGQGGFGYVHRGVLVDGTLVAIKQLKSGSGQGEREFQAEIQTISRVHHRHLVSL 202
Query: 153 IGYCVEGFL-FLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDHSVPV 211
+GYC+ G LVYE++ N L HLH E+ M S RMKIAL A+GL Y+H+ P
Sbjct: 203 LGYCITGAQRLLVYEFVPNKTLEFHLHEKERPVMEWSKRMKIALGAAKGLAYLHEDCNPK 262
Query: 212 YIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVA----GTFGYMPPENA-Y 266
IHRD+K+ NIL+++++ K+ADFGL A +S D HV+ GTFGY+ PE A
Sbjct: 263 TIHRDVKAANILIDDSYEAKLADFGL-----ARSSLDTDTHVSTRIMGTFGYLAPEYASS 317
Query: 267 GRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEV 326
G+++ K DV++ GVVL ELI+ + V KS ++SI ++ L +
Sbjct: 318 GKLTEKSDVFSIGVVLLELITGRRPV---------DKSQPFADDDSIVDWAK--PLMIQA 366
Query: 327 MNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELN 386
+N + LVDPRL ++ I+ +++M A A + K+RP M +V E N
Sbjct: 367 LND-----GNFDGLVDPRLENDFDINEMTRMVACAAASVRHSAKRRPKMSQ-IVRAFEGN 420
Query: 387 SSIDN 391
SID+
Sbjct: 421 ISIDD 425
>At3g55950 receptor kinase - like protein
Length = 814
Score = 201 bits (512), Expect = 4e-52
Identities = 131/326 (40%), Positives = 186/326 (56%), Gaps = 37/326 (11%)
Query: 80 DKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIK------KMKMQATR 132
DK+ EFS+ ELA+AT NFSL KIG G FG VY G+L G+++AIK KMK +
Sbjct: 479 DKAEEFSFSELASATGNFSLENKIGSGSFGVVYRGKLNDGREVAIKRGEVNAKMKKFQEK 538
Query: 133 E--FLSELKVLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLH---NSEKEPMT 186
E F SE+ L+ +HH +LV L+GYC E LVY+YM+NG L HLH N EK
Sbjct: 539 ETAFDSEIAFLSRLHHKHLVRLVGYCEEREEKLLVYDYMKNGALYDHLHDKNNVEKHSSL 598
Query: 187 LST---RMKIALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDA 243
+++ R+KIALD ARG+EY+H+++VP IHRDIKS NILL+ N+ +V+DFGL+ +
Sbjct: 599 INSWKMRIKIALDAARGIEYLHNYAVPPIIHRDIKSSNILLDSNWVARVSDFGLSLMGPV 658
Query: 244 A----NSADNTVHVAGTFGYMPPE-NAYGRISRKIDVYAFGVVLYELISAKAAVIKIDKT 298
N AGT GY+ PE + ++ K DVY GVVL EL++ K A+
Sbjct: 659 LGKDHNPYQRPTKAAGTVGYIDPEYYSLNVLTDKSDVYGLGVVLLELLTGKRAI------ 712
Query: 299 EFEFKSLEIKTNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNY--SIDSISK 356
+ N ++E + V + + D+L ++DPR+G D++
Sbjct: 713 --------FRNNGDVEEEEGCVPVHLVDYSVPAITADELSTILDPRVGSPELGEGDAVEL 764
Query: 357 MAKLAKACINRDPKQRPTMRDVVVSL 382
+A A C+N + + RPTM D+V +L
Sbjct: 765 VAYTAMHCVNAEGRNRPTMTDIVGNL 790
>At2g18470 putative protein kinase
Length = 633
Score = 201 bits (512), Expect = 4e-52
Identities = 146/392 (37%), Positives = 203/392 (51%), Gaps = 53/392 (13%)
Query: 15 SQGSGLVYIRGKDGNGLYVPLPPKYFRKKNGEEESKF--PPEDSMTPSTKDVDKDTNDDN 72
SQG G G G P PP+ +GE+ S + P + P + + N
Sbjct: 214 SQGQNQQSTGGWGGGGPSPPPPPRM--PTSGEDSSMYSGPSRPVLPPPSPALALGFNKST 271
Query: 73 GSKYIWVDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGEL-RGQKIAIKKMKM--- 128
F+Y+ELA AT F+ A +GQGGFG V+ G L G+++A+K +K
Sbjct: 272 ------------FTYQELAAATGGFTDANLLGQGGFGYVHKGVLPSGKEVAVKSLKAGSG 319
Query: 129 QATREFLSELKVLTSVHHWNLVHLIGYCV-EGFLFLVYEYMENGNLSQHLHNSEKEPMTL 187
Q REF +E+ +++ VHH LV L+GYC+ +G LVYE++ N L HLH M
Sbjct: 320 QGEREFQAEVDIISRVHHRYLVSLVGYCIADGQRMLVYEFVPNKTLEYHLHGKNLPVMEF 379
Query: 188 STRMKIALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSA 247
STR++IAL A+GL Y+H+ P IHRDIKS NILL+ NF VADFGL KLT N+
Sbjct: 380 STRLRIALGAAKGLAYLHEDCHPRIIHRDIKSANILLDFNFDAMVADFGLAKLTSDNNTH 439
Query: 248 DNTVHVAGTFGYMPPENA-YGRISRKIDVYAFGVVLYELISAKAAV---IKIDKTEFEFK 303
+T V GTFGY+ PE A G+++ K DV+++GV+L ELI+ K V I +D T
Sbjct: 440 VST-RVMGTFGYLAPEYASSGKLTEKSDVFSYGVMLLELITGKRPVDNSITMDDT----- 493
Query: 304 SLEIKTNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKA 363
+D + L+A E N +L D RL NY+ +++M A A
Sbjct: 494 --------LVDWARPLMARALEDGN--------FNELADARLEGNYNPQEMARMVTCAAA 537
Query: 364 CINRDPKQRPTMRDVV------VSLMELNSSI 389
I ++RP M +V VSL LN +
Sbjct: 538 SIRHSGRKRPKMSQIVRALEGEVSLDALNEGV 569
>At1g24030 protein kinase like protein
Length = 375
Score = 201 bits (511), Expect = 6e-52
Identities = 125/314 (39%), Positives = 181/314 (56%), Gaps = 34/314 (10%)
Query: 82 SPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKM------KMQATREF 134
S ++ +E+ AT +FS +G+GGFG VY G L+ G+ +AIKKM K REF
Sbjct: 61 SSVYTLKEMEEATSSFSDENLLGKGGFGRVYQGTLKTGEVVAIKKMDLPTFKKADGEREF 120
Query: 135 LSELKVLTSVHHWNLVHLIGYCVEG-FLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKI 193
E+ +L+ + H NLV LIGYC +G FLVYEYM+NGNL HL+ ++ ++ R++I
Sbjct: 121 RVEVDILSRLDHPNLVSLIGYCADGKHRFLVYEYMQNGNLQDHLNGIKEAKISWPIRLRI 180
Query: 194 ALDVARGLEYIHDHS---VPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNT 250
AL A+GL Y+H S +P+ +HRD KS N+LL+ N+ K++DFGL KL T
Sbjct: 181 ALGAAKGLAYLHSSSSVGIPI-VHRDFKSTNVLLDSNYNAKISDFGLAKLMPEGKDTCVT 239
Query: 251 VHVAGTFGYMPPE-NAYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKT 309
V GTFGY PE + G+++ + D+YAFGVVL EL++ + AV L
Sbjct: 240 ARVLGTFGYFDPEYTSTGKLTLQSDIYAFGVVLLELLTGRRAV-----------DLTQGP 288
Query: 310 NESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYN-YSIDSISKMAKLAKACINRD 368
NE ++LV ++N LRK++D L N YS+++I+ A LA CI +
Sbjct: 289 NE-----QNLVLQVRNILNDR----KKLRKVIDVELPRNSYSMEAITMFADLASRCIRIE 339
Query: 369 PKQRPTMRDVVVSL 382
K+RP++ D V L
Sbjct: 340 SKERPSVMDCVKEL 353
>At1g07870 putative protein kinase (At1g07870)
Length = 423
Score = 201 bits (510), Expect = 8e-52
Identities = 128/327 (39%), Positives = 184/327 (56%), Gaps = 29/327 (8%)
Query: 81 KSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELR--GQKIAIKKMK---MQATREFL 135
K+ F+++ELA AT NF +G+GGFG+V+ G + Q +AIK++ +Q REF+
Sbjct: 87 KAQTFTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRNGVQGIREFV 146
Query: 136 SELKVLTSVHHWNLVHLIGYCVEGFL-FLVYEYMENGNLSQHLH--NSEKEPMTLSTRMK 192
E+ L+ H NLV LIG+C EG LVYEYM G+L HLH S K+P+ +TRMK
Sbjct: 147 VEVLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDWNTRMK 206
Query: 193 IALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVH 252
IA ARGLEY+HD P I+RD+K NILL E++ K++DFGL K+ + + +
Sbjct: 207 IAAGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKTHVSTR 266
Query: 253 VAGTFGYMPPENAY-GRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNE 311
V GT+GY P+ A G+++ K D+Y+FGVVL ELI+ + A ID T KT +
Sbjct: 267 VMGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKA---IDNT---------KTRK 314
Query: 312 SIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQ 371
+ LF + N K+VDP L Y + + + ++ C+ P
Sbjct: 315 DQNLVGWARPLFKDRRN--------FPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTM 366
Query: 372 RPTMRDVVVSLMELNSSIDNKNSTGSA 398
RP + DVV++L L SS + NS S+
Sbjct: 367 RPVVSDVVLALNFLASSKYDPNSPSSS 393
>At4g32710 putative protein kinase
Length = 731
Score = 200 bits (509), Expect = 1e-51
Identities = 128/324 (39%), Positives = 183/324 (55%), Gaps = 25/324 (7%)
Query: 67 DTNDDNGSKYIWVDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKK 125
DT ++N S FSYEEL+ AT FS +G+GGFG V+ G L+ G ++A+K+
Sbjct: 359 DTKENNSVAKNISMPSGMFSYEELSKATGGFSEENLLGEGGFGYVHKGVLKNGTEVAVKQ 418
Query: 126 MKM---QATREFLSELKVLTSVHHWNLVHLIGYCVEGFL-FLVYEYMENGNLSQHLHNSE 181
+K+ Q REF +E+ ++ VHH +LV L+GYCV G LVYE++ L HLH +
Sbjct: 419 LKIGSYQGEREFQAEVDTISRVHHKHLVSLVGYCVNGDKRLLVYEFVPKDTLEFHLHENR 478
Query: 182 KEPMTLSTRMKIALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLT 241
+ R++IA+ A+GL Y+H+ P IHRDIK+ NILL+ F KV+DFGL K
Sbjct: 479 GSVLEWEMRLRIAVGAAKGLAYLHEDCSPTIIHRDIKAANILLDSKFEAKVSDFGLAKFF 538
Query: 242 DAANSADN--TVHVAGTFGYMPPENA-YGRISRKIDVYAFGVVLYELISAKAAVIKIDKT 298
NS+ + V GTFGYM PE A G+++ K DVY+FGVVL ELI+ + ++ D +
Sbjct: 539 SDTNSSFTHISTRVVGTFGYMAPEYASSGKVTDKSDVYSFGVVLLELITGRPSIFAKDSS 598
Query: 299 EFEFKSLEIKTNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMA 358
TN+S+ ++ L + + +G+ D LVD RL NY ++ MA
Sbjct: 599 ----------TNQSLVDWAR--PLLTKAI--SGESFD---FLVDSRLEKNYDTTQMANMA 641
Query: 359 KLAKACINRDPKQRPTMRDVVVSL 382
A ACI + RP M VV +L
Sbjct: 642 ACAAACIRQSAWLRPRMSQVVRAL 665
>At2g19190 putative receptor-like protein kinase
Length = 876
Score = 200 bits (509), Expect = 1e-51
Identities = 120/306 (39%), Positives = 175/306 (56%), Gaps = 29/306 (9%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMK---MQATREFLSELKVL 141
F Y E+ N T+NF + IG+GGFG+VY+G + G+++A+K + Q +EF +E+ +L
Sbjct: 564 FKYSEVVNITNNFE--RVIGKGGFGKVYHGVINGEQVAVKVLSEESAQGYKEFRAEVDLL 621
Query: 142 TSVHHWNLVHLIGYCVE-GFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARG 200
VHH NL L+GYC E + L+YEYM N NL +L ++ R+KI+LD A+G
Sbjct: 622 MRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGKRSFILSWEERLKISLDAAQG 681
Query: 201 LEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYM 260
LEY+H+ P +HRD+K NILLNE K+ADFGL++ S + VAG+ GY+
Sbjct: 682 LEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEGSGQISTVVAGSIGYL 741
Query: 261 PPENAYGR-ISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSL 319
PE R ++ K DVY+ GVVL E+I+ + A+ KTE K + S D +S+
Sbjct: 742 DPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIAS-SKTE--------KVHIS-DHVRSI 791
Query: 320 VALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV 379
+A D+R +VD RL Y + S KM+++A AC QRPTM VV
Sbjct: 792 LA------------NGDIRGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQVV 839
Query: 380 VSLMEL 385
+ L ++
Sbjct: 840 MELKQI 845
>At1g51850 Putative protein kinase
Length = 875
Score = 200 bits (508), Expect = 1e-51
Identities = 125/340 (36%), Positives = 191/340 (55%), Gaps = 35/340 (10%)
Query: 66 KDTNDDNGSKYIWVDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRG-QKIAIK 124
+D S+ V K+ F+Y ++A T+NF + +G+GGFG VY+G + G +++A+K
Sbjct: 539 EDGRSPRSSEPAIVTKNRRFTYSQVAIMTNNFQ--RILGKGGFGMVYHGFVNGTEQVAVK 596
Query: 125 KMK---MQATREFLSELKVLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHNS 180
+ Q +EF +E+++L VHH NLV L+GYC EG + L+YEYM NG+L +H+ +
Sbjct: 597 ILSHSSSQGYKEFKAEVELLLRVHHKNLVGLVGYCDEGENMALIYEYMANGDLKEHMSGT 656
Query: 181 EKE-PMTLSTRMKIALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTK 239
+ TR+KI ++ A+GLEY+H+ P +HRD+K+ NILLNE+F K+ADFGL++
Sbjct: 657 RNRFTLNWGTRLKIVVESAQGLEYLHNGCKPPMVHRDVKTTNILLNEHFQAKLADFGLSR 716
Query: 240 LTDAANSADNTVHVAGTFGYMPPE-NAYGRISRKIDVYAFGVVLYELISAKAAVIKIDKT 298
+ VAGT GY+ PE ++ K DVY+FG+VL ELI+ + + K
Sbjct: 717 SFPIEGETHVSTVVAGTPGYLDPEYYKTNWLTEKSDVYSFGIVLLELITNRPVIDK---- 772
Query: 299 EFEFKSLEIKTNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMA 358
K +A + VM G D+ ++DP L +Y S+ K
Sbjct: 773 ---------------SREKPHIAEWVGVMLTKG----DINSIMDPNLNEDYDSGSVWKAV 813
Query: 359 KLAKACINRDPKQRPTMRDVVVSLMELNSSIDNKNSTGSA 398
+LA +C+N +RPTM VV+ ELN I ++NS G A
Sbjct: 814 ELAMSCLNPSSARRPTMSQVVI---ELNECIASENSRGGA 850
>At2g19210 putative receptor-like protein kinase
Length = 881
Score = 199 bits (507), Expect = 2e-51
Identities = 114/305 (37%), Positives = 174/305 (56%), Gaps = 27/305 (8%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIK---KMKMQATREFLSELKVL 141
+ Y E+ T+NF + +GQGGFG+VY+G L ++A+K + Q +EF +E+++L
Sbjct: 566 YKYSEVVKVTNNFE--RVLGQGGFGKVYHGVLNDDQVAVKILSESSAQGYKEFRAEVELL 623
Query: 142 TSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARG 200
VHH NL LIGYC EG + L+YE+M NG L +L + ++ R++I+LD A+G
Sbjct: 624 LRVHHKNLTALIGYCHEGKKMALIYEFMANGTLGDYLSGEKSYVLSWEERLQISLDAAQG 683
Query: 201 LEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYM 260
LEY+H+ P + RD+K NIL+NE K+ADFGL++ + +T VAGT GY+
Sbjct: 684 LEYLHNGCKPPIVQRDVKPANILINEKLQAKIADFGLSRSVALDGNNQDTTAVAGTIGYL 743
Query: 261 PPE-NAYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSL 319
PE + ++S K D+Y+FGVVL E++S + + + T + I + +D
Sbjct: 744 DPEYHLTQKLSEKSDIYSFGVVLLEVVSGQPVIARSRTT-----AENIHITDRVD----- 793
Query: 320 VALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV 379
+M TG D+R +VDP+LG + S K+ ++A AC + K RPTM VV
Sbjct: 794 ------LMLSTG----DIRGIVDPKLGERFDAGSAWKITEVAMACASSSSKNRPTMSHVV 843
Query: 380 VSLME 384
L E
Sbjct: 844 AELKE 848
>At4g34440 serine/threonine protein kinase - like
Length = 670
Score = 199 bits (506), Expect = 2e-51
Identities = 129/317 (40%), Positives = 187/317 (58%), Gaps = 34/317 (10%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGEL-RGQKIAIKKMKM---QATREFLSELKV 140
F+Y+EL+ AT+ F+ + +GQGGFG V+ G L G+++A+K +K+ Q REF +E+ +
Sbjct: 300 FTYDELSIATEGFAQSNLLGQGGFGYVHKGVLPSGKEVAVKSLKLGSGQGEREFQAEVDI 359
Query: 141 LTSVHHWNLVHLIGYCVEGFL-FLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVAR 199
++ VHH +LV L+GYC+ G LVYE++ N L HLH + + TR+KIAL AR
Sbjct: 360 ISRVHHRHLVSLVGYCISGGQRLLVYEFIPNNTLEFHLHGKGRPVLDWPTRVKIALGSAR 419
Query: 200 GLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVA----G 255
GL Y+H+ P IHRDIK+ NILL+ +F KVADFGL KL S DN HV+ G
Sbjct: 420 GLAYLHEDCHPRIIHRDIKAANILLDFSFETKVADFGLAKL-----SQDNYTHVSTRVMG 474
Query: 256 TFGYMPPENA-YGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESID 314
TFGY+ PE A G++S K DV++FGV+L ELI+ + + L + +S+
Sbjct: 475 TFGYLAPEYASSGKLSDKSDVFSFGVMLLELITGRPPL-----------DLTGEMEDSLV 523
Query: 315 EYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPT 374
++ + L Q G D +L DPRL NYS + +MA A A I ++RP
Sbjct: 524 DWARPLCL---KAAQDG----DYNQLADPRLELNYSHQEMVQMASCAAAAIRHSARRRPK 576
Query: 375 MRDVVVSLMELNSSIDN 391
M +V +L E + S+D+
Sbjct: 577 MSQIVRAL-EGDMSMDD 592
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.135 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,487,993
Number of Sequences: 26719
Number of extensions: 449575
Number of successful extensions: 5610
Number of sequences better than 10.0: 1013
Number of HSP's better than 10.0 without gapping: 851
Number of HSP's successfully gapped in prelim test: 162
Number of HSP's that attempted gapping in prelim test: 1413
Number of HSP's gapped (non-prelim): 1318
length of query: 398
length of database: 11,318,596
effective HSP length: 101
effective length of query: 297
effective length of database: 8,619,977
effective search space: 2560133169
effective search space used: 2560133169
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC123570.11


