

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122726.13 - phase: 0
(587 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
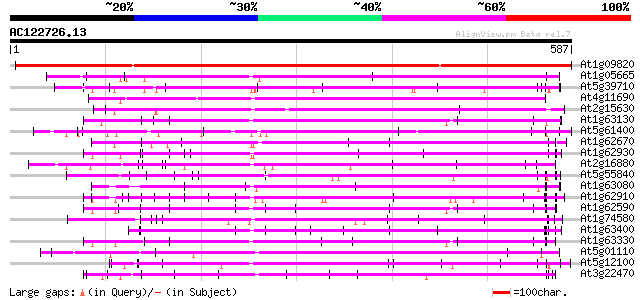
Score E
Sequences producing significant alignments: (bits) Value
At1g09820 hypothetical protein 638 0.0
At1g05665 putative protein 306 2e-83
At5g39710 putative protein 294 1e-79
At4g11690 putative protein 293 2e-79
At2g15630 putative salt-inducible protein 281 9e-76
At1g63130 unknown protein 279 4e-75
At5g61400 putative protein 276 2e-74
At1g62670 PPR-repeat protein 273 2e-73
At1g62930 unknown protein 273 3e-73
At2g16880 putative salt-inducible protein 272 4e-73
At5g55840 putative protein 271 1e-72
At1g63080 unknown protein 270 2e-72
At1g62910 unknown protein 266 2e-71
At1g62590 putative membrane-associated salt-inducible protein (A... 266 2e-71
At1g74580 hypothetical protein 266 3e-71
At1g63400 PPR-repeat protein 265 4e-71
At1g63330 unknown protein 265 4e-71
At5g01110 unknown protein (At5g01110) 264 9e-71
At5g12100 unknown protein (At5g12100) 261 8e-70
At3g22470 hypothetical protein 260 2e-69
>At1g09820 hypothetical protein
Length = 606
Score = 638 bits (1646), Expect = 0.0
Identities = 317/584 (54%), Positives = 420/584 (71%), Gaps = 5/584 (0%)
Query: 7 LITSEFLFTG-PILQSLSIPTISELLSKQHWSELKPHLRVTKPATFLDQLLNAGVDSELV 65
L +S TG P + I++L+ KQHWS+L H+ P QL+++ +D +L
Sbjct: 25 LCSSSSTITGSPCPPRYDVAVIADLIEKQHWSKLGVHVTDINPNELFRQLISSELDPDLC 84
Query: 66 LRFFKWSQKEYRLSYGLEPTSKVLHFLANSKRYSKVRSFLDSFVKN-EKHTVSSVFHSLL 124
LR++ W K +S LE T K+LH LAN+KRYSK+RSFLD FV+N H V S+FH++
Sbjct: 85 LRYYSWLVKNSDISVSLELTFKLLHSLANAKRYSKIRSFLDGFVRNGSDHQVHSIFHAIS 144
Query: 125 LDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENK 184
+ ++I DMLVLAY N +EAF R+ YG+KLS SC PL+ AL+KEN+
Sbjct: 145 MCDN-VCVNSIIADMLVLAYANNSRFELGFEAFKRSGYYGYKLSALSCKPLMIALLKENR 203
Query: 185 IGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNT 244
DVEYVYKEMI+R+I N+ TFN+ IN LC+ GK+NKA D +EDMK +G SPNVV+YNT
Sbjct: 204 SADVEYVYKEMIRRKIQPNVFTFNVVINALCKTGKMNKARDVMEDMKVYGCSPNVVSYNT 263
Query: 245 LVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQ 304
L+DGYCK G GKMYKA+A +KEM+ N + PN TFN LIDGF KD+N+ + K F+EM
Sbjct: 264 LIDGYCKLGGNGKMYKADAVLKEMVENDVSPNLTTFNILIDGFWKDDNLPGSMKVFKEML 323
Query: 305 KQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMK 364
Q +KPN+++YNSLINGLCN GK+ EAI + DKMV G++PN++TYNALINGFCK M+K
Sbjct: 324 DQDVKPNVISYNSLINGLCNGGKISEAISMRDKMVSAGVQPNLITYNALINGFCKNDMLK 383
Query: 365 EATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLI 424
EA +F V Q VP +N +IDAYCK G +++GF+L M EGI+P+V TYNCLI
Sbjct: 384 EALDMFGSVKGQGAVPTTRMYNMLIDAYCKLGKIDDGFALKEEMEREGIVPDVGTYNCLI 443
Query: 425 AGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLK 484
AGLCR +++AAK+L +++ +KGL D+VT++IL++G C+ +SR A LL EM +GLK
Sbjct: 444 AGLCRNGNIEAAKKLFDQLTSKGLP-DLVTFHILMEGYCRKGESRKAAMLLKEMSKMGLK 502
Query: 485 PNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERK-QPNVVTYNVLIKGYCKINKLEAANG 543
P H+TYN +M GYC EG LKAA N+RT+MEKER+ + NV +YNVL++GY + KLE AN
Sbjct: 503 PRHLTYNIVMKGYCKEGNLKAATNMRTQMEKERRLRMNVASYNVLLQGYSQKGKLEDANM 562
Query: 544 LLNEMLEKGLNPNRTTYDIVRLEMLEKGFSPDIEGHLYNISSMS 587
LLNEMLEKGL PNR TY+IV+ EM+++GF PDIEGHL+N+S+ S
Sbjct: 563 LLNEMLEKGLVPNRITYEIVKEEMVDQGFVPDIEGHLFNVSTKS 606
>At1g05665 putative protein
Length = 732
Score = 306 bits (784), Expect = 2e-83
Identities = 183/560 (32%), Positives = 291/560 (51%), Gaps = 39/560 (6%)
Query: 39 LKPHLRVTKPATFLDQLLNAGVDSELVLRFFKWSQKEYRLSYGLEPTSKVLHFLANSKRY 98
LKP+ K + L+ D LVL FF W++ R LE V+H SK
Sbjct: 69 LKPYECKFKTDHLIWVLMKIKCDYRLVLDFFDWARS--RRDSNLESLCIVIHLAVASKDL 126
Query: 99 SKVRSFLDSFVKNEKHTVSSVF---HSLLLDGGRP-GATALIIDMLVLAYVKNLELHCAY 154
+S + SF + K V+ F LL+ + G+ + D+ V L A
Sbjct: 127 KVAQSLISSFWERPKLNVTDSFVQFFDLLVYTYKDWGSDPRVFDVFFQVLVDFGLLREAR 186
Query: 155 EAFTRAKDYGFKLSLTSCNPLLSALVKE-NKIGDVEYVYKEMIKRRIHTNLNTFNIFING 213
F + +YG LS+ SCN L+ L K+ K V++E + + N+ ++NI I+
Sbjct: 187 RVFEKMLNYGLVLSVDSCNVYLTRLSKDCYKTATAIIVFREFPEVGVCWNVASYNIVIHF 246
Query: 214 LCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYK------------- 260
+C+ G++ +A + M+ G +P+V++Y+T+V+GYC+ G K++K
Sbjct: 247 VCQLGRIKEAHHLLLLMELKGYTPDVISYSTVVNGYCRFGELDKVWKLIEVMKRKGLKPN 306
Query: 261 -------------------AEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFE 301
AE EM+ I P+ V + TLIDGFCK ++ AA K F
Sbjct: 307 SYIYGSIIGLLCRICKLAEAEEAFSEMIRQGILPDTVVYTTLIDGFCKRGDIRAASKFFY 366
Query: 302 EMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKK 361
EM + + P+++TY ++I+G C G + EA L+ +M GL+P+ VT+ LING+CK
Sbjct: 367 EMHSRDITPDVLTYTAIISGFCQIGDMVEAGKLFHEMFCKGLEPDSVTFTELINGYCKAG 426
Query: 362 MMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYN 421
MK+A +V + + + PNV+T+ T+ID CKEG ++ L M G+ PN+ TYN
Sbjct: 427 HMKDAFRVHNHMIQAGCSPNVVTYTTLIDGLCKEGDLDSANELLHEMWKIGLQPNIFTYN 486
Query: 422 CLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNL 481
++ GLC+ +++ A +L+ E E GL D VTY L+D CK+ + A+++L EM
Sbjct: 487 SIVNGLCKSGNIEEAVKLVGEFEAAGLNADTVTYTTLMDAYCKSGEMDKAQEILKEMLGK 546
Query: 482 GLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAA 541
GL+P VT+N LM+G+C+ G L+ + M + PN T+N L+K YC N L+AA
Sbjct: 547 GLQPTIVTFNVLMNGFCLHGMLEDGEKLLNWMLAKGIAPNATTFNSLVKQYCIRNNLKAA 606
Query: 542 NGLLNEMLEKGLNPNRTTYD 561
+ +M +G+ P+ TY+
Sbjct: 607 TAIYKDMCSRGVGPDGKTYE 626
Score = 243 bits (621), Expect = 2e-64
Identities = 142/457 (31%), Positives = 238/457 (52%), Gaps = 6/457 (1%)
Query: 121 HSLLLDGGRPGATALIIDM--LVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSA 178
H LLL G T +I +V Y + EL ++ K G K + ++
Sbjct: 257 HHLLLLMELKGYTPDVISYSTVVNGYCRFGELDKVWKLIEVMKRKGLKPNSYIYGSIIGL 316
Query: 179 LVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPN 238
L + K+ + E + EMI++ I + + I+G C+ G + A +M + I+P+
Sbjct: 317 LCRICKLAEAEEAFSEMIRQGILPDTVVYTTLIDGFCKRGDIRAASKFFYEMHSRDITPD 376
Query: 239 VVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKK 298
V+TY ++ G+C+ G M +A EM + P+ VTF LI+G+CK ++ A +
Sbjct: 377 VLTYTAIISGFCQ---IGDMVEAGKLFHEMFCKGLEPDSVTFTELINGYCKAGHMKDAFR 433
Query: 299 AFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFC 358
M + G PN+VTY +LI+GLC G L+ A +L +M +GL+PNI TYN+++NG C
Sbjct: 434 VHNHMIQAGCSPNVVTYTTLIDGLCKEGDLDSANELLHEMWKIGLQPNIFTYNSIVNGLC 493
Query: 359 KKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVS 418
K ++EA K+ + L + +T+ T++DAYCK G M++ + ML +G+ P +
Sbjct: 494 KSGNIEEAVKLVGEFEAAGLNADTVTYTTLMDAYCKSGEMDKAQEILKEMLGKGLQPTIV 553
Query: 419 TYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEM 478
T+N L+ G C L+ ++LLN M KG+ + T+N L+ C + + A + +M
Sbjct: 554 TFNVLMNGFCLHGMLEDGEKLLNWMLAKGIAPNATTFNSLVKQYCIRNNLKAATAIYKDM 613
Query: 479 FNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKL 538
+ G+ P+ TY L+ G+C +K A + M+ + +V TY+VLIKG+ K K
Sbjct: 614 CSRGVGPDGKTYENLVKGHCKARNMKEAWFLFQEMKGKGFSVSVSTYSVLIKGFLKRKKF 673
Query: 539 EAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKGFSPD 575
A + ++M +GL ++ +D + KG PD
Sbjct: 674 LEAREVFDQMRREGLAADKEIFDFFS-DTKYKGKRPD 709
Score = 104 bits (259), Expect = 2e-22
Identities = 83/324 (25%), Positives = 138/324 (41%), Gaps = 31/324 (9%)
Query: 81 GLEPTSKVLHFLANSKRYSKVRSFLDSFVKNEKH-----------TVSSVFHSLLLDGGR 129
GLEP S L N Y K D+F + H T +++ L +G
Sbjct: 407 GLEPDSVTFTELING--YCKAGHMKDAF-RVHNHMIQAGCSPNVVTYTTLIDGLCKEGDL 463
Query: 130 PGATALIIDM--------------LVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPL 175
A L+ +M +V K+ + A + + G + L
Sbjct: 464 DSANELLHEMWKIGLQPNIFTYNSIVNGLCKSGNIEEAVKLVGEFEAAGLNADTVTYTTL 523
Query: 176 LSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGI 235
+ A K ++ + + KEM+ + + + TFN+ +NG C G L E + M A GI
Sbjct: 524 MDAYCKSGEMDKAQEILKEMLGKGLQPTIVTFNVLMNGFCLHGMLEDGEKLLNWMLAKGI 583
Query: 236 SPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAA 295
+PN T+N+LV YC R + + A A K+M + + P+ T+ L+ G CK N+
Sbjct: 584 APNATTFNSLVKQYCIRNN---LKAATAIYKDMCSRGVGPDGKTYENLVKGHCKARNMKE 640
Query: 296 AKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALIN 355
A F+EM+ +G ++ TY+ LI G K EA +++D+M GL + ++ +
Sbjct: 641 AWFLFQEMKGKGFSVSVSTYSVLIKGFLKRKKFLEAREVFDQMRREGLAADKEIFDFFSD 700
Query: 356 GFCKKKMMKEATKVFDDVSKQELV 379
K K D++ + LV
Sbjct: 701 TKYKGKRPDTIVDPIDEIIENYLV 724
>At5g39710 putative protein
Length = 747
Score = 294 bits (752), Expect = 1e-79
Identities = 176/548 (32%), Positives = 279/548 (50%), Gaps = 37/548 (6%)
Query: 48 PATFLDQLLNAGVDSELVLRFFKWSQKEYRLSYGLEPTSKVLHFLANSKRYSKVRSFLDS 107
P + LL + D L+L+F W+ + L LH L K Y + +
Sbjct: 48 PEAASNLLLKSQNDQALILKFLNWANPHQ--FFTLRCKCITLHILTKFKLYKTAQILAED 105
Query: 108 FVKN--EKHTVSSVFHSLLLDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGF 165
+ S VF SL +T+ + D++V +Y + + A A+ +GF
Sbjct: 106 VAAKTLDDEYASLVFKSLQETYDLCYSTSSVFDLVVKSYSRLSLIDKALSIVHLAQAHGF 165
Query: 166 KLSLTSCNPLLSALVKENK-IGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAE 224
+ S N +L A ++ + I E V+KEM++ ++ N+ T+NI I G C AG ++ A
Sbjct: 166 MPGVLSYNAVLDATIRSKRNISFAENVFKEMLESQVSPNVFTYNILIRGFCFAGNIDVAL 225
Query: 225 DAIEDMKAWGISPNVVTYNTLVDGYCK-----------RGSA------------------ 255
+ M+ G PNVVTYNTL+DGYCK R A
Sbjct: 226 TLFDKMETKGCLPNVVTYNTLIDGYCKLRKIDDGFKLLRSMALKGLEPNLISYNVVINGL 285
Query: 256 ---GKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNI 312
G+M + + EM +EVT+NTLI G+CK+ N A EM + GL P++
Sbjct: 286 CREGRMKEVSFVLTEMNRRGYSLDEVTYNTLIKGYCKEGNFHQALVMHAEMLRHGLTPSV 345
Query: 313 VTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDD 372
+TY SLI+ +C G + A++ D+M GL PN TY L++GF +K M EA +V +
Sbjct: 346 ITYTSLIHSMCKAGNMNRAMEFLDQMRVRGLCPNERTYTTLVDGFSQKGYMNEAYRVLRE 405
Query: 373 VSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQD 432
++ P+V+T+N +I+ +C G ME+ ++ M ++G+ P+V +Y+ +++G CR D
Sbjct: 406 MNDNGFSPSVVTYNALINGHCVTGKMEDAIAVLEDMKEKGLSPDVVSYSTVLSGFCRSYD 465
Query: 433 LQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNT 492
+ A + EM KG+K D +TY+ LI G C+ +++ A L EM +GL P+ TY
Sbjct: 466 VDEALRVKREMVEKGIKPDTITYSSLIQGFCEQRRTKEACDLYEEMLRVGLPPDEFTYTA 525
Query: 493 LMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKG 552
L++ YCMEG L+ AL + M ++ P+VVTY+VLI G K ++ A LL ++ +
Sbjct: 526 LINAYCMEGDLEKALQLHNEMVEKGVLPDVVTYSVLINGLNKQSRTREAKRLLLKLFYEE 585
Query: 553 LNPNRTTY 560
P+ TY
Sbjct: 586 SVPSDVTY 593
Score = 263 bits (672), Expect = 2e-70
Identities = 153/497 (30%), Positives = 260/497 (51%), Gaps = 51/497 (10%)
Query: 105 LDSFVKNEKHT--VSSVFHSLLLDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKD 162
LD+ ++++++ +VF +L P I + + N+++ A F + +
Sbjct: 176 LDATIRSKRNISFAENVFKEMLESQVSPNVFTYNILIRGFCFAGNIDV--ALTLFDKMET 233
Query: 163 YGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNK 222
G ++ + N L+ K KI D + + M + + NL ++N+ INGLCR G++ +
Sbjct: 234 KGCLPNVVTYNTLIDGYCKLRKIDDGFKLLRSMALKGLEPNLISYNVVINGLCREGRMKE 293
Query: 223 AEDAIEDMKAWGISPNVVTYNTLVDGYCKRGS---------------------------- 254
+ +M G S + VTYNTL+ GYCK G+
Sbjct: 294 VSFVLTEMNRRGYSLDEVTYNTLIKGYCKEGNFHQALVMHAEMLRHGLTPSVITYTSLIH 353
Query: 255 ----AGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKP 310
AG M +A F+ +M +CPNE T+ TL+DGF + + A + EM G P
Sbjct: 354 SMCKAGNMNRAMEFLDQMRVRGLCPNERTYTTLVDGFSQKGYMNEAYRVLREMNDNGFSP 413
Query: 311 NIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVF 370
++VTYN+LING C GK+E+AI + + M GL P++V+Y+ +++GFC+ + EA +V
Sbjct: 414 SVVTYNALINGHCVTGKMEDAIAVLEDMKEKGLSPDVVSYSTVLSGFCRSYDVDEALRVK 473
Query: 371 DDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRK 430
++ ++ + P+ IT++++I +C++ +E L ML G+ P+ TY LI C +
Sbjct: 474 REMVEKGIKPDTITYSSLIQGFCEQRRTKEACDLYEEMLRVGLPPDEFTYTALINAYCME 533
Query: 431 QDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTY 490
DL+ A +L NEM KG+ DVVTY++LI+GL K ++R A++LL ++F P+ VTY
Sbjct: 534 GDLEKALQLHNEMVEKGVLPDVVTYSVLINGLNKQSRTREAKRLLLKLFYEESVPSDVTY 593
Query: 491 NTLMD---------------GYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKI 535
+TL++ G+CM+G + A V M + +P+ YN++I G+C+
Sbjct: 594 HTLIENCSNIEFKSVVSLIKGFCMKGMMTEADQVFESMLGKNHKPDGTAYNIMIHGHCRA 653
Query: 536 NKLEAANGLLNEMLEKG 552
+ A L EM++ G
Sbjct: 654 GDIRKAYTLYKEMVKSG 670
Score = 210 bits (535), Expect = 2e-54
Identities = 117/333 (35%), Positives = 181/333 (54%), Gaps = 17/333 (5%)
Query: 260 KAEAFMKEMLANKICPNEVTFNTLIDGFCKDE-NVAAAKKAFEEMQKQGLKPNIVTYNSL 318
KA + + A+ P +++N ++D + + N++ A+ F+EM + + PN+ TYN L
Sbjct: 152 KALSIVHLAQAHGFMPGVLSYNAVLDATIRSKRNISFAENVFKEMLESQVSPNVFTYNIL 211
Query: 319 INGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQEL 378
I G C G ++ A+ L+DKM G PN+VTYN LI+G+CK + + + K+ ++ + L
Sbjct: 212 IRGFCFAGNIDVALTLFDKMETKGCLPNVVTYNTLIDGYCKLRKIDDGFKLLRSMALKGL 271
Query: 379 VPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKE 438
PN+I++N +I+ C+EG M+E + + M G + TYN LI G C++ + A
Sbjct: 272 EPNLISYNVVINGLCREGRMKEVSFVLTEMNRRGYSLDEVTYNTLIKGYCKEGNFHQALV 331
Query: 439 LLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYC 498
+ EM GL V+TY LI +CK A + L++M GL PN TY TL+DG+
Sbjct: 332 MHAEMLRHGLTPSVITYTSLIHSMCKAGNMNRAMEFLDQMRVRGLCPNERTYTTLVDGFS 391
Query: 499 MEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRT 558
+G + A V M P+VVTYN LI G+C K+E A +L +M EKGL+P+
Sbjct: 392 QKGYMNEAYRVLREMNDNGFSPSVVTYNALINGHCVTGKMEDAIAVLEDMKEKGLSPDVV 451
Query: 559 TYDIV----------------RLEMLEKGFSPD 575
+Y V + EM+EKG PD
Sbjct: 452 SYSTVLSGFCRSYDVDEALRVKREMVEKGIKPD 484
Score = 207 bits (528), Expect = 1e-53
Identities = 123/443 (27%), Positives = 218/443 (48%), Gaps = 53/443 (11%)
Query: 164 GFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKA 223
G+ L + N L+ KE ++ EM++ + ++ T+ I+ +C+AG +N+A
Sbjct: 305 GYSLDEVTYNTLIKGYCKEGNFHQALVMHAEMLRHGLTPSVITYTSLIHSMCKAGNMNRA 364
Query: 224 EDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTL 283
+ ++ M+ G+ PN TY TLVDG+ ++G + Y+ ++EM N P+ VT+N L
Sbjct: 365 MEFLDQMRVRGLCPNERTYTTLVDGFSQKGYMNEAYRV---LREMNDNGFSPSVVTYNAL 421
Query: 284 IDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGL 343
I+G C + A E+M+++GL P++V+Y+++++G C + ++EA+ + +MV G+
Sbjct: 422 INGHCVTGKMEDAIAVLEDMKEKGLSPDVVSYSTVLSGFCRSYDVDEALRVKREMVEKGI 481
Query: 344 KPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFS 403
KP+ +TY++LI GFC+++ KEA +++++ + L P+ T+ +I+AYC EG +E+
Sbjct: 482 KPDTITYSSLIQGFCEQRRTKEACDLYEEMLRVGLPPDEFTYTALINAYCMEGDLEKALQ 541
Query: 404 LCSSMLDEGILPNVSTY----------------------------------------NC- 422
L + M+++G+LP+V TY NC
Sbjct: 542 LHNEMVEKGVLPDVVTYSVLINGLNKQSRTREAKRLLLKLFYEESVPSDVTYHTLIENCS 601
Query: 423 ---------LIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEK 473
LI G C K + A ++ M K K D YNI+I G C+ R A
Sbjct: 602 NIEFKSVVSLIKGFCMKGMMTEADQVFESMLGKNHKPDGTAYNIMIHGHCRAGDIRKAYT 661
Query: 474 LLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYC 533
L EM G + VT L+ EGK+ +V + + + VL++
Sbjct: 662 LYKEMVKSGFLLHTVTVIALVKALHKEGKVNELNSVIVHVLRSCELSEAEQAKVLVEINH 721
Query: 534 KINKLEAANGLLNEMLEKGLNPN 556
+ ++ +L EM + G PN
Sbjct: 722 REGNMDVVLDVLAEMAKDGFLPN 744
Score = 157 bits (397), Expect = 2e-38
Identities = 86/266 (32%), Positives = 140/266 (52%), Gaps = 17/266 (6%)
Query: 328 LEEAIDLWDKMVGLGLKPNIVTYNALINGFCK-KKMMKEATKVFDDVSKQELVPNVITFN 386
+++A+ + G P +++YNA+++ + K+ + A VF ++ + ++ PNV T+N
Sbjct: 150 IDKALSIVHLAQAHGFMPGVLSYNAVLDATIRSKRNISFAENVFKEMLESQVSPNVFTYN 209
Query: 387 TMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENK 446
+I +C G ++ +L M +G LPNV TYN LI G C+ + + +LL M K
Sbjct: 210 ILIRGFCFAGNIDVALTLFDKMETKGCLPNVVTYNTLIDGYCKLRKIDDGFKLLRSMALK 269
Query: 447 GLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAA 506
GL+ ++++YN++I+GLC+ + + +L EM G + VTYNTL+ GYC EG A
Sbjct: 270 GLEPNLISYNVVINGLCREGRMKEVSFVLTEMNRRGYSLDEVTYNTLIKGYCKEGNFHQA 329
Query: 507 LNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDI---- 562
L + M + P+V+TY LI CK + A L++M +GL PN TY
Sbjct: 330 LVMHAEMLRHGLTPSVITYTSLIHSMCKAGNMNRAMEFLDQMRVRGLCPNERTYTTLVDG 389
Query: 563 ------------VRLEMLEKGFSPDI 576
V EM + GFSP +
Sbjct: 390 FSQKGYMNEAYRVLREMNDNGFSPSV 415
Score = 117 bits (292), Expect = 2e-26
Identities = 95/408 (23%), Positives = 177/408 (43%), Gaps = 42/408 (10%)
Query: 78 LSYGLEPT----SKVLHFLANSKRYSKVRSFLDSF-----VKNEKHTVSSVFHSLLLDGG 128
L +GL P+ + ++H + + ++ FLD NE+ T +++ G
Sbjct: 337 LRHGLTPSVITYTSLIHSMCKAGNMNRAMEFLDQMRVRGLCPNER-TYTTLVDGFSQKGY 395
Query: 129 RPGATALIIDM-------LVLAYVKNLELHC-------AYEAFTRAKDYGFKLSLTSCNP 174
A ++ +M V+ Y + HC A K+ G + S +
Sbjct: 396 MNEAYRVLREMNDNGFSPSVVTYNALINGHCVTGKMEDAIAVLEDMKEKGLSPDVVSYST 455
Query: 175 LLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWG 234
+LS + + + V +EM+++ I + T++ I G C + +A D E+M G
Sbjct: 456 VLSGFCRSYDVDEALRVKREMVEKGIKPDTITYSSLIQGFCEQRRTKEACDLYEEMLRVG 515
Query: 235 ISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVA 294
+ P+ TY L++ YC G + KA EM+ + P+ VT++ LI+G K
Sbjct: 516 LPPDEFTYTALINAYCMEGD---LEKALQLHNEMVEKGVLPDVVTYSVLINGLNKQSRTR 572
Query: 295 AAKKAFEEMQKQGLKPNIVTYN---------------SLINGLCNNGKLEEAIDLWDKMV 339
AK+ ++ + P+ VTY+ SLI G C G + EA +++ M+
Sbjct: 573 EAKRLLLKLFYEESVPSDVTYHTLIENCSNIEFKSVVSLIKGFCMKGMMTEADQVFESML 632
Query: 340 GLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMME 399
G KP+ YN +I+G C+ +++A ++ ++ K + + +T ++ A KEG +
Sbjct: 633 GKNHKPDGTAYNIMIHGHCRAGDIRKAYTLYKEMVKSGFLLHTVTVIALVKALHKEGKVN 692
Query: 400 EGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKG 447
E S+ +L L L+ R+ ++ ++L EM G
Sbjct: 693 ELNSVIVHVLRSCELSEAEQAKVLVEINHREGNMDVVLDVLAEMAKDG 740
>At4g11690 putative protein
Length = 566
Score = 293 bits (749), Expect = 2e-79
Identities = 166/482 (34%), Positives = 264/482 (54%), Gaps = 9/482 (1%)
Query: 83 EPTSKVLHFLANSKRYSKVRSFLDSFVKNEKH----TVSSVFHSLLLDGGRPGATALIID 138
E S +L L + +S +S L + + H T SS+ H L + + +
Sbjct: 40 ESISILLRLLLSGNLFSHAQSLLLQVISGKIHSQFFTSSSLLH-YLTESETSKTKFRLYE 98
Query: 139 MLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKR 198
+++ +YV++ L+ + F D GF N LL+ +V + + E K
Sbjct: 99 VIINSYVQSQSLNLSISYFNEMVDNGFVPGSNCFNYLLTFVVGSSSFNQWWSFFNEN-KS 157
Query: 199 RIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKM 258
++ ++ +F I I G C AG++ K+ D + ++ +G SPNVV Y TL+DG CK+G ++
Sbjct: 158 KVVLDVYSFGILIKGCCEAGEIEKSFDLLIELTEFGFSPNVVIYTTLIDGCCKKG---EI 214
Query: 259 YKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSL 318
KA+ EM + NE T+ LI+G K+ + +E+MQ+ G+ PN+ TYN +
Sbjct: 215 EKAKDLFFEMGKLGLVANERTYTVLINGLFKNGVKKQGFEMYEKMQEDGVFPNLYTYNCV 274
Query: 319 INGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQEL 378
+N LC +G+ ++A ++D+M G+ NIVTYN LI G C++ + EA KV D + +
Sbjct: 275 MNQLCKDGRTKDAFQVFDEMRERGVSCNIVTYNTLIGGLCREMKLNEANKVVDQMKSDGI 334
Query: 379 VPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKE 438
PN+IT+NT+ID +C G + + SLC + G+ P++ TYN L++G CRK D A +
Sbjct: 335 NPNLITYNTLIDGFCGVGKLGKALSLCRDLKSRGLSPSLVTYNILVSGFCRKGDTSGAAK 394
Query: 439 LLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYC 498
++ EME +G+K VTY ILID ++D A +L M LGL P+ TY+ L+ G+C
Sbjct: 395 MVKEMEERGIKPSKVTYTILIDTFARSDNMEKAIQLRLSMEELGLVPDVHTYSVLIHGFC 454
Query: 499 MEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRT 558
++G++ A + M ++ +PN V YN +I GYCK A LL EM EK L PN
Sbjct: 455 IKGQMNEASRLFKSMVEKNCEPNEVIYNTMILGYCKEGSSYRALKLLKEMEEKELAPNVA 514
Query: 559 TY 560
+Y
Sbjct: 515 SY 516
>At2g15630 putative salt-inducible protein
Length = 627
Score = 281 bits (718), Expect = 9e-76
Identities = 169/481 (35%), Positives = 258/481 (53%), Gaps = 12/481 (2%)
Query: 101 VRSFLDSFVKNEKHTVSSVFHSLLLDGGR-PGATALIIDMLVLAYVKNLELHCAYEAFTR 159
V L V + K+++ ++F L+L R + ++ D+LV + + A E F
Sbjct: 121 VTQLLKEVVTSRKNSIRNLFDELVLAHDRLETKSTILFDLLVRCCCQLRMVDEAIECFYL 180
Query: 160 AKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGK 219
K+ GF +CN +L+ L + N+I + Y +M + I +N+ TFNI IN LC+ GK
Sbjct: 181 MKEKGFYPKTETCNHILTLLSRLNRIENAWVFYADMYRMEIKSNVYTFNIMINVLCKEGK 240
Query: 220 LNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVT 279
L KA+ + M+ +GI P +VTYNTLV G+ RG ++ A + EM + P+ T
Sbjct: 241 LKKAKGFLGIMEVFGIKPTIVTYNTLVQGFSLRG---RIEGARLIISEMKSKGFQPDMQT 297
Query: 280 FNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMV 339
+N ++ C N A + EM++ GL P+ V+YN LI G NNG LE A D+MV
Sbjct: 298 YNPILSWMC---NEGRASEVLREMKEIGLVPDSVSYNILIRGCSNNGDLEMAFAYRDEMV 354
Query: 340 GLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMME 399
G+ P TYN LI+G + ++ A + ++ ++ +V + +T+N +I+ YC+ G +
Sbjct: 355 KQGMVPTFYTYNTLIHGLFMENKIEAAEILIREIREKGIVLDSVTYNILINGYCQHGDAK 414
Query: 400 EGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILI 459
+ F+L M+ +GI P TY LI LCRK + A EL ++ KG+K D+V N L+
Sbjct: 415 KAFALHDEMMTDGIQPTQFTYTSLIYVLCRKNKTREADELFEKVVGKGMKPDLVMMNTLM 474
Query: 460 DGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQ 519
DG C A LL EM + + P+ VTYN LM G C EGK + A + M++ +
Sbjct: 475 DGHCAIGNMDRAFSLLKEMDMMSINPDDVTYNCLMRGLCGEGKFEEARELMGEMKRRGIK 534
Query: 520 PNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKGFSPDIEGH 579
P+ ++YN LI GY K + A + +EML G NP TY+ L KG S + EG
Sbjct: 535 PDHISYNTLISGYSKKGDTKHAFMVRDEMLSLGFNPTLLTYN-----ALLKGLSKNQEGE 589
Query: 580 L 580
L
Sbjct: 590 L 590
Score = 174 bits (440), Expect = 2e-43
Identities = 109/399 (27%), Positives = 203/399 (50%), Gaps = 20/399 (5%)
Query: 88 VLHFLANSKRYSKVRSFLDSF----VKNEKHTVSSVFHSLLLDGGRPGATALIIDMLVLA 143
+++ L + K + FL +K T +++ L G GA +I +M
Sbjct: 231 MINVLCKEGKLKKAKGFLGIMEVFGIKPTIVTYNTLVQGFSLRGRIEGARLIISEMKSKG 290
Query: 144 YVKNLELH-------C----AYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVY 192
+ +++ + C A E K+ G S N L+ + ++ + Y
Sbjct: 291 FQPDMQTYNPILSWMCNEGRASEVLREMKEIGLVPDSVSYNILIRGCSNNGDL-EMAFAY 349
Query: 193 K-EMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCK 251
+ EM+K+ + T+N I+GL K+ AE I +++ GI + VTYN L++GYC+
Sbjct: 350 RDEMVKQGMVPTFYTYNTLIHGLFMENKIEAAEILIREIREKGIVLDSVTYNILINGYCQ 409
Query: 252 RGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPN 311
G A K + A EM+ + I P + T+ +LI C+ A + FE++ +G+KP+
Sbjct: 410 HGDAKKAF---ALHDEMMTDGIQPTQFTYTSLIYVLCRKNKTREADELFEKVVGKGMKPD 466
Query: 312 IVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFD 371
+V N+L++G C G ++ A L +M + + P+ VTYN L+ G C + +EA ++
Sbjct: 467 LVMMNTLMDGHCAIGNMDRAFSLLKEMDMMSINPDDVTYNCLMRGLCGEGKFEEARELMG 526
Query: 372 DVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQ 431
++ ++ + P+ I++NT+I Y K+G + F + ML G P + TYN L+ GL + Q
Sbjct: 527 EMKRRGIKPDHISYNTLISGYSKKGDTKHAFMVRDEMLSLGFNPTLLTYNALLKGLSKNQ 586
Query: 432 DLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRN 470
+ + A+ELL EM+++G+ + ++ +I+ + D ++
Sbjct: 587 EGELAEELLREMKSEGIVPNDSSFCSVIEAMSNLDAKKS 625
>At1g63130 unknown protein
Length = 630
Score = 279 bits (713), Expect = 4e-75
Identities = 144/439 (32%), Positives = 244/439 (54%), Gaps = 3/439 (0%)
Query: 139 MLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKR 198
+L+ + + +L A + G++ + + N LL+ N+I D + +M++
Sbjct: 121 ILINCFCRRSQLSLALAVLAKMMKLGYEPDIVTLNSLLNGFCHGNRISDAVSLVGQMVEM 180
Query: 199 RIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKM 258
+ TFN I+GL R + ++A ++ M G P++VTY +V+G CKRG
Sbjct: 181 GYQPDSFTFNTLIHGLFRHNRASEAVALVDRMVVKGCQPDLVTYGIVVNGLCKRGDIDL- 239
Query: 259 YKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSL 318
A + +K+M KI P V +NT+ID C +NV A F EM +G++PN+VTYNSL
Sbjct: 240 --ALSLLKKMEQGKIEPGVVIYNTIIDALCNYKNVNDALNLFTEMDNKGIRPNVVTYNSL 297
Query: 319 INGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQEL 378
I LCN G+ +A L M+ + PN+VT++ALI+ F K+ + EA K++D++ K+ +
Sbjct: 298 IRCLCNYGRWSDASRLLSDMIERKINPNVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSI 357
Query: 379 VPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKE 438
P++ T++++I+ +C ++E + M+ + PNV TYN LI G C+ + + E
Sbjct: 358 DPDIFTYSSLINGFCMHDRLDEAKHMFELMISKDCFPNVVTYNTLIKGFCKAKRVDEGME 417
Query: 439 LLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYC 498
L EM +GL G+ VTY LI G + + NA+ + +M + G+ P+ +TY+ L+DG C
Sbjct: 418 LFREMSQRGLVGNTVTYTTLIHGFFQARECDNAQIVFKQMVSDGVLPDIMTYSILLDGLC 477
Query: 499 MEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRT 558
GK++ AL V +++ + +P++ TYN++I+G CK K+E L + KG+ PN
Sbjct: 478 NNGKVETALVVFEYLQRSKMEPDIYTYNIMIEGMCKAGKVEDGWDLFCSLSLKGVKPNVV 537
Query: 559 TYDIVRLEMLEKGFSPDIE 577
TY + KG + +
Sbjct: 538 TYTTMMSGFCRKGLKEEAD 556
Score = 243 bits (621), Expect = 2e-64
Identities = 146/479 (30%), Positives = 245/479 (50%), Gaps = 32/479 (6%)
Query: 78 LSYGLEPTSKVLHFLAN--------SKRYSKVRSFLDSFVKNEKHTVSSVFHSLLLDGGR 129
+ G EP L+ L N S S V ++ + + T +++ H L
Sbjct: 143 MKLGYEPDIVTLNSLLNGFCHGNRISDAVSLVGQMVEMGYQPDSFTFNTLIHGLFRHNRA 202
Query: 130 PGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVE 189
A AL+ M+V G + L + +++ L K I
Sbjct: 203 SEAVALVDRMVVK---------------------GCQPDLVTYGIVVNGLCKRGDIDLAL 241
Query: 190 YVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGY 249
+ K+M + +I + +N I+ LC +N A + +M GI PNVVTYN+L+
Sbjct: 242 SLLKKMEQGKIEPGVVIYNTIIDALCNYKNVNDALNLFTEMDNKGIRPNVVTYNSLIRCL 301
Query: 250 CKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLK 309
C + G+ A + +M+ KI PN VTF+ LID F K+ + A+K ++EM K+ +
Sbjct: 302 C---NYGRWSDASRLLSDMIERKINPNVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSID 358
Query: 310 PNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKV 369
P+I TY+SLING C + +L+EA +++ M+ PN+VTYN LI GFCK K + E ++
Sbjct: 359 PDIFTYSSLINGFCMHDRLDEAKHMFELMISKDCFPNVVTYNTLIKGFCKAKRVDEGMEL 418
Query: 370 FDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCR 429
F ++S++ LV N +T+ T+I + + + + M+ +G+LP++ TY+ L+ GLC
Sbjct: 419 FREMSQRGLVGNTVTYTTLIHGFFQARECDNAQIVFKQMVSDGVLPDIMTYSILLDGLCN 478
Query: 430 KQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVT 489
++ A + ++ ++ D+ TYNI+I+G+CK K + L + G+KPN VT
Sbjct: 479 NGKVETALVVFEYLQRSKMEPDIYTYNIMIEGMCKAGKVEDGWDLFCSLSLKGVKPNVVT 538
Query: 490 YNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEM 548
Y T+M G+C +G + A + M++E P+ TYN LI+ + + A+ L+ EM
Sbjct: 539 YTTMMSGFCRKGLKEEADALFREMKEEGPLPDSGTYNTLIRAHLRDGDKAASAELIREM 597
Score = 233 bits (595), Expect = 2e-61
Identities = 140/444 (31%), Positives = 236/444 (52%), Gaps = 29/444 (6%)
Query: 151 HCAY--EAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFN 208
HC++ F+ + K+S+ N L K+ D ++ +M+K R ++ F+
Sbjct: 34 HCSFWVRDFSGVRYDYRKISINRLNDL--------KLDDAVNLFGDMVKSRPFPSIVEFS 85
Query: 209 IFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEM 268
++ + + K + E M+ GIS N+ TY+ L++ +C+R ++ A A + +M
Sbjct: 86 KLLSAIAKMNKFDLVISLGEQMQNLGISHNLYTYSILINCFCRR---SQLSLALAVLAKM 142
Query: 269 LANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKL 328
+ P+ VT N+L++GFC ++ A +M + G +P+ T+N+LI+GL + +
Sbjct: 143 MKLGYEPDIVTLNSLLNGFCHGNRISDAVSLVGQMVEMGYQPDSFTFNTLIHGLFRHNRA 202
Query: 329 EEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTM 388
EA+ L D+MV G +P++VTY ++NG CK+ + A + + + ++ P V+ +NT+
Sbjct: 203 SEAVALVDRMVVKGCQPDLVTYGIVVNGLCKRGDIDLALSLLKKMEQGKIEPGVVIYNTI 262
Query: 389 IDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGL 448
IDA C + + +L + M ++GI PNV TYN LI LC A LL++M + +
Sbjct: 263 IDALCNYKNVNDALNLFTEMDNKGIRPNVVTYNSLIRCLCNYGRWSDASRLLSDMIERKI 322
Query: 449 KGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALN 508
+VVT++ LID K K AEKL +EM + P+ TY++L++G+CM +L A +
Sbjct: 323 NPNVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSIDPDIFTYSSLINGFCMHDRLDEAKH 382
Query: 509 VRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTY-------- 560
+ M + PNVVTYN LIKG+CK +++ L EM ++GL N TY
Sbjct: 383 MFELMISKDCFPNVVTYNTLIKGFCKAKRVDEGMELFREMSQRGLVGNTVTYTTLIHGFF 442
Query: 561 --------DIVRLEMLEKGFSPDI 576
IV +M+ G PDI
Sbjct: 443 QARECDNAQIVFKQMVSDGVLPDI 466
Score = 201 bits (512), Expect = 7e-52
Identities = 113/305 (37%), Positives = 180/305 (58%), Gaps = 9/305 (2%)
Query: 168 SLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAI 227
++ + + L+ A VKE K+ + E +Y EMIKR I ++ T++ ING C +L++A+
Sbjct: 325 NVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSIDPDIFTYSSLINGFCMHDRLDEAKHMF 384
Query: 228 EDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGF 287
E M + PNVVTYNTL+ G+CK A ++ + +EM + N VT+ TLI GF
Sbjct: 385 ELMISKDCFPNVVTYNTLIKGFCK---AKRVDEGMELFREMSQRGLVGNTVTYTTLIHGF 441
Query: 288 CKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNI 347
+ A+ F++M G+ P+I+TY+ L++GLCNNGK+E A+ +++ + ++P+I
Sbjct: 442 FQARECDNAQIVFKQMVSDGVLPDIMTYSILLDGLCNNGKVETALVVFEYLQRSKMEPDI 501
Query: 348 VTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSS 407
TYN +I G CK +++ +F +S + + PNV+T+ TM+ +C++G+ EE +L
Sbjct: 502 YTYNIMIEGMCKAGKVEDGWDLFCSLSLKGVKPNVVTYTTMMSGFCRKGLKEEADALFRE 561
Query: 408 MLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTY----NILIDGLC 463
M +EG LP+ TYN LI R D A+ EL+ EM + GD T N+L DG
Sbjct: 562 MKEEGPLPDSGTYNTLIRAHLRDGDKAASAELIREMRSCRFVGDASTIGLVTNMLHDG-- 619
Query: 464 KNDKS 468
+ DKS
Sbjct: 620 RLDKS 624
>At5g61400 putative protein
Length = 654
Score = 276 bits (707), Expect = 2e-74
Identities = 164/529 (31%), Positives = 262/529 (49%), Gaps = 46/529 (8%)
Query: 72 SQKEYRLSYGLEPTSKVLHFLANSKRYSKVRSFLDSFVK-----NEKHTVSSVFHSLLLD 126
S+ S L+ S V+H L + +Y+ R + S ++ +E +S + L D
Sbjct: 65 SRSRVSKSNDLQSFSAVIHVLTGAHKYTLARCLIKSLIERLKRHSEPSNMSHRLFNALED 124
Query: 127 GGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIG 186
P + + +L++ + LE+ EA +++ +C +L+ LV+ +
Sbjct: 125 IQSPKFSIGVFSLLIMEF---LEMGLFEEALWVSREMKCSPDSKACLSILNGLVRRRRFD 181
Query: 187 DVEYVYKEMIKRR-----------------------------------IHTNLNTFNIFI 211
V Y+ MI R I N+ + I+I
Sbjct: 182 SVWVDYQLMISRGLVPDVHIYFVLFQCCFKQGLYSKKEKLLDEMTSLGIKPNVYIYTIYI 241
Query: 212 NGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLAN 271
LCR K+ +AE E MK G+ PN+ TY+ ++DGYCK G+ + Y KE+L
Sbjct: 242 LDLCRDNKMEEAEKMFELMKKHGVLPNLYTYSAMIDGYCKTGNVRQAY---GLYKEILVA 298
Query: 272 KICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEA 331
++ PN V F TL+DGFCK + A+ F M K G+ PN+ YN LI+G C +G + EA
Sbjct: 299 ELLPNVVVFGTLVDGFCKARELVTARSLFVHMVKFGVDPNLYVYNCLIHGHCKSGNMLEA 358
Query: 332 IDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDA 391
+ L +M L L P++ TY LING C + + EA ++F + + + P+ T+N++I
Sbjct: 359 VGLLSEMESLNLSPDVFTYTILINGLCIEDQVAEANRLFQKMKNERIFPSSATYNSLIHG 418
Query: 392 YCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGD 451
YCKE ME+ LCS M G+ PN+ T++ LI G C +D++AA L EM KG+ D
Sbjct: 419 YCKEYNMEQALDLCSEMTASGVEPNIITFSTLIDGYCNVRDIKAAMGLYFEMTIKGIVPD 478
Query: 452 VVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRT 511
VVTY LID K + A +L ++M G+ PN T+ L+DG+ EG+L A++
Sbjct: 479 VVTYTALIDAHFKEANMKEALRLYSDMLEAGIHPNDHTFACLVDGFWKEGRLSVAIDFYQ 538
Query: 512 RMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTY 560
++R N V + LI+G C+ + A+ ++M G+ P+ +Y
Sbjct: 539 ENNQQRSCWNHVGFTCLIEGLCQNGYILRASRFFSDMRSCGITPDICSY 587
Score = 226 bits (575), Expect = 4e-59
Identities = 145/558 (25%), Positives = 271/558 (47%), Gaps = 43/558 (7%)
Query: 26 TISELLSKQHWSELKPHLRVTKPATFLDQLLNA-----------GVDSELVLRFFK---- 70
T++ L K LK H ++P+ +L NA GV S L++ F +
Sbjct: 92 TLARCLIKSLIERLKRH---SEPSNMSHRLFNALEDIQSPKFSIGVFSLLIMEFLEMGLF 148
Query: 71 ----WSQKEYRLSYGLEPTSKVLHFLANSKRYSKV----RSFLDSFVKNEKHTVSSVFHS 122
W +E + S + +L+ L +R+ V + + + + H +F
Sbjct: 149 EEALWVSREMKCSPDSKACLSILNGLVRRRRFDSVWVDYQLMISRGLVPDVHIYFVLFQC 208
Query: 123 LLLDGGRPGATALIIDMLVLAYVKNLELHCAY--------------EAFTRAKDYGFKLS 168
G L+ +M L N+ ++ Y + F K +G +
Sbjct: 209 CFKQGLYSKKEKLLDEMTSLGIKPNVYIYTIYILDLCRDNKMEEAEKMFELMKKHGVLPN 268
Query: 169 LTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIE 228
L + + ++ K + +YKE++ + N+ F ++G C+A +L A
Sbjct: 269 LYTYSAMIDGYCKTGNVRQAYGLYKEILVAELLPNVVVFGTLVDGFCKARELVTARSLFV 328
Query: 229 DMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFC 288
M +G+ PN+ YN L+ G+CK +G M +A + EM + + P+ T+ LI+G C
Sbjct: 329 HMVKFGVDPNLYVYNCLIHGHCK---SGNMLEAVGLLSEMESLNLSPDVFTYTILINGLC 385
Query: 289 KDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIV 348
++ VA A + F++M+ + + P+ TYNSLI+G C +E+A+DL +M G++PNI+
Sbjct: 386 IEDQVAEANRLFQKMKNERIFPSSATYNSLIHGYCKEYNMEQALDLCSEMTASGVEPNII 445
Query: 349 TYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSM 408
T++ LI+G+C + +K A ++ +++ + +VP+V+T+ +IDA+ KE M+E L S M
Sbjct: 446 TFSTLIDGYCNVRDIKAAMGLYFEMTIKGIVPDVVTYTALIDAHFKEANMKEALRLYSDM 505
Query: 409 LDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKS 468
L+ GI PN T+ CL+ G ++ L A + E + + V + LI+GLC+N
Sbjct: 506 LEAGIHPNDHTFACLVDGFWKEGRLSVAIDFYQENNQQRSCWNHVGFTCLIEGLCQNGYI 565
Query: 469 RNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVL 528
A + ++M + G+ P+ +Y +++ G+ E ++ + ++ M K PN++ +L
Sbjct: 566 LRASRFFSDMRSCGITPDICSYVSMLKGHLQEKRITDTMMLQCDMIKTGILPNLLVNQLL 625
Query: 529 IKGYCKINKLEAANGLLN 546
+ Y +++A L N
Sbjct: 626 ARFYQANGYVKSACFLTN 643
Score = 212 bits (540), Expect = 4e-55
Identities = 124/389 (31%), Positives = 208/389 (52%), Gaps = 22/389 (5%)
Query: 203 NLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAE 262
++ F++ I G +A +MK SP+ ++++G +R ++
Sbjct: 131 SIGVFSLLIMEFLEMGLFEEALWVSREMKC---SPDSKACLSILNGLVRRRRFDSVW--- 184
Query: 263 AFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGL 322
+ M++ + P+ + L K + +K +EM G+KPN+ Y I L
Sbjct: 185 VDYQLMISRGLVPDVHIYFVLFQCCFKQGLYSKKEKLLDEMTSLGIKPNVYIYTIYILDL 244
Query: 323 CNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNV 382
C + K+EEA +++ M G+ PN+ TY+A+I+G+CK +++A ++ ++ EL+PNV
Sbjct: 245 CRDNKMEEAEKMFELMKKHGVLPNLYTYSAMIDGYCKTGNVRQAYGLYKEILVAELLPNV 304
Query: 383 ITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNE 442
+ F T++D +CK + SL M+ G+ PN+ YNCLI G C+ ++ A LL+E
Sbjct: 305 VVFGTLVDGFCKARELVTARSLFVHMVKFGVDPNLYVYNCLIHGHCKSGNMLEAVGLLSE 364
Query: 443 MENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGK 502
ME+ L DV TY ILI+GLC D+ A +L +M N + P+ TYN+L+ GYC E
Sbjct: 365 MESLNLSPDVFTYTILINGLCIEDQVAEANRLFQKMKNERIFPSSATYNSLIHGYCKEYN 424
Query: 503 LKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTY-- 560
++ AL++ + M +PN++T++ LI GYC + ++AA GL EM KG+ P+ TY
Sbjct: 425 MEQALDLCSEMTASGVEPNIITFSTLIDGYCNVRDIKAAMGLYFEMTIKGIVPDVVTYTA 484
Query: 561 ------------DIVRL--EMLEKGFSPD 575
+ +RL +MLE G P+
Sbjct: 485 LIDAHFKEANMKEALRLYSDMLEAGIHPN 513
Score = 116 bits (291), Expect = 3e-26
Identities = 90/364 (24%), Positives = 165/364 (44%), Gaps = 38/364 (10%)
Query: 77 RLSYGLEPTSKVLHFLANSKRYSKVRSFLDSFVK-NEKHTVSSVFHSLLLDGGRPGATAL 135
R +YGL V L N + + +D F K E T S+F ++ G P
Sbjct: 286 RQAYGLYKEILVAELLPNVVVFG---TLVDGFCKARELVTARSLFVHMVKFGVDPNL--Y 340
Query: 136 IIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEM 195
+ + L+ + K+ + A + + + + L++ L E+++ + ++++M
Sbjct: 341 VYNCLIHGHCKSGNMLEAVGLLSEMESLNLSPDVFTYTILINGLCIEDQVAEANRLFQKM 400
Query: 196 IKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCK---- 251
RI + T+N I+G C+ + +A D +M A G+ PN++T++TL+DGYC
Sbjct: 401 KNERIFPSSATYNSLIHGYCKEYNMEQALDLCSEMTASGVEPNIITFSTLIDGYCNVRDI 460
Query: 252 RGSAGKMYKA---------------------EAFMKE-------MLANKICPNEVTFNTL 283
+ + G ++ EA MKE ML I PN+ TF L
Sbjct: 461 KAAMGLYFEMTIKGIVPDVVTYTALIDAHFKEANMKEALRLYSDMLEAGIHPNDHTFACL 520
Query: 284 IDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGL 343
+DGF K+ ++ A ++E +Q N V + LI GLC NG + A + M G+
Sbjct: 521 VDGFWKEGRLSVAIDFYQENNQQRSCWNHVGFTCLIEGLCQNGYILRASRFFSDMRSCGI 580
Query: 344 KPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFS 403
P+I +Y +++ G ++K + + + D+ K ++PN++ + Y G ++
Sbjct: 581 TPDICSYVSMLKGHLQEKRITDTMMLQCDMIKTGILPNLLVNQLLARFYQANGYVKSACF 640
Query: 404 LCSS 407
L +S
Sbjct: 641 LTNS 644
Score = 102 bits (254), Expect = 6e-22
Identities = 60/181 (33%), Positives = 96/181 (52%), Gaps = 2/181 (1%)
Query: 408 MLDEGILPNVSTYNCLIAGLCRKQDLQAAKE-LLNEMENKGLKGDVVTYNILIDGLCKND 466
M+ G++P+V Y L C KQ L + KE LL+EM + G+K +V Y I I LC+++
Sbjct: 190 MISRGLVPDVHIYFVLFQ-CCFKQGLYSKKEKLLDEMTSLGIKPNVYIYTIYILDLCRDN 248
Query: 467 KSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYN 526
K AEK+ M G+ PN TY+ ++DGYC G ++ A + + PNVV +
Sbjct: 249 KMEEAEKMFELMKKHGVLPNLYTYSAMIDGYCKTGNVRQAYGLYKEILVAELLPNVVVFG 308
Query: 527 VLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKGFSPDIEGHLYNISSM 586
L+ G+CK +L A L M++ G++PN Y+ + + G + G L + S+
Sbjct: 309 TLVDGFCKARELVTARSLFVHMVKFGVDPNLYVYNCLIHGHCKSGNMLEAVGLLSEMESL 368
Query: 587 S 587
+
Sbjct: 369 N 369
>At1g62670 PPR-repeat protein
Length = 630
Score = 273 bits (698), Expect = 2e-73
Identities = 139/433 (32%), Positives = 246/433 (56%), Gaps = 3/433 (0%)
Query: 139 MLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKR 198
+L+ + + +L A + G++ ++ + + LL+ +I + + +M
Sbjct: 121 ILINCFCRRSQLPLALAVLGKMMKLGYEPNIVTLSSLLNGYCHSKRISEAVALVDQMFVT 180
Query: 199 RIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKM 258
N TFN I+GL K ++A I+ M A G P++VTY +V+G CKRG
Sbjct: 181 GYQPNTVTFNTLIHGLFLHNKASEAMALIDRMVAKGCQPDLVTYGVVVNGLCKRGDTDLA 240
Query: 259 YKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSL 318
+ + +M K+ P + +NT+IDG CK +++ A F+EM+ +G++PN+VTY+SL
Sbjct: 241 FN---LLNKMEQGKLEPGVLIYNTIIDGLCKYKHMDDALNLFKEMETKGIRPNVVTYSSL 297
Query: 319 INGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQEL 378
I+ LCN G+ +A L M+ + P++ T++ALI+ F K+ + EA K++D++ K+ +
Sbjct: 298 ISCLCNYGRWSDASRLLSDMIERKINPDVFTFSALIDAFVKEGKLVEAEKLYDEMVKRSI 357
Query: 379 VPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKE 438
P+++T++++I+ +C ++E + M+ + P+V TYN LI G C+ + ++ E
Sbjct: 358 DPSIVTYSSLINGFCMHDRLDEAKQMFEFMVSKHCFPDVVTYNTLIKGFCKYKRVEEGME 417
Query: 439 LLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYC 498
+ EM +GL G+ VTYNILI GL + A+++ EM + G+ PN +TYNTL+DG C
Sbjct: 418 VFREMSQRGLVGNTVTYNILIQGLFQAGDCDMAQEIFKEMVSDGVPPNIMTYNTLLDGLC 477
Query: 499 MEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRT 558
GKL+ A+ V +++ + +P + TYN++I+G CK K+E L + KG+ P+
Sbjct: 478 KNGKLEKAMVVFEYLQRSKMEPTIYTYNIMIEGMCKAGKVEDGWDLFCNLSLKGVKPDVV 537
Query: 559 TYDIVRLEMLEKG 571
Y+ + KG
Sbjct: 538 AYNTMISGFCRKG 550
Score = 245 bits (626), Expect = 4e-65
Identities = 153/536 (28%), Positives = 265/536 (48%), Gaps = 51/536 (9%)
Query: 86 SKVLHFLANSKRYSKVRSFLDSF----VKNEKHTVSSVFHSLLLDGGRPGATALIIDMLV 141
SK+L +A ++ V S + + + +T S + + P A A++ M+
Sbjct: 85 SKLLSAIAKMNKFDVVISLGEQMQNLGIPHNHYTYSILINCFCRRSQLPLALAVLGKMMK 144
Query: 142 LAYVKN-------LELHCAYEAFTRAKDY-------GFKLSLTSCNPLLSALVKENKIGD 187
L Y N L +C + + A G++ + + N L+ L NK +
Sbjct: 145 LGYEPNIVTLSSLLNGYCHSKRISEAVALVDQMFVTGYQPNTVTFNTLIHGLFLHNKASE 204
Query: 188 VEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVD 247
+ M+ + +L T+ + +NGLC+ G + A + + M+ + P V+ YNT++D
Sbjct: 205 AMALIDRMVAKGCQPDLVTYGVVVNGLCKRGDTDLAFNLLNKMEQGKLEPGVLIYNTIID 264
Query: 248 GYCK----------------RG----------------SAGKMYKAEAFMKEMLANKICP 275
G CK +G + G+ A + +M+ KI P
Sbjct: 265 GLCKYKHMDDALNLFKEMETKGIRPNVVTYSSLISCLCNYGRWSDASRLLSDMIERKINP 324
Query: 276 NEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLW 335
+ TF+ LID F K+ + A+K ++EM K+ + P+IVTY+SLING C + +L+EA ++
Sbjct: 325 DVFTFSALIDAFVKEGKLVEAEKLYDEMVKRSIDPSIVTYSSLINGFCMHDRLDEAKQMF 384
Query: 336 DKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKE 395
+ MV P++VTYN LI GFCK K ++E +VF ++S++ LV N +T+N +I +
Sbjct: 385 EFMVSKHCFPDVVTYNTLIKGFCKYKRVEEGMEVFREMSQRGLVGNTVTYNILIQGLFQA 444
Query: 396 GMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTY 455
G + + M+ +G+ PN+ TYN L+ GLC+ L+ A + ++ ++ + TY
Sbjct: 445 GDCDMAQEIFKEMVSDGVPPNIMTYNTLLDGLCKNGKLEKAMVVFEYLQRSKMEPTIYTY 504
Query: 456 NILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEK 515
NI+I+G+CK K + L + G+KP+ V YNT++ G+C +G + A + M++
Sbjct: 505 NIMIEGMCKAGKVEDGWDLFCNLSLKGVKPDVVAYNTMISGFCRKGSKEEADALFKEMKE 564
Query: 516 ERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
+ PN YN LI+ + EA+ L+ EM G + +T +V ML G
Sbjct: 565 DGTLPNSGCYNTLIRARLRDGDREASAELIKEMRSCGFAGDASTIGLV-TNMLHDG 619
Score = 238 bits (606), Expect = 9e-63
Identities = 132/392 (33%), Positives = 221/392 (55%), Gaps = 3/392 (0%)
Query: 180 VKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNV 239
+ E K+ D ++ EM+K R ++ F+ ++ + + K + E M+ GI N
Sbjct: 57 LSELKLDDAVALFGEMVKSRPFPSIIEFSKLLSAIAKMNKFDVVISLGEQMQNLGIPHNH 116
Query: 240 VTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKA 299
TY+ L++ +C+R ++ A A + +M+ PN VT ++L++G+C + ++ A
Sbjct: 117 YTYSILINCFCRR---SQLPLALAVLGKMMKLGYEPNIVTLSSLLNGYCHSKRISEAVAL 173
Query: 300 FEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCK 359
++M G +PN VT+N+LI+GL + K EA+ L D+MV G +P++VTY ++NG CK
Sbjct: 174 VDQMFVTGYQPNTVTFNTLIHGLFLHNKASEAMALIDRMVAKGCQPDLVTYGVVVNGLCK 233
Query: 360 KKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVST 419
+ A + + + + +L P V+ +NT+ID CK M++ +L M +GI PNV T
Sbjct: 234 RGDTDLAFNLLNKMEQGKLEPGVLIYNTIIDGLCKYKHMDDALNLFKEMETKGIRPNVVT 293
Query: 420 YNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMF 479
Y+ LI+ LC A LL++M + + DV T++ LID K K AEKL +EM
Sbjct: 294 YSSLISCLCNYGRWSDASRLLSDMIERKINPDVFTFSALIDAFVKEGKLVEAEKLYDEMV 353
Query: 480 NLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLE 539
+ P+ VTY++L++G+CM +L A + M + P+VVTYN LIKG+CK ++E
Sbjct: 354 KRSIDPSIVTYSSLINGFCMHDRLDEAKQMFEFMVSKHCFPDVVTYNTLIKGFCKYKRVE 413
Query: 540 AANGLLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
+ EM ++GL N TY+I+ + + G
Sbjct: 414 EGMEVFREMSQRGLVGNTVTYNILIQGLFQAG 445
Score = 115 bits (287), Expect = 9e-26
Identities = 67/209 (32%), Positives = 111/209 (53%), Gaps = 1/209 (0%)
Query: 355 NGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGIL 414
NG + K+ +A +F ++ K P++I F+ ++ A K + SL M + GI
Sbjct: 55 NGLSELKL-DDAVALFGEMVKSRPFPSIIEFSKLLSAIAKMNKFDVVISLGEQMQNLGIP 113
Query: 415 PNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKL 474
N TY+ LI CR+ L A +L +M G + ++VT + L++G C + + A L
Sbjct: 114 HNHYTYSILINCFCRRSQLPLALAVLGKMMKLGYEPNIVTLSSLLNGYCHSKRISEAVAL 173
Query: 475 LNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCK 534
+++MF G +PN VT+NTL+ G + K A+ + RM + QP++VTY V++ G CK
Sbjct: 174 VDQMFVTGYQPNTVTFNTLIHGLFLHNKASEAMALIDRMVAKGCQPDLVTYGVVVNGLCK 233
Query: 535 INKLEAANGLLNEMLEKGLNPNRTTYDIV 563
+ A LLN+M + L P Y+ +
Sbjct: 234 RGDTDLAFNLLNKMEQGKLEPGVLIYNTI 262
Score = 94.0 bits (232), Expect = 2e-19
Identities = 53/185 (28%), Positives = 97/185 (51%), Gaps = 1/185 (0%)
Query: 398 MEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNI 457
+++ +L M+ P++ ++ L++ + + L +M+N G+ + TY+I
Sbjct: 62 LDDAVALFGEMVKSRPFPSIIEFSKLLSAIAKMNKFDVVISLGEQMQNLGIPHNHYTYSI 121
Query: 458 LIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKER 517
LI+ C+ + A +L +M LG +PN VT ++L++GYC ++ A+ + +M
Sbjct: 122 LINCFCRRSQLPLALAVLGKMMKLGYEPNIVTLSSLLNGYCHSKRISEAVALVDQMFVTG 181
Query: 518 KQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKGFSPDIE 577
QPN VT+N LI G NK A L++ M+ KG P+ TY +V + ++G D+
Sbjct: 182 YQPNTVTFNTLIHGLFLHNKASEAMALIDRMVAKGCQPDLVTYGVVVNGLCKRG-DTDLA 240
Query: 578 GHLYN 582
+L N
Sbjct: 241 FNLLN 245
>At1g62930 unknown protein
Length = 659
Score = 273 bits (697), Expect = 3e-73
Identities = 145/440 (32%), Positives = 242/440 (54%), Gaps = 3/440 (0%)
Query: 138 DMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIK 197
++L+ + + +L A + G++ + + + LL+ +I + + +M
Sbjct: 119 NILINCFCRRSQLPLALAVLGKMMKLGYEPDIVTLSSLLNGYCHGKRISEAVALVDQMFV 178
Query: 198 RRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGK 257
N TFN I+GL K ++A I+ M A G P++ TY T+V+G CKRG
Sbjct: 179 MEYQPNTVTFNTLIHGLFLHNKASEAVALIDRMVARGCQPDLFTYGTVVNGLCKRGDIDL 238
Query: 258 MYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNS 317
A + +K+M KI + V + T+ID C +NV A F EM +G++PN+VTYNS
Sbjct: 239 ---ALSLLKKMEKGKIEADVVIYTTIIDALCNYKNVNDALNLFTEMDNKGIRPNVVTYNS 295
Query: 318 LINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQE 377
LI LCN G+ +A L M+ + PN+VT++ALI+ F K+ + EA K++D++ K+
Sbjct: 296 LIRCLCNYGRWSDASRLLSDMIERKINPNVVTFSALIDAFVKEGKLVEAEKLYDEMIKRS 355
Query: 378 LVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAK 437
+ P++ T++++I+ +C ++E + M+ + PNV TYN LI G C+ + ++
Sbjct: 356 IDPDIFTYSSLINGFCMHDRLDEAKHMFELMISKDCFPNVVTYNTLIKGFCKAKRVEEGM 415
Query: 438 ELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGY 497
EL EM +GL G+ VTYN LI GL + A+K+ +M + G+ P+ +TY+ L+DG
Sbjct: 416 ELFREMSQRGLVGNTVTYNTLIQGLFQAGDCDMAQKIFKKMVSDGVPPDIITYSILLDGL 475
Query: 498 CMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNR 557
C GKL+ AL V ++K + +P++ TYN++I+G CK K+E L + KG+ PN
Sbjct: 476 CKYGKLEKALVVFEYLQKSKMEPDIYTYNIMIEGMCKAGKVEDGWDLFCSLSLKGVKPNV 535
Query: 558 TTYDIVRLEMLEKGFSPDIE 577
Y + KG + +
Sbjct: 536 IIYTTMISGFCRKGLKEEAD 555
Score = 246 bits (628), Expect = 3e-65
Identities = 152/494 (30%), Positives = 255/494 (50%), Gaps = 32/494 (6%)
Query: 78 LSYGLEPT----SKVLHFLANSKRYSKVRSFLDSFVKNEKH----TVSSVFHSLLLDGGR 129
+ G EP S +L+ + KR S+ + +D E T +++ H L L
Sbjct: 142 MKLGYEPDIVTLSSLLNGYCHGKRISEAVALVDQMFVMEYQPNTVTFNTLIHGLFLHNKA 201
Query: 130 PGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVE 189
A ALI M+ C + FT YG +++ L K I
Sbjct: 202 SEAVALIDRMVARG--------CQPDLFT----YG---------TVVNGLCKRGDIDLAL 240
Query: 190 YVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGY 249
+ K+M K +I ++ + I+ LC +N A + +M GI PNVVTYN+L+
Sbjct: 241 SLLKKMEKGKIEADVVIYTTIIDALCNYKNVNDALNLFTEMDNKGIRPNVVTYNSLIRCL 300
Query: 250 CKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLK 309
C + G+ A + +M+ KI PN VTF+ LID F K+ + A+K ++EM K+ +
Sbjct: 301 C---NYGRWSDASRLLSDMIERKINPNVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSID 357
Query: 310 PNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKV 369
P+I TY+SLING C + +L+EA +++ M+ PN+VTYN LI GFCK K ++E ++
Sbjct: 358 PDIFTYSSLINGFCMHDRLDEAKHMFELMISKDCFPNVVTYNTLIKGFCKAKRVEEGMEL 417
Query: 370 FDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCR 429
F ++S++ LV N +T+NT+I + G + + M+ +G+ P++ TY+ L+ GLC+
Sbjct: 418 FREMSQRGLVGNTVTYNTLIQGLFQAGDCDMAQKIFKKMVSDGVPPDIITYSILLDGLCK 477
Query: 430 KQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVT 489
L+ A + ++ ++ D+ TYNI+I+G+CK K + L + G+KPN +
Sbjct: 478 YGKLEKALVVFEYLQKSKMEPDIYTYNIMIEGMCKAGKVEDGWDLFCSLSLKGVKPNVII 537
Query: 490 YNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEML 549
Y T++ G+C +G + A + M+++ PN TYN LI+ + A+ L+ EM
Sbjct: 538 YTTMISGFCRKGLKEEADALFREMKEDGTLPNSGTYNTLIRARLRDGDKAASAELIKEMR 597
Query: 550 EKGLNPNRTTYDIV 563
G + +T ++
Sbjct: 598 SCGFVGDASTISML 611
Score = 222 bits (566), Expect = 4e-58
Identities = 125/388 (32%), Positives = 217/388 (55%), Gaps = 3/388 (0%)
Query: 184 KIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYN 243
K+ D ++ EM++ R ++ FN ++ + + K + E M+ IS ++ +YN
Sbjct: 60 KLDDAVDLFGEMVQSRPLPSIVEFNKLLSAIAKMNKFDLVISLGERMQNLRISYDLYSYN 119
Query: 244 TLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEM 303
L++ +C+R ++ A A + +M+ P+ VT ++L++G+C + ++ A ++M
Sbjct: 120 ILINCFCRR---SQLPLALAVLGKMMKLGYEPDIVTLSSLLNGYCHGKRISEAVALVDQM 176
Query: 304 QKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMM 363
+PN VT+N+LI+GL + K EA+ L D+MV G +P++ TY ++NG CK+ +
Sbjct: 177 FVMEYQPNTVTFNTLIHGLFLHNKASEAVALIDRMVARGCQPDLFTYGTVVNGLCKRGDI 236
Query: 364 KEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCL 423
A + + K ++ +V+ + T+IDA C + + +L + M ++GI PNV TYN L
Sbjct: 237 DLALSLLKKMEKGKIEADVVIYTTIIDALCNYKNVNDALNLFTEMDNKGIRPNVVTYNSL 296
Query: 424 IAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGL 483
I LC A LL++M + + +VVT++ LID K K AEKL +EM +
Sbjct: 297 IRCLCNYGRWSDASRLLSDMIERKINPNVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSI 356
Query: 484 KPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANG 543
P+ TY++L++G+CM +L A ++ M + PNVVTYN LIKG+CK ++E
Sbjct: 357 DPDIFTYSSLINGFCMHDRLDEAKHMFELMISKDCFPNVVTYNTLIKGFCKAKRVEEGME 416
Query: 544 LLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
L EM ++GL N TY+ + + + G
Sbjct: 417 LFREMSQRGLVGNTVTYNTLIQGLFQAG 444
Score = 191 bits (485), Expect = 1e-48
Identities = 118/383 (30%), Positives = 193/383 (49%), Gaps = 22/383 (5%)
Query: 190 YVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGY 249
Y Y+E + R + +L KL+ A D +M P++V +N L+
Sbjct: 45 YDYREKLSRNVLLDL--------------KLDDAVDLFGEMVQSRPLPSIVEFNKLLSAI 90
Query: 250 CKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLK 309
K + M+ + +I + ++N LI+ FC+ + A +M K G +
Sbjct: 91 AKMNKFDLVISLGERMQNL---RISYDLYSYNILINCFCRRSQLPLALAVLGKMMKLGYE 147
Query: 310 PNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKV 369
P+IVT +SL+NG C+ ++ EA+ L D+M + +PN VT+N LI+G EA +
Sbjct: 148 PDIVTLSSLLNGYCHGKRISEAVALVDQMFVMEYQPNTVTFNTLIHGLFLHNKASEAVAL 207
Query: 370 FDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCR 429
D + + P++ T+ T+++ CK G ++ SL M I +V Y +I LC
Sbjct: 208 IDRMVARGCQPDLFTYGTVVNGLCKRGDIDLALSLLKKMEKGKIEADVVIYTTIIDALCN 267
Query: 430 KQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVT 489
+++ A L EM+NKG++ +VVTYN LI LC + +A +LL++M + PN VT
Sbjct: 268 YKNVNDALNLFTEMDNKGIRPNVVTYNSLIRCLCNYGRWSDASRLLSDMIERKINPNVVT 327
Query: 490 YNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEML 549
++ L+D + EGKL A + M K P++ TY+ LI G+C ++L+ A + M+
Sbjct: 328 FSALIDAFVKEGKLVEAEKLYDEMIKRSIDPDIFTYSSLINGFCMHDRLDEAKHMFELMI 387
Query: 550 EKGLNPNRTTYDIVRLEMLEKGF 572
K PN TY+ L KGF
Sbjct: 388 SKDCFPNVVTYN-----TLIKGF 405
Score = 169 bits (428), Expect = 4e-42
Identities = 96/298 (32%), Positives = 158/298 (52%), Gaps = 32/298 (10%)
Query: 168 SLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAI 227
++ + + L+ A VKE K+ + E +Y EMIKR I ++ T++ ING C +L++A+
Sbjct: 324 NVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSIDPDIFTYSSLINGFCMHDRLDEAKHMF 383
Query: 228 EDMKAWGISPNVVTYNTLVDGYCK----------------RG----------------SA 255
E M + PNVVTYNTL+ G+CK RG A
Sbjct: 384 ELMISKDCFPNVVTYNTLIKGFCKAKRVEEGMELFREMSQRGLVGNTVTYNTLIQGLFQA 443
Query: 256 GKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTY 315
G A+ K+M+++ + P+ +T++ L+DG CK + A FE +QK ++P+I TY
Sbjct: 444 GDCDMAQKIFKKMVSDGVPPDIITYSILLDGLCKYGKLEKALVVFEYLQKSKMEPDIYTY 503
Query: 316 NSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSK 375
N +I G+C GK+E+ DL+ + G+KPN++ Y +I+GFC+K + +EA +F ++ +
Sbjct: 504 NIMIEGMCKAGKVEDGWDLFCSLSLKGVKPNVIIYTTMISGFCRKGLKEEADALFREMKE 563
Query: 376 QELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDL 433
+PN T+NT+I A ++G L M G + + ST + L KQ +
Sbjct: 564 DGTLPNSGTYNTLIRARLRDGDKAASAELIKEMRSCGFVGDASTISMLSGNAVLKQSI 621
Score = 95.9 bits (237), Expect = 6e-20
Identities = 58/226 (25%), Positives = 104/226 (45%), Gaps = 3/226 (1%)
Query: 140 LVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRR 199
L+ + K + E F G + + N L+ L + + ++K+M+
Sbjct: 401 LIKGFCKAKRVEEGMELFREMSQRGLVGNTVTYNTLIQGLFQAGDCDMAQKIFKKMVSDG 460
Query: 200 IHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMY 259
+ ++ T++I ++GLC+ GKL KA E ++ + P++ TYN +++G CK AGK+
Sbjct: 461 VPPDIITYSILLDGLCKYGKLEKALVVFEYLQKSKMEPDIYTYNIMIEGMCK---AGKVE 517
Query: 260 KAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLI 319
+ + PN + + T+I GFC+ A F EM++ G PN TYN+LI
Sbjct: 518 DGWDLFCSLSLKGVKPNVIIYTTMISGFCRKGLKEEADALFREMKEDGTLPNSGTYNTLI 577
Query: 320 NGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKE 365
+G + +L +M G + T + L K+ + E
Sbjct: 578 RARLRDGDKAASAELIKEMRSCGFVGDASTISMLSGNAVLKQSILE 623
>At2g16880 putative salt-inducible protein
Length = 743
Score = 272 bits (695), Expect = 4e-73
Identities = 177/569 (31%), Positives = 300/569 (52%), Gaps = 23/569 (4%)
Query: 20 QSLSIPTISELLS--KQHWSE-LKPHL-RVTKPATFLDQLLNA---GVDSELVLRFFKWS 72
+S + T++ +L+ K H+ E L P++ ++T+P L LL++ E ++ FF+W+
Sbjct: 8 ESQLLKTLTSILTSEKTHFLETLNPYIPQITQP--LLTSLLSSPSLAKKPETLVSFFQWA 65
Query: 73 QKEYRLSYGLE---PTSKVLHFLANSKRYSKVRSFLDSFVKNEKHTVSSVFHSLLLDGGR 129
Q ++ + P V+ L + +++ +S L S+++ ++S + +SLL
Sbjct: 66 QTSIPEAFPSDSPLPLISVVRSLLSHHKFADAKSLLVSYIRTSDASLS-LCNSLLHPNLH 124
Query: 130 --PGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVK---ENK 184
P + + D+ + AY+ + H A + F + K +L +CN LL LV+
Sbjct: 125 LSPPPSKALFDIALSAYLHEGKPHVALQIFQKMIRLKLKPNLLTCNTLLIGLVRYPSSFS 184
Query: 185 IGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDM-KAWGISPNVVTYN 243
I V+ +M+K + N+ TFN+ +NG C GKL A +E M + ++P+ VTYN
Sbjct: 185 ISSAREVFDDMVKIGVSLNVQTFNVLVNGYCLEGKLEDALGMLERMVSEFKVNPDNVTYN 244
Query: 244 TLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEM 303
T++ K+ G++ + + +M N + PN VT+N L+ G+CK ++ A + E M
Sbjct: 245 TILKAMSKK---GRLSDLKELLLDMKKNGLVPNRVTYNNLVYGYCKLGSLKEAFQIVELM 301
Query: 304 QKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMM 363
++ + P++ TYN LINGLCN G + E ++L D M L L+P++VTYN LI+G + +
Sbjct: 302 KQTNVLPDLCTYNILINGLCNAGSMREGLELMDAMKSLKLQPDVVTYNTLIDGCFELGLS 361
Query: 364 KEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLD-EGILPNVSTYNC 422
EA K+ + + + N +T N + CKE E ++D G P++ TY+
Sbjct: 362 LEARKLMEQMENDGVKANQVTHNISLKWLCKEEKREAVTRKVKELVDMHGFSPDIVTYHT 421
Query: 423 LIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLG 482
LI + DL A E++ EM KG+K + +T N ++D LCK K A LLN G
Sbjct: 422 LIKAYLKVGDLSGALEMMREMGQKGIKMNTITLNTILDALCKERKLDEAHNLLNSAHKRG 481
Query: 483 LKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAAN 542
+ VTY TL+ G+ E K++ AL + M+K + P V T+N LI G C K E A
Sbjct: 482 FIVDEVTYGTLIMGFFREEKVEKALEMWDEMKKVKITPTVSTFNSLIGGLCHHGKTELAM 541
Query: 543 GLLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
+E+ E GL P+ +T++ + L ++G
Sbjct: 542 EKFDELAESGLLPDDSTFNSIILGYCKEG 570
Score = 198 bits (503), Expect = 8e-51
Identities = 123/424 (29%), Positives = 214/424 (50%), Gaps = 34/424 (8%)
Query: 161 KDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKL 220
K G + + N L+ K + + + + M + + +L T+NI INGLC AG +
Sbjct: 267 KKNGLVPNRVTYNNLVYGYCKLGSLKEAFQIVELMKQTNVLPDLCTYNILINGLCNAGSM 326
Query: 221 NKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFM--------------- 265
+ + ++ MK+ + P+VVTYNTL+DG + G + + K M
Sbjct: 327 REGLELMDAMKSLKLQPDVVTYNTLIDGCFELGLSLEARKLMEQMENDGVKANQVTHNIS 386
Query: 266 ---------KEMLANKI---------CPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQG 307
+E + K+ P+ VT++TLI + K +++ A + EM ++G
Sbjct: 387 LKWLCKEEKREAVTRKVKELVDMHGFSPDIVTYHTLIKAYLKVGDLSGALEMMREMGQKG 446
Query: 308 LKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEAT 367
+K N +T N++++ LC KL+EA +L + G + VTY LI GF +++ +++A
Sbjct: 447 IKMNTITLNTILDALCKERKLDEAHNLLNSAHKRGFIVDEVTYGTLIMGFFREEKVEKAL 506
Query: 368 KVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGL 427
+++D++ K ++ P V TFN++I C G E + + G+LP+ ST+N +I G
Sbjct: 507 EMWDEMKKVKITPTVSTFNSLIGGLCHHGKTELAMEKFDELAESGLLPDDSTFNSIILGY 566
Query: 428 CRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNH 487
C++ ++ A E NE K D T NIL++GLCK + A N + + +
Sbjct: 567 CKEGRVEKAFEFYNESIKHSFKPDNYTCNILLNGLCKEGMTEKALNFFNTLIE-EREVDT 625
Query: 488 VTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNE 547
VTYNT++ +C + KLK A ++ + ME++ +P+ TYN I + KL + LL +
Sbjct: 626 VTYNTMISAFCKDKKLKEAYDLLSEMEEKGLEPDRFTYNSFISLLMEDGKLSETDELLKK 685
Query: 548 MLEK 551
K
Sbjct: 686 FSGK 689
Score = 178 bits (451), Expect = 9e-45
Identities = 110/377 (29%), Positives = 196/377 (51%), Gaps = 17/377 (4%)
Query: 164 GFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRR-IHTNLNTFNIFINGLCRAGKLNK 222
G K + + N L L KE K V KE++ ++ T++ I + G L+
Sbjct: 375 GVKANQVTHNISLKWLCKEEKREAVTRKVKELVDMHGFSPDIVTYHTLIKAYLKVGDLSG 434
Query: 223 AEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNT 282
A + + +M GI N +T NT++D CK K+ +A + +EVT+ T
Sbjct: 435 ALEMMREMGQKGIKMNTITLNTILDALCKER---KLDEAHNLLNSAHKRGFIVDEVTYGT 491
Query: 283 LIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLG 342
LI GF ++E V A + ++EM+K + P + T+NSLI GLC++GK E A++ +D++ G
Sbjct: 492 LIMGFFREEKVEKALEMWDEMKKVKITPTVSTFNSLIGGLCHHGKTELAMEKFDELAESG 551
Query: 343 LKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGF 402
L P+ T+N++I G+CK+ +++A + +++ K P+ T N +++ CKEGM E+
Sbjct: 552 LLPDDSTFNSIILGYCKEGRVEKAFEFYNESIKHSFKPDNYTCNILLNGLCKEGMTEKAL 611
Query: 403 SLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGL 462
+ +++++E + V TYN +I+ C+ + L+ A +LL+EME KGL+ D TYN I L
Sbjct: 612 NFFNTLIEEREVDTV-TYNTMISAFCKDKKLKEAYDLLSEMEEKGLEPDRFTYNSFISLL 670
Query: 463 CKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNV 522
++ K ++LL + ++ + +++ N T KE
Sbjct: 671 MEDGKLSETDELLKK------------FSGKFGSMKRDLQVETEKNPATSESKEELNTEA 718
Query: 523 VTYNVLIKGYCKINKLE 539
+ Y+ +I C +L+
Sbjct: 719 IAYSDVIDELCSRGRLK 735
Score = 145 bits (366), Expect = 6e-35
Identities = 91/328 (27%), Positives = 159/328 (47%), Gaps = 16/328 (4%)
Query: 140 LVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRR 199
L+ AY+K +L A E G K++ + N +L AL KE K+ + + KR
Sbjct: 422 LIKAYLKVGDLSGALEMMREMGQKGIKMNTITLNTILDALCKERKLDEAHNLLNSAHKRG 481
Query: 200 IHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMY 259
+ T+ I G R K+ KA + ++MK I+P V T+N+L+ G C GK
Sbjct: 482 FIVDEVTYGTLIMGFFREEKVEKALEMWDEMKKVKITPTVSTFNSLIGGLCHH---GKTE 538
Query: 260 KAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLI 319
A E+ + + P++ TFN++I G+CK+ V A + + E K KP+ T N L+
Sbjct: 539 LAMEKFDELAESGLLPDDSTFNSIILGYCKEGRVEKAFEFYNESIKHSFKPDNYTCNILL 598
Query: 320 NGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELV 379
NGLC G E+A++ ++ ++ + + VTYN +I+ FCK K +KEA + ++ ++ L
Sbjct: 599 NGLCKEGMTEKALNFFNTLIE-EREVDTVTYNTMISAFCKDKKLKEAYDLLSEMEEKGLE 657
Query: 380 PNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKEL 439
P+ T+N+ I ++G + E L + ++ + R ++ K
Sbjct: 658 PDRFTYNSFISLLMEDGKLSETDEL------------LKKFSGKFGSMKRDLQVETEKNP 705
Query: 440 LNEMENKGLKGDVVTYNILIDGLCKNDK 467
+ L + + Y+ +ID LC +
Sbjct: 706 ATSESKEELNTEAIAYSDVIDELCSRGR 733
Score = 120 bits (302), Expect = 2e-27
Identities = 80/289 (27%), Positives = 138/289 (47%), Gaps = 27/289 (9%)
Query: 135 LIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKE 194
+ ++ ++ A K +L A+ A GF + + L+ +E K+ ++ E
Sbjct: 452 ITLNTILDALCKERKLDEAHNLLNSAHKRGFIVDEVTYGTLIMGFFREEKVEKALEMWDE 511
Query: 195 MIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGS 254
M K +I ++TFN I GLC GK A + +++ G+ P+ T+N+++ GYCK
Sbjct: 512 MKKVKITPTVSTFNSLIGGLCHHGKTELAMEKFDELAESGLLPDDSTFNSIILGYCKE-- 569
Query: 255 AGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVT 314
G++ KA F E + + P+ T N L++G CK+ A F + ++ + + VT
Sbjct: 570 -GRVEKAFEFYNESIKHSFKPDNYTCNILLNGLCKEGMTEKALNFFNTLIEE-REVDTVT 627
Query: 315 YNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALIN------------------- 355
YN++I+ C + KL+EA DL +M GL+P+ TYN+ I+
Sbjct: 628 YNTMISAFCKDKKLKEAYDLLSEMEEKGLEPDRFTYNSFISLLMEDGKLSETDELLKKFS 687
Query: 356 ---GFCKKKMMKEATK-VFDDVSKQELVPNVITFNTMIDAYCKEGMMEE 400
G K+ + E K SK+EL I ++ +ID C G ++E
Sbjct: 688 GKFGSMKRDLQVETEKNPATSESKEELNTEAIAYSDVIDELCSRGRLKE 736
>At5g55840 putative protein
Length = 1274
Score = 271 bits (692), Expect = 1e-72
Identities = 160/504 (31%), Positives = 257/504 (50%), Gaps = 6/504 (1%)
Query: 60 VDSELVLRFFKWSQKEYRLS--YGLEPTSKVLHFLANSKRYSKVRSFLDSFVKNEKHTVS 117
V +L L+F KW K+ L + ++ H L ++ Y R L + S
Sbjct: 48 VHGKLALKFLKWVVKQPGLETDHIVQLVCITTHILVRARMYDPARHILKELSLMSGKS-S 106
Query: 118 SVFHSLLLDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLS 177
VF +L+ + + D+L+ Y++ + + E F YGF S+ +CN +L
Sbjct: 107 FVFGALMTTYRLCNSNPSVYDILIRVYLREGMIQDSLEIFRLMGLYGFNPSVYTCNAILG 166
Query: 178 ALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISP 237
++VK + V KEM+KR+I ++ TFNI IN LC G K+ ++ M+ G +P
Sbjct: 167 SVVKSGEDVSVWSFLKEMLKRKICPDVATFNILINVLCAEGSFEKSSYLMQKMEKSGYAP 226
Query: 238 NVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAK 297
+VTYNT++ YCK+G + A + M + + + T+N LI C+ +A
Sbjct: 227 TIVTYNTVLHWYCKKG---RFKAAIELLDHMKSKGVDADVCTYNMLIHDLCRSNRIAKGY 283
Query: 298 KAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGF 357
+M+K+ + PN VTYN+LING N GK+ A L ++M+ GL PN VT+NALI+G
Sbjct: 284 LLLRDMRKRMIHPNEVTYNTLINGFSNEGKVLIASQLLNEMLSFGLSPNHVTFNALIDGH 343
Query: 358 CKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNV 417
+ KEA K+F + + L P+ +++ ++D CK + M G+
Sbjct: 344 ISEGNFKEALKMFYMMEAKGLTPSEVSYGVLLDGLCKNAEFDLARGFYMRMKRNGVCVGR 403
Query: 418 STYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNE 477
TY +I GLC+ L A LLNEM G+ D+VTY+ LI+G CK + + A++++
Sbjct: 404 ITYTGMIDGLCKNGFLDEAVVLLNEMSKDGIDPDIVTYSALINGFCKVGRFKTAKEIVCR 463
Query: 478 MFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINK 537
++ +GL PN + Y+TL+ C G LK A+ + M E + T+NVL+ CK K
Sbjct: 464 IYRVGLSPNGIIYSTLIYNCCRMGCLKEAIRIYEAMILEGHTRDHFTFNVLVTSLCKAGK 523
Query: 538 LEAANGLLNEMLEKGLNPNRTTYD 561
+ A + M G+ PN ++D
Sbjct: 524 VAEAEEFMRCMTSDGILPNTVSFD 547
Score = 235 bits (599), Expect = 6e-62
Identities = 134/428 (31%), Positives = 221/428 (51%), Gaps = 3/428 (0%)
Query: 144 YVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTN 203
Y K A E K G + + N L+ L + N+I + ++M KR IH N
Sbjct: 238 YCKKGRFKAAIELLDHMKSKGVDADVCTYNMLIHDLCRSNRIAKGYLLLRDMRKRMIHPN 297
Query: 204 LNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEA 263
T+N ING GK+ A + +M ++G+SPN VT+N L+DG+ G+ + K
Sbjct: 298 EVTYNTLINGFSNEGKVLIASQLLNEMLSFGLSPNHVTFNALIDGHISEGNFKEALKMFY 357
Query: 264 FMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLC 323
M+ A + P+EV++ L+DG CK+ A+ + M++ G+ +TY +I+GLC
Sbjct: 358 MME---AKGLTPSEVSYGVLLDGLCKNAEFDLARGFYMRMKRNGVCVGRITYTGMIDGLC 414
Query: 324 NNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVI 383
NG L+EA+ L ++M G+ P+IVTY+ALINGFCK K A ++ + + L PN I
Sbjct: 415 KNGFLDEAVVLLNEMSKDGIDPDIVTYSALINGFCKVGRFKTAKEIVCRIYRVGLSPNGI 474
Query: 384 TFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEM 443
++T+I C+ G ++E + +M+ EG + T+N L+ LC+ + A+E + M
Sbjct: 475 IYSTLIYNCCRMGCLKEAIRIYEAMILEGHTRDHFTFNVLVTSLCKAGKVAEAEEFMRCM 534
Query: 444 ENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKL 503
+ G+ + V+++ LI+G + + A + +EM +G P TY +L+ G C G L
Sbjct: 535 TSDGILPNTVSFDCLINGYGNSGEGLKAFSVFDEMTKVGHHPTFFTYGSLLKGLCKGGHL 594
Query: 504 KAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIV 563
+ A + + V YN L+ CK L A L EM+++ + P+ TY +
Sbjct: 595 REAEKFLKSLHAVPAAVDTVMYNTLLTAMCKSGNLAKAVSLFGEMVQRSILPDSYTYTSL 654
Query: 564 RLEMLEKG 571
+ KG
Sbjct: 655 ISGLCRKG 662
Score = 203 bits (516), Expect = 2e-52
Identities = 119/409 (29%), Positives = 202/409 (49%), Gaps = 38/409 (9%)
Query: 198 RRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGK 257
R ++N + ++I I R G + + + M +G +P+V T N ++ K G
Sbjct: 117 RLCNSNPSVYDILIRVYLREGMIQDSLEIFRLMGLYGFNPSVYTCNAILGSVVKSGEDVS 176
Query: 258 MYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNS 317
++ +F+KEML KICP+ TFN LI+ C + + + ++M+K G P IVTYN+
Sbjct: 177 VW---SFLKEMLKRKICPDVATFNILINVLCAEGSFEKSSYLMQKMEKSGYAPTIVTYNT 233
Query: 318 LINGLCNNGKLEEAIDLWDKMVGLG----------------------------------- 342
+++ C G+ + AI+L D M G
Sbjct: 234 VLHWYCKKGRFKAAIELLDHMKSKGVDADVCTYNMLIHDLCRSNRIAKGYLLLRDMRKRM 293
Query: 343 LKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGF 402
+ PN VTYN LINGF + + A+++ +++ L PN +TFN +ID + EG +E
Sbjct: 294 IHPNEVTYNTLINGFSNEGKVLIASQLLNEMLSFGLSPNHVTFNALIDGHISEGNFKEAL 353
Query: 403 SLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGL 462
+ M +G+ P+ +Y L+ GLC+ + A+ M+ G+ +TY +IDGL
Sbjct: 354 KMFYMMEAKGLTPSEVSYGVLLDGLCKNAEFDLARGFYMRMKRNGVCVGRITYTGMIDGL 413
Query: 463 CKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNV 522
CKN A LLNEM G+ P+ VTY+ L++G+C G+ K A + R+ + PN
Sbjct: 414 CKNGFLDEAVVLLNEMSKDGIDPDIVTYSALINGFCKVGRFKTAKEIVCRIYRVGLSPNG 473
Query: 523 VTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
+ Y+ LI C++ L+ A + M+ +G + T++++ + + G
Sbjct: 474 IIYSTLIYNCCRMGCLKEAIRIYEAMILEGHTRDHFTFNVLVTSLCKAG 522
Score = 170 bits (431), Expect = 2e-42
Identities = 114/436 (26%), Positives = 203/436 (46%), Gaps = 6/436 (1%)
Query: 126 DGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKI 185
DG P + D L+ Y + E A+ F G + + LL L K +
Sbjct: 537 DGILPNTVSF--DCLINGYGNSGEGLKAFSVFDEMTKVGHHPTFFTYGSLLKGLCKGGHL 594
Query: 186 GDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTL 245
+ E K + + +N + +C++G L KA +M I P+ TY +L
Sbjct: 595 REAEKFLKSLHAVPAAVDTVMYNTLLTAMCKSGNLAKAVSLFGEMVQRSILPDSYTYTSL 654
Query: 246 VDGYCKRGSAGKMYKAEAFMKEMLAN-KICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQ 304
+ G C++G K A F KE A + PN+V + +DG K A E+M
Sbjct: 655 ISGLCRKG---KTVIAILFAKEAEARGNVLPNKVMYTCFVDGMFKAGQWKAGIYFREQMD 711
Query: 305 KQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMK 364
G P+IVT N++I+G GK+E+ DL +M PN+ TYN L++G+ K+K +
Sbjct: 712 NLGHTPDIVTTNAMIDGYSRMGKIEKTNDLLPEMGNQNGGPNLTTYNILLHGYSKRKDVS 771
Query: 365 EATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLI 424
+ ++ + ++P+ +T ++++ C+ M+E G + + + G+ + T+N LI
Sbjct: 772 TSFLLYRSIILNGILPDKLTCHSLVLGICESNMLEIGLKILKAFICRGVEVDRYTFNMLI 831
Query: 425 AGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLK 484
+ C ++ A +L+ M + G+ D T + ++ L +N + + + +L+EM G+
Sbjct: 832 SKCCANGEINWAFDLVKVMTSLGISLDKDTCDAMVSVLNRNHRFQESRMVLHEMSKQGIS 891
Query: 485 PNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGL 544
P Y L++G C G +K A V+ M + P V + +++ K K + A L
Sbjct: 892 PESRKYIGLINGLCRVGDIKTAFVVKEEMIAHKICPPNVAESAMVRALAKCGKADEATLL 951
Query: 545 LNEMLEKGLNPNRTTY 560
L ML+ L P ++
Sbjct: 952 LRFMLKMKLVPTIASF 967
Score = 169 bits (429), Expect = 3e-42
Identities = 99/329 (30%), Positives = 163/329 (49%), Gaps = 23/329 (6%)
Query: 268 MLANKIC-PNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNG 326
M ++C N ++ LI + ++ + + + F M G P++ T N+++ + +G
Sbjct: 113 MTTYRLCNSNPSVYDILIRVYLREGMIQDSLEIFRLMGLYGFNPSVYTCNAILGSVVKSG 172
Query: 327 KLEEAIDLWD---KMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVI 383
E + +W +M+ + P++ T+N LIN C + ++++ + + K P ++
Sbjct: 173 ---EDVSVWSFLKEMLKRKICPDVATFNILINVLCAEGSFEKSSYLMQKMEKSGYAPTIV 229
Query: 384 TFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEM 443
T+NT++ YCK+G + L M +G+ +V TYN LI LCR + LL +M
Sbjct: 230 TYNTVLHWYCKKGRFKAAIELLDHMKSKGVDADVCTYNMLIHDLCRSNRIAKGYLLLRDM 289
Query: 444 ENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKL 503
+ + + VTYN LI+G K A +LLNEM + GL PNHVT+N L+DG+ EG
Sbjct: 290 RKRMIHPNEVTYNTLINGFSNEGKVLIASQLLNEMLSFGLSPNHVTFNALIDGHISEGNF 349
Query: 504 KAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYD-- 561
K AL + ME + P+ V+Y VL+ G CK + + A G M G+ R TY
Sbjct: 350 KEALKMFYMMEAKGLTPSEVSYGVLLDGLCKNAEFDLARGFYMRMKRNGVCVGRITYTGM 409
Query: 562 --------------IVRLEMLEKGFSPDI 576
++ EM + G PDI
Sbjct: 410 IDGLCKNGFLDEAVVLLNEMSKDGIDPDI 438
Score = 156 bits (394), Expect = 3e-38
Identities = 123/454 (27%), Positives = 197/454 (43%), Gaps = 58/454 (12%)
Query: 173 NPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKA 232
N LL+A+ K + ++ EM++R I + T+ I+GLCR GK A ++ +A
Sbjct: 617 NTLLTAMCKSGNLAKAVSLFGEMVQRSILPDSYTYTSLISGLCRKGKTVIAILFAKEAEA 676
Query: 233 WG-ISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANK-ICPNEVTFNTLIDGFCKD 290
G + PN V Y VDG K G +KA + +E + N P+ VT N +IDG+ +
Sbjct: 677 RGNVLPNKVMYTCFVDGMFKAGQ----WKAGIYFREQMDNLGHTPDIVTTNAMIDGYSRM 732
Query: 291 ENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTY 350
+ EM Q PN+ TYN L++G + + L+ ++ G+ P+ +T
Sbjct: 733 GKIEKTNDLLPEMGNQNGGPNLTTYNILLHGYSKRKDVSTSFLLYRSIILNGILPDKLTC 792
Query: 351 NALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLD 410
++L+ G C+ M++ K+ + + + TFN +I C G + F L M
Sbjct: 793 HSLVLGICESNMLEIGLKILKAFICRGVEVDRYTFNMLISKCCANGEINWAFDLVKVMTS 852
Query: 411 EGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLC------- 463
GI + T + +++ L R Q ++ +L+EM +G+ + Y LI+GLC
Sbjct: 853 LGISLDKDTCDAMVSVLNRNHRFQESRMVLHEMSKQGISPESRKYIGLINGLCRVGDIKT 912
Query: 464 ----------------------------KNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMD 495
K K+ A LL M + L P ++ TLM
Sbjct: 913 AFVVKEEMIAHKICPPNVAESAMVRALAKCGKADEATLLLRFMLKMKLVPTIASFTTLMH 972
Query: 496 GYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNP 555
C G + AL +R M + ++V+YNVLI G C + A L EM G
Sbjct: 973 LCCKNGNVIEALELRVVMSNCGLKLDLVSYNVLITGLCAKGDMALAFELYEEMKGDGFLA 1032
Query: 556 NRTTY-----------------DIVRLEMLEKGF 572
N TTY DI+ ++L +GF
Sbjct: 1033 NATTYKALIRGLLARETAFSGADIILKDLLARGF 1066
Score = 144 bits (363), Expect = 1e-34
Identities = 96/363 (26%), Positives = 172/363 (46%), Gaps = 5/363 (1%)
Query: 192 YKEMIKRRIHT-NLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYC 250
++E + HT ++ T N I+G R GK+ K D + +M PN+ TYN L+ GY
Sbjct: 706 FREQMDNLGHTPDIVTTNAMIDGYSRMGKIEKTNDLLPEMGNQNGGPNLTTYNILLHGYS 765
Query: 251 KRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKP 310
KR + + ++ N I P+++T ++L+ G C+ + K + +G++
Sbjct: 766 KRKDVSTSF---LLYRSIILNGILPDKLTCHSLVLGICESNMLEIGLKILKAFICRGVEV 822
Query: 311 NIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVF 370
+ T+N LI+ C NG++ A DL M LG+ + T +A+++ + +E+ V
Sbjct: 823 DRYTFNMLISKCCANGEINWAFDLVKVMTSLGISLDKDTCDAMVSVLNRNHRFQESRMVL 882
Query: 371 DDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRK 430
++SKQ + P + +I+ C+ G ++ F + M+ I P + ++ L +
Sbjct: 883 HEMSKQGISPESRKYIGLINGLCRVGDIKTAFVVKEEMIAHKICPPNVAESAMVRALAKC 942
Query: 431 QDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTY 490
A LL M L + ++ L+ CKN A +L M N GLK + V+Y
Sbjct: 943 GKADEATLLLRFMLKMKLVPTIASFTTLMHLCCKNGNVIEALELRVVMSNCGLKLDLVSY 1002
Query: 491 NTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKG-YCKINKLEAANGLLNEML 549
N L+ G C +G + A + M+ + N TY LI+G + A+ +L ++L
Sbjct: 1003 NVLITGLCAKGDMALAFELYEEMKGDGFLANATTYKALIRGLLARETAFSGADIILKDLL 1062
Query: 550 EKG 552
+G
Sbjct: 1063 ARG 1065
Score = 29.6 bits (65), Expect = 4.9
Identities = 21/72 (29%), Positives = 33/72 (45%), Gaps = 1/72 (1%)
Query: 164 GFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFING-LCRAGKLNK 222
G KL L S N L++ L + + +Y+EM N T+ I G L R +
Sbjct: 994 GLKLDLVSYNVLITGLCAKGDMALAFELYEEMKGDGFLANATTYKALIRGLLARETAFSG 1053
Query: 223 AEDAIEDMKAWG 234
A+ ++D+ A G
Sbjct: 1054 ADIILKDLLARG 1065
>At1g63080 unknown protein
Length = 614
Score = 270 bits (690), Expect = 2e-72
Identities = 150/490 (30%), Positives = 262/490 (52%), Gaps = 20/490 (4%)
Query: 86 SKVLHFLANSKRYSKVRSFLDSFVKNEKHTVSSVFHSLLLDGGRPGATALIIDMLVLAYV 145
SK+L +A K++ V SF EK + V H+L ++++
Sbjct: 69 SKLLSAIAKMKKFDLVISF------GEKMEILGVSHNLYT-----------YNIMINCLC 111
Query: 146 KNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLN 205
+ +L A + G+ S+ + N LL+ N+I + + +M++ +
Sbjct: 112 RRSQLSFALAILGKMMKLGYGPSIVTLNSLLNGFCHGNRISEAVALVDQMVEMGYQPDTV 171
Query: 206 TFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFM 265
TF ++GL + K ++A +E M G P++VTY +++G CKRG A +
Sbjct: 172 TFTTLVHGLFQHNKASEAVALVERMVVKGCQPDLVTYGAVINGLCKRGEPDL---ALNLL 228
Query: 266 KEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNN 325
+M KI + V ++T+ID CK +V A F EM +G++P++ TY+SLI+ LCN
Sbjct: 229 NKMEKGKIEADVVIYSTVIDSLCKYRHVDDALNLFTEMDNKGIRPDVFTYSSLISCLCNY 288
Query: 326 GKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITF 385
G+ +A L M+ + PN+VT+N+LI+ F K+ + EA K+FD++ ++ + PN++T+
Sbjct: 289 GRWSDASRLLSDMLERKINPNVVTFNSLIDAFAKEGKLIEAEKLFDEMIQRSIDPNIVTY 348
Query: 386 NTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMEN 445
N++I+ +C ++E + + M+ + LP+V TYN LI G C+ + + EL +M
Sbjct: 349 NSLINGFCMHDRLDEAQQIFTLMVSKDCLPDVVTYNTLINGFCKAKKVVDGMELFRDMSR 408
Query: 446 KGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKA 505
+GL G+ VTY LI G + NA+ + +M + G+ PN +TYNTL+DG C GKL+
Sbjct: 409 RGLVGNTVTYTTLIHGFFQASDCDNAQMVFKQMVSDGVHPNIMTYNTLLDGLCKNGKLEK 468
Query: 506 ALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRL 565
A+ V ++K + +P++ TYN++ +G CK K+E L + KG+ P+ Y+ +
Sbjct: 469 AMVVFEYLQKSKMEPDIYTYNIMSEGMCKAGKVEDGWDLFCSLSLKGVKPDVIAYNTMIS 528
Query: 566 EMLEKGFSPD 575
+KG +
Sbjct: 529 GFCKKGLKEE 538
Score = 232 bits (592), Expect = 4e-61
Identities = 126/377 (33%), Positives = 216/377 (56%), Gaps = 3/377 (0%)
Query: 184 KIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYN 243
K+ + ++ EM+K R ++ F+ ++ + + K + E M+ G+S N+ TYN
Sbjct: 45 KLDEAVDLFGEMVKSRPFPSIVEFSKLLSAIAKMKKFDLVISFGEKMEILGVSHNLYTYN 104
Query: 244 TLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEM 303
+++ C+R ++ A A + +M+ P+ VT N+L++GFC ++ A ++M
Sbjct: 105 IMINCLCRR---SQLSFALAILGKMMKLGYGPSIVTLNSLLNGFCHGNRISEAVALVDQM 161
Query: 304 QKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMM 363
+ G +P+ VT+ +L++GL + K EA+ L ++MV G +P++VTY A+ING CK+
Sbjct: 162 VEMGYQPDTVTFTTLVHGLFQHNKASEAVALVERMVVKGCQPDLVTYGAVINGLCKRGEP 221
Query: 364 KEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCL 423
A + + + K ++ +V+ ++T+ID+ CK +++ +L + M ++GI P+V TY+ L
Sbjct: 222 DLALNLLNKMEKGKIEADVVIYSTVIDSLCKYRHVDDALNLFTEMDNKGIRPDVFTYSSL 281
Query: 424 IAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGL 483
I+ LC A LL++M + + +VVT+N LID K K AEKL +EM +
Sbjct: 282 ISCLCNYGRWSDASRLLSDMLERKINPNVVTFNSLIDAFAKEGKLIEAEKLFDEMIQRSI 341
Query: 484 KPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANG 543
PN VTYN+L++G+CM +L A + T M + P+VVTYN LI G+CK K+
Sbjct: 342 DPNIVTYNSLINGFCMHDRLDEAQQIFTLMVSKDCLPDVVTYNTLINGFCKAKKVVDGME 401
Query: 544 LLNEMLEKGLNPNRTTY 560
L +M +GL N TY
Sbjct: 402 LFRDMSRRGLVGNTVTY 418
Score = 186 bits (473), Expect = 2e-47
Identities = 106/320 (33%), Positives = 173/320 (53%), Gaps = 5/320 (1%)
Query: 247 DGYCKRGSAG-----KMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFE 301
DGY ++ S K+ +A EM+ ++ P+ V F+ L+ K + E
Sbjct: 30 DGYREKLSRNALLHLKLDEAVDLFGEMVKSRPFPSIVEFSKLLSAIAKMKKFDLVISFGE 89
Query: 302 EMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKK 361
+M+ G+ N+ TYN +IN LC +L A+ + KM+ LG P+IVT N+L+NGFC
Sbjct: 90 KMEILGVSHNLYTYNIMINCLCRRSQLSFALAILGKMMKLGYGPSIVTLNSLLNGFCHGN 149
Query: 362 MMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYN 421
+ EA + D + + P+ +TF T++ + E +L M+ +G P++ TY
Sbjct: 150 RISEAVALVDQMVEMGYQPDTVTFTTLVHGLFQHNKASEAVALVERMVVKGCQPDLVTYG 209
Query: 422 CLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNL 481
+I GLC++ + A LLN+ME ++ DVV Y+ +ID LCK +A L EM N
Sbjct: 210 AVINGLCKRGEPDLALNLLNKMEKGKIEADVVIYSTVIDSLCKYRHVDDALNLFTEMDNK 269
Query: 482 GLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAA 541
G++P+ TY++L+ C G+ A + + M + + PNVVT+N LI + K KL A
Sbjct: 270 GIRPDVFTYSSLISCLCNYGRWSDASRLLSDMLERKINPNVVTFNSLIDAFAKEGKLIEA 329
Query: 542 NGLLNEMLEKGLNPNRTTYD 561
L +EM+++ ++PN TY+
Sbjct: 330 EKLFDEMIQRSIDPNIVTYN 349
Score = 106 bits (264), Expect = 4e-23
Identities = 71/231 (30%), Positives = 112/231 (47%), Gaps = 18/231 (7%)
Query: 363 MKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNC 422
+ EA +F ++ K P+++ F+ ++ A K + S M G+ N+ TYN
Sbjct: 46 LDEAVDLFGEMVKSRPFPSIVEFSKLLSAIAKMKKFDLVISFGEKMEILGVSHNLYTYNI 105
Query: 423 LIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLG 482
+I LCR+ L A +L +M G +VT N L++G C ++ A L+++M +G
Sbjct: 106 MINCLCRRSQLSFALAILGKMMKLGYGPSIVTLNSLLNGFCHGNRISEAVALVDQMVEMG 165
Query: 483 LKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAAN 542
+P+ VT+ TL+ G K A+ + RM + QP++VTY +I G CK + + A
Sbjct: 166 YQPDTVTFTTLVHGLFQHNKASEAVALVERMVVKGCQPDLVTYGAVINGLCKRGEPDLAL 225
Query: 543 GLLNEMLEKG---------------LNPNRTTYDIVRL--EMLEKGFSPDI 576
LLN+M EKG L R D + L EM KG PD+
Sbjct: 226 NLLNKM-EKGKIEADVVIYSTVIDSLCKYRHVDDALNLFTEMDNKGIRPDV 275
Score = 30.8 bits (68), Expect = 2.2
Identities = 17/62 (27%), Positives = 32/62 (51%)
Query: 502 KLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYD 561
KL A+++ M K R P++V ++ L+ K+ K + +M G++ N TY+
Sbjct: 45 KLDEAVDLFGEMVKSRPFPSIVEFSKLLSAIAKMKKFDLVISFGEKMEILGVSHNLYTYN 104
Query: 562 IV 563
I+
Sbjct: 105 IM 106
>At1g62910 unknown protein
Length = 1133
Score = 266 bits (681), Expect = 2e-71
Identities = 138/442 (31%), Positives = 244/442 (54%), Gaps = 3/442 (0%)
Query: 139 MLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKR 198
+ + + + +L A + G++ + + + LL+ +I D + +M++
Sbjct: 123 IFINCFCRRSQLSLALAVLAKMMKLGYEPDIVTLSSLLNGYCHSKRISDAVALVDQMVEM 182
Query: 199 RIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKM 258
+ TF I+GL K ++A ++ M G P++VTY T+V+G CKRG
Sbjct: 183 GYKPDTFTFTTLIHGLFLHNKASEAVALVDQMVQRGCQPDLVTYGTVVNGLCKRGDIDL- 241
Query: 259 YKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSL 318
A + +K+M KI + V +NT+IDG CK +++ A F EM +G++P++ TY+SL
Sbjct: 242 --ALSLLKKMEKGKIEADVVIYNTIIDGLCKYKHMDDALNLFTEMDNKGIRPDVFTYSSL 299
Query: 319 INGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQEL 378
I+ LCN G+ +A L M+ + PN+VT++ALI+ F K+ + EA K++D++ K+ +
Sbjct: 300 ISCLCNYGRWSDASRLLSDMIERKINPNVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSI 359
Query: 379 VPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKE 438
P++ T++++I+ +C ++E + M+ + PNV TY+ LI G C+ + ++ E
Sbjct: 360 DPDIFTYSSLINGFCMHDRLDEAKHMFELMISKDCFPNVVTYSTLIKGFCKAKRVEEGME 419
Query: 439 LLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYC 498
L EM +GL G+ VTY LI G + NA+ + +M ++G+ PN +TYN L+DG C
Sbjct: 420 LFREMSQRGLVGNTVTYTTLIHGFFQARDCDNAQMVFKQMVSVGVHPNILTYNILLDGLC 479
Query: 499 MEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRT 558
GKL A+ V +++ +P++ TYN++I+G CK K+E L + KG++PN
Sbjct: 480 KNGKLAKAMVVFEYLQRSTMEPDIYTYNIMIEGMCKAGKVEDGWELFCNLSLKGVSPNVI 539
Query: 559 TYDIVRLEMLEKGFSPDIEGHL 580
Y+ + KG + + L
Sbjct: 540 AYNTMISGFCRKGSKEEADSLL 561
Score = 257 bits (656), Expect = 1e-68
Identities = 144/452 (31%), Positives = 242/452 (52%), Gaps = 20/452 (4%)
Query: 86 SKVLHFLANSKRYSKVRSFLDSFVKNEKHTVSSVFHSLLLDGGRPGATALIIDMLVLAYV 145
SK+L +A ++ V SF EK + + H+L ++L+ +
Sbjct: 687 SKLLSAIAKMNKFDLVISF------GEKMEILGISHNLYT-----------YNILINCFC 729
Query: 146 KNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLN 205
+ L A + G++ + + N LL+ N+I D + +M++ +
Sbjct: 730 RCSRLSLALALLGKMMKLGYEPDIVTLNSLLNGFCHGNRISDAVALVDQMVEMGYKPDTV 789
Query: 206 TFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFM 265
TF I+GL K ++A I+ M G P++VTY +V+G CKRG A +
Sbjct: 790 TFTTLIHGLFLHNKASEAVALIDRMVQRGCQPDLVTYGAVVNGLCKRGDTDL---ALNLL 846
Query: 266 KEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNN 325
+M A KI N V ++T+ID CK + A F EM+ +G++PN++TY+SLI+ LCN
Sbjct: 847 NKMEAAKIEANVVIYSTVIDSLCKYRHEDDALNLFTEMENKGVRPNVITYSSLISCLCNY 906
Query: 326 GKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITF 385
G+ +A L M+ + PN+VT++ALI+ F KK + +A K+++++ K+ + PN+ T+
Sbjct: 907 GRWSDASRLLSDMIERKINPNLVTFSALIDAFVKKGKLVKAEKLYEEMIKRSIDPNIFTY 966
Query: 386 NTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMEN 445
+++I+ +C + E + M+ + LPNV TYN LI G C+ + + EL EM
Sbjct: 967 SSLINGFCMLDRLGEAKQMLELMIRKDCLPNVVTYNTLINGFCKAKRVDKGMELFREMSQ 1026
Query: 446 KGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKA 505
+GL G+ VTY LI G + NA+ + +M ++G+ PN +TYN L+DG C GKL
Sbjct: 1027 RGLVGNTVTYTTLIHGFFQARDCDNAQMVFKQMVSVGVHPNILTYNILLDGLCKNGKLAK 1086
Query: 506 ALNVRTRMEKERKQPNVVTYNVLIKGYCKINK 537
A+ V +++ +P++ TYN++I+G CK K
Sbjct: 1087 AMVVFEYLQRSTMEPDIYTYNIMIEGMCKAGK 1118
Score = 251 bits (641), Expect = 8e-67
Identities = 148/490 (30%), Positives = 256/490 (52%), Gaps = 32/490 (6%)
Query: 78 LSYGLEPT----SKVLHFLANSKRYSKVRSFLDSFV----KNEKHTVSSVFHSLLLDGGR 129
+ G EP S +L+ +SKR S + +D V K + T +++ H L L
Sbjct: 145 MKLGYEPDIVTLSSLLNGYCHSKRISDAVALVDQMVEMGYKPDTFTFTTLIHGLFLHNKA 204
Query: 130 PGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVE 189
A AL+ M+ G + L + +++ L K I
Sbjct: 205 SEAVALVDQMV---------------------QRGCQPDLVTYGTVVNGLCKRGDIDLAL 243
Query: 190 YVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGY 249
+ K+M K +I ++ +N I+GLC+ ++ A + +M GI P+V TY++L+
Sbjct: 244 SLLKKMEKGKIEADVVIYNTIIDGLCKYKHMDDALNLFTEMDNKGIRPDVFTYSSLISCL 303
Query: 250 CKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLK 309
C + G+ A + +M+ KI PN VTF+ LID F K+ + A+K ++EM K+ +
Sbjct: 304 C---NYGRWSDASRLLSDMIERKINPNVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSID 360
Query: 310 PNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKV 369
P+I TY+SLING C + +L+EA +++ M+ PN+VTY+ LI GFCK K ++E ++
Sbjct: 361 PDIFTYSSLINGFCMHDRLDEAKHMFELMISKDCFPNVVTYSTLIKGFCKAKRVEEGMEL 420
Query: 370 FDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCR 429
F ++S++ LV N +T+ T+I + + + + M+ G+ PN+ TYN L+ GLC+
Sbjct: 421 FREMSQRGLVGNTVTYTTLIHGFFQARDCDNAQMVFKQMVSVGVHPNILTYNILLDGLCK 480
Query: 430 KQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVT 489
L A + ++ ++ D+ TYNI+I+G+CK K + +L + G+ PN +
Sbjct: 481 NGKLAKAMVVFEYLQRSTMEPDIYTYNIMIEGMCKAGKVEDGWELFCNLSLKGVSPNVIA 540
Query: 490 YNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEML 549
YNT++ G+C +G + A ++ +M+++ PN TYN LI+ + EA+ L+ EM
Sbjct: 541 YNTMISGFCRKGSKEEADSLLKKMKEDGPLPNSGTYNTLIRARLRDGDREASAELIKEMR 600
Query: 550 EKGLNPNRTT 559
G + +T
Sbjct: 601 SCGFAGDAST 610
Score = 232 bits (591), Expect = 5e-61
Identities = 143/445 (32%), Positives = 228/445 (51%), Gaps = 48/445 (10%)
Query: 168 SLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAI 227
++ + + L+ A VKE K+ + E +Y EMIKR I ++ T++ ING C +L++A+
Sbjct: 327 NVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSIDPDIFTYSSLINGFCMHDRLDEAKHMF 386
Query: 228 EDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGF 287
E M + PNVVTY+TL+ G+CK A ++ + +EM + N VT+ TLI GF
Sbjct: 387 ELMISKDCFPNVVTYSTLIKGFCK---AKRVEEGMELFREMSQRGLVGNTVTYTTLIHGF 443
Query: 288 CKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNI 347
+ + A+ F++M G+ PNI+TYN L++GLC NGKL +A+ +++ + ++P+I
Sbjct: 444 FQARDCDNAQMVFKQMVSVGVHPNILTYNILLDGLCKNGKLAKAMVVFEYLQRSTMEPDI 503
Query: 348 VTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSS 407
TYN +I G CK +++ ++F ++S + + PNVI +NTMI +C++G EE SL
Sbjct: 504 YTYNIMIEGMCKAGKVEDGWELFCNLSLKGVSPNVIAYNTMISGFCRKGSKEEADSLLKK 563
Query: 408 MLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILI-------- 459
M ++G LPN TYN LI R D +A+ EL+ EM + G GD T +
Sbjct: 564 MKEDGPLPNSGTYNTLIRARLRDGDREASAELIKEMRSCGFAGDASTIGLSSKFKREIEK 623
Query: 460 ---DGLCKN---------DKSRNAEKLLNEMFNLGLK--------------------PNH 487
+ CKN +S E+ GL P+
Sbjct: 624 PNRENACKNIIFSEEICSSESCGESYDYREVLRTGLSDIELDDAIGLFGVMAQSRPFPSI 683
Query: 488 VTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNE 547
+ ++ L+ K ++ +ME N+ TYN+LI +C+ ++L A LL +
Sbjct: 684 IEFSKLLSAIAKMNKFDLVISFGEKMEILGISHNLYTYNILINCFCRCSRLSLALALLGK 743
Query: 548 MLEKGLNPNRTTYDIVRLEMLEKGF 572
M++ G P DIV L L GF
Sbjct: 744 MMKLGYEP-----DIVTLNSLLNGF 763
Score = 226 bits (577), Expect = 2e-59
Identities = 123/377 (32%), Positives = 215/377 (56%), Gaps = 3/377 (0%)
Query: 184 KIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYN 243
K+ D ++ +M+K R ++ FN ++ + + K E M+ GIS ++ TY+
Sbjct: 63 KVDDAVDLFGDMVKSRPFPSIVEFNKLLSAVAKMNKFELVISLGEQMQTLGISHDLYTYS 122
Query: 244 TLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEM 303
++ +C+R ++ A A + +M+ P+ VT ++L++G+C + ++ A ++M
Sbjct: 123 IFINCFCRR---SQLSLALAVLAKMMKLGYEPDIVTLSSLLNGYCHSKRISDAVALVDQM 179
Query: 304 QKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMM 363
+ G KP+ T+ +LI+GL + K EA+ L D+MV G +P++VTY ++NG CK+ +
Sbjct: 180 VEMGYKPDTFTFTTLIHGLFLHNKASEAVALVDQMVQRGCQPDLVTYGTVVNGLCKRGDI 239
Query: 364 KEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCL 423
A + + K ++ +V+ +NT+ID CK M++ +L + M ++GI P+V TY+ L
Sbjct: 240 DLALSLLKKMEKGKIEADVVIYNTIIDGLCKYKHMDDALNLFTEMDNKGIRPDVFTYSSL 299
Query: 424 IAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGL 483
I+ LC A LL++M + + +VVT++ LID K K AEKL +EM +
Sbjct: 300 ISCLCNYGRWSDASRLLSDMIERKINPNVVTFSALIDAFVKEGKLVEAEKLYDEMIKRSI 359
Query: 484 KPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANG 543
P+ TY++L++G+CM +L A ++ M + PNVVTY+ LIKG+CK ++E
Sbjct: 360 DPDIFTYSSLINGFCMHDRLDEAKHMFELMISKDCFPNVVTYSTLIKGFCKAKRVEEGME 419
Query: 544 LLNEMLEKGLNPNRTTY 560
L EM ++GL N TY
Sbjct: 420 LFREMSQRGLVGNTVTY 436
Score = 224 bits (571), Expect = 1e-58
Identities = 128/409 (31%), Positives = 223/409 (54%), Gaps = 12/409 (2%)
Query: 152 CAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFI 211
C+ E+ + DY +L + + ++ D ++ M + R ++ F+ +
Sbjct: 640 CSSESCGESYDY---------REVLRTGLSDIELDDAIGLFGVMAQSRPFPSIIEFSKLL 690
Query: 212 NGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLAN 271
+ + + K + E M+ GIS N+ TYN L++ +C+ ++ A A + +M+
Sbjct: 691 SAIAKMNKFDLVISFGEKMEILGISHNLYTYNILINCFCR---CSRLSLALALLGKMMKL 747
Query: 272 KICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEA 331
P+ VT N+L++GFC ++ A ++M + G KP+ VT+ +LI+GL + K EA
Sbjct: 748 GYEPDIVTLNSLLNGFCHGNRISDAVALVDQMVEMGYKPDTVTFTTLIHGLFLHNKASEA 807
Query: 332 IDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDA 391
+ L D+MV G +P++VTY A++NG CK+ A + + + ++ NV+ ++T+ID+
Sbjct: 808 VALIDRMVQRGCQPDLVTYGAVVNGLCKRGDTDLALNLLNKMEAAKIEANVVIYSTVIDS 867
Query: 392 YCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGD 451
CK ++ +L + M ++G+ PNV TY+ LI+ LC A LL++M + + +
Sbjct: 868 LCKYRHEDDALNLFTEMENKGVRPNVITYSSLISCLCNYGRWSDASRLLSDMIERKINPN 927
Query: 452 VVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRT 511
+VT++ LID K K AEKL EM + PN TY++L++G+CM +L A +
Sbjct: 928 LVTFSALIDAFVKKGKLVKAEKLYEEMIKRSIDPNIFTYSSLINGFCMLDRLGEAKQMLE 987
Query: 512 RMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTY 560
M ++ PNVVTYN LI G+CK +++ L EM ++GL N TY
Sbjct: 988 LMIRKDCLPNVVTYNTLINGFCKAKRVDKGMELFREMSQRGLVGNTVTY 1036
Score = 222 bits (566), Expect = 4e-58
Identities = 138/439 (31%), Positives = 222/439 (50%), Gaps = 38/439 (8%)
Query: 168 SLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAI 227
S+ + LLSA+ K NK V ++M I NL T+NI IN CR +L+ A +
Sbjct: 682 SIIEFSKLLSAIAKMNKFDLVISFGEKMEILGISHNLYTYNILINCFCRCSRLSLALALL 741
Query: 228 EDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGF 287
M G P++VT N+L++G+C ++ A A + +M+ P+ VTF TLI G
Sbjct: 742 GKMMKLGYEPDIVTLNSLLNGFCH---GNRISDAVALVDQMVEMGYKPDTVTFTTLIHGL 798
Query: 288 CKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNG---------------KLE--- 329
+ A + M ++G +P++VTY +++NGLC G K+E
Sbjct: 799 FLHNKASEAVALIDRMVQRGCQPDLVTYGAVVNGLCKRGDTDLALNLLNKMEAAKIEANV 858
Query: 330 -----------------EAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDD 372
+A++L+ +M G++PN++TY++LI+ C +A+++ D
Sbjct: 859 VIYSTVIDSLCKYRHEDDALNLFTEMENKGVRPNVITYSSLISCLCNYGRWSDASRLLSD 918
Query: 373 VSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQD 432
+ ++++ PN++TF+ +IDA+ K+G + + L M+ I PN+ TY+ LI G C
Sbjct: 919 MIERKINPNLVTFSALIDAFVKKGKLVKAEKLYEEMIKRSIDPNIFTYSSLINGFCMLDR 978
Query: 433 LQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNT 492
L AK++L M K +VVTYN LI+G CK + +L EM GL N VTY T
Sbjct: 979 LGEAKQMLELMIRKDCLPNVVTYNTLINGFCKAKRVDKGMELFREMSQRGLVGNTVTYTT 1038
Query: 493 LMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKG 552
L+ G+ A V +M PN++TYN+L+ G CK KL A + +
Sbjct: 1039 LIHGFFQARDCDNAQMVFKQMVSVGVHPNILTYNILLDGLCKNGKLAKAMVVFEYLQRST 1098
Query: 553 LNPNRTTYDIVRLEMLEKG 571
+ P+ TY+I+ M + G
Sbjct: 1099 MEPDIYTYNIMIEGMCKAG 1117
Score = 196 bits (497), Expect = 4e-50
Identities = 116/364 (31%), Positives = 190/364 (51%), Gaps = 8/364 (2%)
Query: 209 IFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEM 268
I N L K++ A D DM P++V +N L+ K K + ++M
Sbjct: 53 ILRNRLSDIIKVDDAVDLFGDMVKSRPFPSIVEFNKLLSAVAKMN---KFELVISLGEQM 109
Query: 269 LANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKL 328
I + T++ I+ FC+ ++ A +M K G +P+IVT +SL+NG C++ ++
Sbjct: 110 QTLGISHDLYTYSIFINCFCRRSQLSLALAVLAKMMKLGYEPDIVTLSSLLNGYCHSKRI 169
Query: 329 EEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTM 388
+A+ L D+MV +G KP+ T+ LI+G EA + D + ++ P+++T+ T+
Sbjct: 170 SDAVALVDQMVEMGYKPDTFTFTTLIHGLFLHNKASEAVALVDQMVQRGCQPDLVTYGTV 229
Query: 389 IDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGL 448
++ CK G ++ SL M I +V YN +I GLC+ + + A L EM+NKG+
Sbjct: 230 VNGLCKRGDIDLALSLLKKMEKGKIEADVVIYNTIIDGLCKYKHMDDALNLFTEMDNKGI 289
Query: 449 KGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALN 508
+ DV TY+ LI LC + +A +LL++M + PN VT++ L+D + EGKL A
Sbjct: 290 RPDVFTYSSLISCLCNYGRWSDASRLLSDMIERKINPNVVTFSALIDAFVKEGKLVEAEK 349
Query: 509 VRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEML 568
+ M K P++ TY+ LI G+C ++L+ A + M+ K PN TY L
Sbjct: 350 LYDEMIKRSIDPDIFTYSSLINGFCMHDRLDEAKHMFELMISKDCFPNVVTY-----STL 404
Query: 569 EKGF 572
KGF
Sbjct: 405 IKGF 408
Score = 193 bits (491), Expect = 2e-49
Identities = 106/325 (32%), Positives = 178/325 (54%), Gaps = 3/325 (0%)
Query: 237 PNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAA 296
P+++ ++ L+ K K +F ++M I N T+N LI+ FC+ ++ A
Sbjct: 681 PSIIEFSKLLSAIAKMN---KFDLVISFGEKMEILGISHNLYTYNILINCFCRCSRLSLA 737
Query: 297 KKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALING 356
+M K G +P+IVT NSL+NG C+ ++ +A+ L D+MV +G KP+ VT+ LI+G
Sbjct: 738 LALLGKMMKLGYEPDIVTLNSLLNGFCHGNRISDAVALVDQMVEMGYKPDTVTFTTLIHG 797
Query: 357 FCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPN 416
EA + D + ++ P+++T+ +++ CK G + +L + M I N
Sbjct: 798 LFLHNKASEAVALIDRMVQRGCQPDLVTYGAVVNGLCKRGDTDLALNLLNKMEAAKIEAN 857
Query: 417 VSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLN 476
V Y+ +I LC+ + A L EMENKG++ +V+TY+ LI LC + +A +LL+
Sbjct: 858 VVIYSTVIDSLCKYRHEDDALNLFTEMENKGVRPNVITYSSLISCLCNYGRWSDASRLLS 917
Query: 477 EMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKIN 536
+M + PN VT++ L+D + +GKL A + M K PN+ TY+ LI G+C ++
Sbjct: 918 DMIERKINPNLVTFSALIDAFVKKGKLVKAEKLYEEMIKRSIDPNIFTYSSLINGFCMLD 977
Query: 537 KLEAANGLLNEMLEKGLNPNRTTYD 561
+L A +L M+ K PN TY+
Sbjct: 978 RLGEAKQMLELMIRKDCLPNVVTYN 1002
Score = 185 bits (470), Expect = 5e-47
Identities = 124/490 (25%), Positives = 224/490 (45%), Gaps = 39/490 (7%)
Query: 119 VFHSLLLDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSA 178
VF ++ G P L ++L+ KN +L A F + + + + N ++
Sbjct: 455 VFKQMVSVGVHPNI--LTYNILLDGLCKNGKLAKAMVVFEYLQRSTMEPDIYTYNIMIEG 512
Query: 179 LVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPN 238
+ K K+ D ++ + + + N+ +N I+G CR G +A+ ++ MK G PN
Sbjct: 513 MCKAGKVEDGWELFCNLSLKGVSPNVIAYNTMISGFCRKGSKEEADSLLKKMKEDGPLPN 572
Query: 239 VVTYNTLVDGYCKRGSA------------------------GKMYKAEA----------- 263
TYNTL+ + G +K E
Sbjct: 573 SGTYNTLIRARLRDGDREASAELIKEMRSCGFAGDASTIGLSSKFKREIEKPNRENACKN 632
Query: 264 --FMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLING 321
F +E+ +++ C + ++ D + A F M + P+I+ ++ L++
Sbjct: 633 IIFSEEICSSESCGESYDYREVLRTGLSDIELDDAIGLFGVMAQSRPFPSIIEFSKLLSA 692
Query: 322 LCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPN 381
+ K + I +KM LG+ N+ TYN LIN FC+ + A + + K P+
Sbjct: 693 IAKMNKFDLVISFGEKMEILGISHNLYTYNILINCFCRCSRLSLALALLGKMMKLGYEPD 752
Query: 382 VITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLN 441
++T N++++ +C + + +L M++ G P+ T+ LI GL A L++
Sbjct: 753 IVTLNSLLNGFCHGNRISDAVALVDQMVEMGYKPDTVTFTTLIHGLFLHNKASEAVALID 812
Query: 442 EMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEG 501
M +G + D+VTY +++GLCK + A LLN+M ++ N V Y+T++D C
Sbjct: 813 RMVQRGCQPDLVTYGAVVNGLCKRGDTDLALNLLNKMEAAKIEANVVIYSTVIDSLCKYR 872
Query: 502 KLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYD 561
ALN+ T ME + +PNV+TY+ LI C + A+ LL++M+E+ +NPN T+
Sbjct: 873 HEDDALNLFTEMENKGVRPNVITYSSLISCLCNYGRWSDASRLLSDMIERKINPNLVTFS 932
Query: 562 IVRLEMLEKG 571
+ ++KG
Sbjct: 933 ALIDAFVKKG 942
Score = 168 bits (425), Expect = 9e-42
Identities = 92/278 (33%), Positives = 155/278 (55%), Gaps = 5/278 (1%)
Query: 125 LDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENK 184
++ + A +I ++ + K A FT ++ G + ++ + + L+S L +
Sbjct: 849 MEAAKIEANVVIYSTVIDSLCKYRHEDDALNLFTEMENKGVRPNVITYSSLISCLCNYGR 908
Query: 185 IGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNT 244
D + +MI+R+I+ NL TF+ I+ + GKL KAE E+M I PN+ TY++
Sbjct: 909 WSDASRLLSDMIERKINPNLVTFSALIDAFVKKGKLVKAEKLYEEMIKRSIDPNIFTYSS 968
Query: 245 LVDGYCKRGSAGKMYKAEAFMKEMLANKIC-PNEVTFNTLIDGFCKDENVAAAKKAFEEM 303
L++G+C G+ + M E++ K C PN VT+NTLI+GFCK + V + F EM
Sbjct: 969 LINGFCMLDRLGEAKQ----MLELMIRKDCLPNVVTYNTLINGFCKAKRVDKGMELFREM 1024
Query: 304 QKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMM 363
++GL N VTY +LI+G + A ++ +MV +G+ PNI+TYN L++G CK +
Sbjct: 1025 SQRGLVGNTVTYTTLIHGFFQARDCDNAQMVFKQMVSVGVHPNILTYNILLDGLCKNGKL 1084
Query: 364 KEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEG 401
+A VF+ + + + P++ T+N MI+ CK G + G
Sbjct: 1085 AKAMVVFEYLQRSTMEPDIYTYNIMIEGMCKAGKWKMG 1122
>At1g62590 putative membrane-associated salt-inducible protein
(AL021637); similar to EST gb|AA728420
Length = 634
Score = 266 bits (681), Expect = 2e-71
Identities = 140/424 (33%), Positives = 238/424 (56%), Gaps = 3/424 (0%)
Query: 138 DMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIK 197
++L+ + + ++ A + G++ S+ + + LL+ +I D + +M++
Sbjct: 124 NILINCFCRRSQISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVE 183
Query: 198 RRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGK 257
+ TF I+GL K ++A ++ M G PN+VTY +V+G CKRG
Sbjct: 184 MGYRPDTITFTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDTDL 243
Query: 258 MYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNS 317
A + +M A KI + V FNT+ID CK +V A F+EM+ +G++PN+VTY+S
Sbjct: 244 ---ALNLLNKMEAAKIEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKGIRPNVVTYSS 300
Query: 318 LINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQE 377
LI+ LC+ G+ +A L M+ + PN+VT+NALI+ F K+ EA K++DD+ K+
Sbjct: 301 LISCLCSYGRWSDASQLLSDMIEKKINPNLVTFNALIDAFVKEGKFVEAEKLYDDMIKRS 360
Query: 378 LVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAK 437
+ P++ T+N++++ +C +++ + M+ + P+V TYN LI G C+ + ++
Sbjct: 361 IDPDIFTYNSLVNGFCMHDRLDKAKQMFEFMVSKDCFPDVVTYNTLIKGFCKSKRVEDGT 420
Query: 438 ELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGY 497
EL EM ++GL GD VTY LI GL + NA+K+ +M + G+ P+ +TY+ L+DG
Sbjct: 421 ELFREMSHRGLVGDTVTYTTLIQGLFHDGDCDNAQKVFKQMVSDGVPPDIMTYSILLDGL 480
Query: 498 CMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNR 557
C GKL+ AL V M+K + ++ Y +I+G CK K++ L + KG+ PN
Sbjct: 481 CNNGKLEKALEVFDYMQKSEIKLDIYIYTTMIEGMCKAGKVDDGWDLFCSLSLKGVKPNV 540
Query: 558 TTYD 561
TY+
Sbjct: 541 VTYN 544
Score = 243 bits (621), Expect = 2e-64
Identities = 144/479 (30%), Positives = 250/479 (52%), Gaps = 32/479 (6%)
Query: 78 LSYGLEPT----SKVLHFLANSKRYSKVRSFLDSFV----KNEKHTVSSVFHSLLLDGGR 129
+ G EP+ S +L+ + KR S + +D V + + T +++ H L L
Sbjct: 147 MKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTITFTTLIHGLFLHNKA 206
Query: 130 PGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVE 189
A AL+ M+ G + +L + +++ L K
Sbjct: 207 SEAVALVDRMV---------------------QRGCQPNLVTYGVVVNGLCKRGDTDLAL 245
Query: 190 YVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGY 249
+ +M +I ++ FN I+ LC+ ++ A + ++M+ GI PNVVTY++L+
Sbjct: 246 NLLNKMEAAKIEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKGIRPNVVTYSSLISCL 305
Query: 250 CKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLK 309
C S G+ A + +M+ KI PN VTFN LID F K+ A+K +++M K+ +
Sbjct: 306 C---SYGRWSDASQLLSDMIEKKINPNLVTFNALIDAFVKEGKFVEAEKLYDDMIKRSID 362
Query: 310 PNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKV 369
P+I TYNSL+NG C + +L++A +++ MV P++VTYN LI GFCK K +++ T++
Sbjct: 363 PDIFTYNSLVNGFCMHDRLDKAKQMFEFMVSKDCFPDVVTYNTLIKGFCKSKRVEDGTEL 422
Query: 370 FDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCR 429
F ++S + LV + +T+ T+I +G + + M+ +G+ P++ TY+ L+ GLC
Sbjct: 423 FREMSHRGLVGDTVTYTTLIQGLFHDGDCDNAQKVFKQMVSDGVPPDIMTYSILLDGLCN 482
Query: 430 KQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVT 489
L+ A E+ + M+ +K D+ Y +I+G+CK K + L + G+KPN VT
Sbjct: 483 NGKLEKALEVFDYMQKSEIKLDIYIYTTMIEGMCKAGKVDDGWDLFCSLSLKGVKPNVVT 542
Query: 490 YNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEM 548
YNT++ G C + L+ A + +M+++ PN TYN LI+ + + A+ L+ EM
Sbjct: 543 YNTMISGLCSKRLLQEAYALLKKMKEDGPLPNSGTYNTLIRAHLRDGDKAASAELIREM 601
Score = 222 bits (565), Expect = 5e-58
Identities = 136/446 (30%), Positives = 231/446 (51%), Gaps = 19/446 (4%)
Query: 115 TVSSVFHSLLLDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNP 174
T S + H L G P ID+ + Y + RA F
Sbjct: 12 TTSRIVHRNLQGKGNPRIAPSSIDLCGMCY------------WGRA----FSSGSGDYRE 55
Query: 175 LLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWG 234
+L + + K+ D ++ M+K R ++ FN ++ + + K + E M+
Sbjct: 56 ILRNGLHDMKLDDAIGLFGGMVKSRPLPSIVEFNKLLSAIAKMKKFDVVISLGEKMQRLE 115
Query: 235 ISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVA 294
I + TYN L++ +C+R ++ A A + +M+ P+ VT ++L++G+C + ++
Sbjct: 116 IVHGLYTYNILINCFCRR---SQISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRIS 172
Query: 295 AAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALI 354
A ++M + G +P+ +T+ +LI+GL + K EA+ L D+MV G +PN+VTY ++
Sbjct: 173 DAVALVDQMVEMGYRPDTITFTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVV 232
Query: 355 NGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGIL 414
NG CK+ A + + + ++ +V+ FNT+ID+ CK +++ +L M +GI
Sbjct: 233 NGLCKRGDTDLALNLLNKMEAAKIEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKGIR 292
Query: 415 PNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKL 474
PNV TY+ LI+ LC A +LL++M K + ++VT+N LID K K AEKL
Sbjct: 293 PNVVTYSSLISCLCSYGRWSDASQLLSDMIEKKINPNLVTFNALIDAFVKEGKFVEAEKL 352
Query: 475 LNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCK 534
++M + P+ TYN+L++G+CM +L A + M + P+VVTYN LIKG+CK
Sbjct: 353 YDDMIKRSIDPDIFTYNSLVNGFCMHDRLDKAKQMFEFMVSKDCFPDVVTYNTLIKGFCK 412
Query: 535 INKLEAANGLLNEMLEKGLNPNRTTY 560
++E L EM +GL + TY
Sbjct: 413 SKRVEDGTELFREMSHRGLVGDTVTY 438
Score = 206 bits (524), Expect = 3e-53
Identities = 114/305 (37%), Positives = 177/305 (57%), Gaps = 9/305 (2%)
Query: 168 SLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAI 227
+L + N L+ A VKE K + E +Y +MIKR I ++ T+N +NG C +L+KA+
Sbjct: 329 NLVTFNALIDAFVKEGKFVEAEKLYDDMIKRSIDPDIFTYNSLVNGFCMHDRLDKAKQMF 388
Query: 228 EDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGF 287
E M + P+VVTYNTL+ G+CK + ++ +EM + + VT+ TLI G
Sbjct: 389 EFMVSKDCFPDVVTYNTLIKGFCK---SKRVEDGTELFREMSHRGLVGDTVTYTTLIQGL 445
Query: 288 CKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNI 347
D + A+K F++M G+ P+I+TY+ L++GLCNNGKLE+A++++D M +K +I
Sbjct: 446 FHDGDCDNAQKVFKQMVSDGVPPDIMTYSILLDGLCNNGKLEKALEVFDYMQKSEIKLDI 505
Query: 348 VTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSS 407
Y +I G CK + + +F +S + + PNV+T+NTMI C + +++E ++L
Sbjct: 506 YIYTTMIEGMCKAGKVDDGWDLFCSLSLKGVKPNVVTYNTMISGLCSKRLLQEAYALLKK 565
Query: 408 MLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTY----NILIDGLC 463
M ++G LPN TYN LI R D A+ EL+ EM + GD T N+L DG
Sbjct: 566 MKEDGPLPNSGTYNTLIRAHLRDGDKAASAELIREMRSCRFVGDASTIGLVANMLHDG-- 623
Query: 464 KNDKS 468
+ DKS
Sbjct: 624 RLDKS 628
Score = 185 bits (470), Expect = 5e-47
Identities = 104/336 (30%), Positives = 177/336 (51%), Gaps = 8/336 (2%)
Query: 237 PNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAA 296
P++V +N L+ K + M+ + +I T+N LI+ FC+ ++ A
Sbjct: 83 PSIVEFNKLLSAIAKMKKFDVVISLGEKMQRL---EIVHGLYTYNILINCFCRRSQISLA 139
Query: 297 KKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALING 356
+M K G +P+IVT +SL+NG C+ ++ +A+ L D+MV +G +P+ +T+ LI+G
Sbjct: 140 LALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTITFTTLIHG 199
Query: 357 FCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPN 416
EA + D + ++ PN++T+ +++ CK G + +L + M I +
Sbjct: 200 LFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDTDLALNLLNKMEAAKIEAD 259
Query: 417 VSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLN 476
V +N +I LC+ + + A L EME KG++ +VVTY+ LI LC + +A +LL+
Sbjct: 260 VVIFNTIIDSLCKYRHVDDALNLFKEMETKGIRPNVVTYSSLISCLCSYGRWSDASQLLS 319
Query: 477 EMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKIN 536
+M + PN VT+N L+D + EGK A + M K P++ TYN L+ G+C +
Sbjct: 320 DMIEKKINPNLVTFNALIDAFVKEGKFVEAEKLYDDMIKRSIDPDIFTYNSLVNGFCMHD 379
Query: 537 KLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKGF 572
+L+ A + M+ K P D+V L KGF
Sbjct: 380 RLDKAKQMFEFMVSKDCFP-----DVVTYNTLIKGF 410
Score = 181 bits (458), Expect = 1e-45
Identities = 97/294 (32%), Positives = 160/294 (53%)
Query: 268 MLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGK 327
M+ ++ P+ V FN L+ K + E+MQ+ + + TYN LIN C +
Sbjct: 76 MVKSRPLPSIVEFNKLLSAIAKMKKFDVVISLGEKMQRLEIVHGLYTYNILINCFCRRSQ 135
Query: 328 LEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNT 387
+ A+ L KM+ LG +P+IVT ++L+NG+C K + +A + D + + P+ ITF T
Sbjct: 136 ISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTITFTT 195
Query: 388 MIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKG 447
+I E +L M+ G PN+ TY ++ GLC++ D A LLN+ME
Sbjct: 196 LIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDTDLALNLLNKMEAAK 255
Query: 448 LKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAAL 507
++ DVV +N +ID LCK +A L EM G++PN VTY++L+ C G+ A
Sbjct: 256 IEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKGIRPNVVTYSSLISCLCSYGRWSDAS 315
Query: 508 NVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYD 561
+ + M +++ PN+VT+N LI + K K A L ++M+++ ++P+ TY+
Sbjct: 316 QLLSDMIEKKINPNLVTFNALIDAFVKEGKFVEAEKLYDDMIKRSIDPDIFTYN 369
Score = 152 bits (384), Expect = 5e-37
Identities = 83/272 (30%), Positives = 147/272 (53%)
Query: 300 FEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCK 359
F M K P+IV +N L++ + K + I L +KM L + + TYN LIN FC+
Sbjct: 73 FGGMVKSRPLPSIVEFNKLLSAIAKMKKFDVVISLGEKMQRLEIVHGLYTYNILINCFCR 132
Query: 360 KKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVST 419
+ + A + + K P+++T +++++ YC + + +L M++ G P+ T
Sbjct: 133 RSQISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTIT 192
Query: 420 YNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMF 479
+ LI GL A L++ M +G + ++VTY ++++GLCK + A LLN+M
Sbjct: 193 FTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDTDLALNLLNKME 252
Query: 480 NLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLE 539
++ + V +NT++D C + ALN+ ME + +PNVVTY+ LI C +
Sbjct: 253 AAKIEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKGIRPNVVTYSSLISCLCSYGRWS 312
Query: 540 AANGLLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
A+ LL++M+EK +NPN T++ + +++G
Sbjct: 313 DASQLLSDMIEKKINPNLVTFNALIDAFVKEG 344
Score = 88.6 bits (218), Expect = 9e-18
Identities = 47/174 (27%), Positives = 92/174 (52%)
Query: 398 MEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNI 457
+++ L M+ LP++ +N L++ + + + L +M+ + + TYNI
Sbjct: 66 LDDAIGLFGGMVKSRPLPSIVEFNKLLSAIAKMKKFDVVISLGEKMQRLEIVHGLYTYNI 125
Query: 458 LIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKER 517
LI+ C+ + A LL +M LG +P+ VT ++L++GYC ++ A+ + +M +
Sbjct: 126 LINCFCRRSQISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMG 185
Query: 518 KQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
+P+ +T+ LI G NK A L++ M+++G PN TY +V + ++G
Sbjct: 186 YRPDTITFTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRG 239
Score = 29.3 bits (64), Expect = 6.4
Identities = 18/74 (24%), Positives = 33/74 (44%)
Query: 490 YNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEML 549
Y ++ + KL A+ + M K R P++V +N L+ K+ K + L +M
Sbjct: 53 YREILRNGLHDMKLDDAIGLFGGMVKSRPLPSIVEFNKLLSAIAKMKKFDVVISLGEKMQ 112
Query: 550 EKGLNPNRTTYDIV 563
+ TY+I+
Sbjct: 113 RLEIVHGLYTYNIL 126
>At1g74580 hypothetical protein
Length = 763
Score = 266 bits (679), Expect = 3e-71
Identities = 144/418 (34%), Positives = 235/418 (55%), Gaps = 4/418 (0%)
Query: 154 YEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFING 213
YE F + G L L++ N LL L K+ + + E + ++IKR + NL T+N+FI G
Sbjct: 201 YELFGKMLASGVSLCLSTFNKLLRVLCKKGDVKECEKLLDKVIKRGVLPNLFTYNLFIQG 260
Query: 214 LCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKI 273
LC+ G+L+ A + + G P+V+TYN L+ G CK K +AE ++ +M+ +
Sbjct: 261 LCQRGELDGAVRMVGCLIEQGPKPDVITYNNLIYGLCKNS---KFQEAEVYLGKMVNEGL 317
Query: 274 CPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAID 333
P+ T+NTLI G+CK V A++ + G P+ TY SLI+GLC+ G+ A+
Sbjct: 318 EPDSYTYNTLIAGYCKGGMVQLAERIVGDAVFNGFVPDQFTYRSLIDGLCHEGETNRALA 377
Query: 334 LWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYC 393
L+++ +G G+KPN++ YN LI G + M+ EA ++ +++S++ L+P V TFN +++ C
Sbjct: 378 LFNEALGKGIKPNVILYNTLIKGLSNQGMILEAAQLANEMSEKGLIPEVQTFNILVNGLC 437
Query: 394 KEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVV 453
K G + + L M+ +G P++ T+N LI G + ++ A E+L+ M + G+ DV
Sbjct: 438 KMGCVSDADGLVKVMISKGYFPDIFTFNILIHGYSTQLKMENALEILDVMLDNGVDPDVY 497
Query: 454 TYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRM 513
TYN L++GLCK K + + M G PN T+N L++ C KL AL + M
Sbjct: 498 TYNSLLNGLCKTSKFEDVMETYKTMVEKGCAPNLFTFNILLESLCRYRKLDEALGLLEEM 557
Query: 514 EKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEK-GLNPNRTTYDIVRLEMLEK 570
+ + P+ VT+ LI G+CK L+ A L +M E ++ + TY+I+ EK
Sbjct: 558 KNKSVNPDAVTFGTLIDGFCKNGDLDGAYTLFRKMEEAYKVSSSTPTYNIIIHAFTEK 615
Score = 235 bits (600), Expect = 4e-62
Identities = 158/539 (29%), Positives = 246/539 (45%), Gaps = 56/539 (10%)
Query: 61 DSELVLRFFKWSQKEYRLSYGLEPTSKVLHFLANSKRYSKVRSFLDSFVKNE-KHTVSSV 119
D L F +KE + L V+ L ++ + L +N H + V
Sbjct: 19 DPMKALEMFNSMRKEVGFKHTLSTYRSVIEKLGYYGKFEAMEEVLVDMRENVGNHMLEGV 78
Query: 120 FHSLLLDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSAL 179
+ + + GR G ++ A F R Y + ++ S N ++S L
Sbjct: 79 YVGAMKNYGRKG-----------------KVQEAVNVFERMDFYDCEPTVFSYNAIMSVL 121
Query: 180 VKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNV 239
V VY M R I ++ +F I + C+ + + A + +M + G NV
Sbjct: 122 VDSGYFDQAHKVYMRMRDRGITPDVYSFTIRMKSFCKTSRPHAALRLLNNMSSQGCEMNV 181
Query: 240 VTYNTLVDGY-----------------------------------CKRGSAGKMYKAEAF 264
V Y T+V G+ CK+G + + E
Sbjct: 182 VAYCTVVGGFYEENFKAEGYELFGKMLASGVSLCLSTFNKLLRVLCKKGD---VKECEKL 238
Query: 265 MKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCN 324
+ +++ + PN T+N I G C+ + A + + +QG KP+++TYN+LI GLC
Sbjct: 239 LDKVIKRGVLPNLFTYNLFIQGLCQRGELDGAVRMVGCLIEQGPKPDVITYNNLIYGLCK 298
Query: 325 NGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVIT 384
N K +EA KMV GL+P+ TYN LI G+CK M++ A ++ D VP+ T
Sbjct: 299 NSKFQEAEVYLGKMVNEGLEPDSYTYNTLIAGYCKGGMVQLAERIVGDAVFNGFVPDQFT 358
Query: 385 FNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEME 444
+ ++ID C EG +L + L +GI PNV YN LI GL + + A +L NEM
Sbjct: 359 YRSLIDGLCHEGETNRALALFNEALGKGIKPNVILYNTLIKGLSNQGMILEAAQLANEMS 418
Query: 445 NKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLK 504
KGL +V T+NIL++GLCK +A+ L+ M + G P+ T+N L+ GY + K++
Sbjct: 419 EKGLIPEVQTFNILVNGLCKMGCVSDADGLVKVMISKGYFPDIFTFNILIHGYSTQLKME 478
Query: 505 AALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIV 563
AL + M P+V TYN L+ G CK +K E M+EKG PN T++I+
Sbjct: 479 NALEILDVMLDNGVDPDVYTYNSLLNGLCKTSKFEDVMETYKTMVEKGCAPNLFTFNIL 537
Score = 209 bits (532), Expect = 3e-54
Identities = 128/421 (30%), Positives = 209/421 (49%), Gaps = 8/421 (1%)
Query: 161 KDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKL 220
K+ GFK +L++ ++ L K +E V +M R N +++ + G+
Sbjct: 32 KEVGFKHTLSTYRSVIEKLGYYGKFEAMEEVLVDM--RENVGNHMLEGVYVGAMKNYGRK 89
Query: 221 NKAEDAI---EDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNE 277
K ++A+ E M + P V +YN ++ G + +K M++ I P+
Sbjct: 90 GKVQEAVNVFERMDFYDCEPTVFSYNAIMSVLVDSGYFDQAHKVYMRMRD---RGITPDV 146
Query: 278 VTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDK 337
+F + FCK AA + M QG + N+V Y +++ G E +L+ K
Sbjct: 147 YSFTIRMKSFCKTSRPHAALRLLNNMSSQGCEMNVVAYCTVVGGFYEENFKAEGYELFGK 206
Query: 338 MVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGM 397
M+ G+ + T+N L+ CKK +KE K+ D V K+ ++PN+ T+N I C+ G
Sbjct: 207 MLASGVSLCLSTFNKLLRVLCKKGDVKECEKLLDKVIKRGVLPNLFTYNLFIQGLCQRGE 266
Query: 398 MEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNI 457
++ + ++++G P+V TYN LI GLC+ Q A+ L +M N+GL+ D TYN
Sbjct: 267 LDGAVRMVGCLIEQGPKPDVITYNNLIYGLCKNSKFQEAEVYLGKMVNEGLEPDSYTYNT 326
Query: 458 LIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKER 517
LI G CK + AE+++ + G P+ TY +L+DG C EG+ AL + +
Sbjct: 327 LIAGYCKGGMVQLAERIVGDAVFNGFVPDQFTYRSLIDGLCHEGETNRALALFNEALGKG 386
Query: 518 KQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKGFSPDIE 577
+PNV+ YN LIKG + A L NEM EKGL P T++I+ + + G D +
Sbjct: 387 IKPNVILYNTLIKGLSNQGMILEAAQLANEMSEKGLIPEVQTFNILVNGLCKMGCVSDAD 446
Query: 578 G 578
G
Sbjct: 447 G 447
Score = 159 bits (401), Expect = 5e-39
Identities = 106/385 (27%), Positives = 181/385 (46%), Gaps = 46/385 (11%)
Query: 149 ELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFN 208
E + A F A G K ++ N L+ L + I + + EM ++ + + TFN
Sbjct: 371 ETNRALALFNEALGKGIKPNVILYNTLIKGLSNQGMILEAAQLANEMSEKGLIPEVQTFN 430
Query: 209 IFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEM 268
I +NGLC+ G ++ A+ ++ M + G P++ T+N L+ GY + KM A + M
Sbjct: 431 ILVNGLCKMGCVSDADGLVKVMISKGYFPDIFTFNILIHGY---STQLKMENALEILDVM 487
Query: 269 LANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKL 328
L N + P+ T+N+L++G CK + ++ M ++G PN+ T+N L+ LC KL
Sbjct: 488 LDNGVDPDVYTYNSLLNGLCKTSKFEDVMETYKTMVEKGCAPNLFTFNILLESLCRYRKL 547
Query: 329 EEAIDLWDKMVGLGLKPNIVTYNALINGFCKK----------KMMKEATKV--------- 369
+EA+ L ++M + P+ VT+ LI+GFCK + M+EA KV
Sbjct: 548 DEALGLLEEMKNKSVNPDAVTFGTLIDGFCKNGDLDGAYTLFRKMEEAYKVSSSTPTYNI 607
Query: 370 -----------------FDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEG 412
F ++ + L P+ T+ M+D +CK G + G+ M++ G
Sbjct: 608 IIHAFTEKLNVTMAEKLFQEMVDRCLGPDGYTYRLMVDGFCKTGNVNLGYKFLLEMMENG 667
Query: 413 ILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSR-NA 471
+P+++T +I LC + + A +++ M KGL + V + +C DK A
Sbjct: 668 FIPSLTTLGRVINCLCVEDRVYEAAGIIHRMVQKGLVPEAV------NTICDVDKKEVAA 721
Query: 472 EKLLNEMFNLGLKPNHVTYNTLMDG 496
KL+ E + Y L DG
Sbjct: 722 PKLVLEDLLKKSCITYYAYELLFDG 746
Score = 137 bits (346), Expect = 1e-32
Identities = 83/258 (32%), Positives = 133/258 (51%), Gaps = 4/258 (1%)
Query: 138 DMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIK 197
++L+ Y L++ A E D G + + N LL+ L K +K DV YK M++
Sbjct: 465 NILIHGYSTQLKMENALEILDVMLDNGVDPDVYTYNSLLNGLCKTSKFEDVMETYKTMVE 524
Query: 198 RRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGK 257
+ NL TFNI + LCR KL++A +E+MK ++P+ VT+ TL+DG+CK G
Sbjct: 525 KGCAPNLFTFNILLESLCRYRKLDEALGLLEEMKNKSVNPDAVTFGTLIDGFCKNGDLDG 584
Query: 258 MYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNS 317
Y M+E A K+ + T+N +I F + NV A+K F+EM + L P+ TY
Sbjct: 585 AYTLFRKMEE--AYKVSSSTPTYNIIIHAFTEKLNVTMAEKLFQEMVDRCLGPDGYTYRL 642
Query: 318 LINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQE 377
+++G C G + +M+ G P++ T +IN C + + EA + + ++
Sbjct: 643 MVDGFCKTGNVNLGYKFLLEMMENGFIPSLTTLGRVINCLCVEDRVYEAAGIIHRMVQKG 702
Query: 378 LVPNVITFNTMIDAYCKE 395
LVP + NT+ D KE
Sbjct: 703 LVPEAV--NTICDVDKKE 718
Score = 48.5 bits (114), Expect = 1e-05
Identities = 37/137 (27%), Positives = 67/137 (48%), Gaps = 3/137 (2%)
Query: 428 CRKQDLQAAKELLNEMENK-GLKGDVVTYNILIDGLCKNDKSRNAEKLLNEM-FNLGLKP 485
C+K ++A E+ N M + G K + TY +I+ L K E++L +M N+G
Sbjct: 16 CQKDPMKAL-EMFNSMRKEVGFKHTLSTYRSVIEKLGYYGKFEAMEEVLVDMRENVGNHM 74
Query: 486 NHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLL 545
Y M Y +GK++ A+NV RM+ +P V +YN ++ + A+ +
Sbjct: 75 LEGVYVGAMKNYGRKGKVQEAVNVFERMDFYDCEPTVFSYNAIMSVLVDSGYFDQAHKVY 134
Query: 546 NEMLEKGLNPNRTTYDI 562
M ++G+ P+ ++ I
Sbjct: 135 MRMRDRGITPDVYSFTI 151
>At1g63400 PPR-repeat protein
Length = 577
Score = 265 bits (678), Expect = 4e-71
Identities = 138/440 (31%), Positives = 241/440 (54%), Gaps = 3/440 (0%)
Query: 138 DMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIK 197
++L+ + + ++ A + G++ S+ + + LL+ +I D + +M++
Sbjct: 124 NILINCFCRRSQISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVE 183
Query: 198 RRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGK 257
+ TF I+GL K ++A ++ M G PN+VTY +V+G CKRG
Sbjct: 184 MGYRPDTITFTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDIDL 243
Query: 258 MYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNS 317
+ + +M A KI N V ++T+ID CK + A F EM+ +G++PN++TY+S
Sbjct: 244 AFN---LLNKMEAAKIEANVVIYSTVIDSLCKYRHEDDALNLFTEMENKGVRPNVITYSS 300
Query: 318 LINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQE 377
LI+ LCN + +A L M+ + PN+VT+NALI+ F K+ + EA K++D++ K+
Sbjct: 301 LISCLCNYERWSDASRLLSDMIERKINPNVVTFNALIDAFVKEGKLVEAEKLYDEMIKRS 360
Query: 378 LVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAK 437
+ P++ T++++I+ +C ++E + M+ + PNV TYN LI G C+ + +
Sbjct: 361 IDPDIFTYSSLINGFCMHDRLDEAKHMFELMISKDCFPNVVTYNTLINGFCKAKRIDEGV 420
Query: 438 ELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGY 497
EL EM +GL G+ VTY LI G + NA+ + +M + G+ PN +TYNTL+DG
Sbjct: 421 ELFREMSQRGLVGNTVTYTTLIHGFFQARDCDNAQMVFKQMVSDGVHPNIMTYNTLLDGL 480
Query: 498 CMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNR 557
C GKL+ A+ V +++ + +P + TYN++I+G CK K+E L + KG+ P+
Sbjct: 481 CKNGKLEKAMVVFEYLQRSKMEPTIYTYNIMIEGMCKAGKVEDGWDLFCSLSLKGVKPDV 540
Query: 558 TTYDIVRLEMLEKGFSPDIE 577
Y+ + KG + +
Sbjct: 541 IIYNTMISGFCRKGLKEEAD 560
Score = 243 bits (620), Expect = 2e-64
Identities = 129/423 (30%), Positives = 224/423 (52%), Gaps = 3/423 (0%)
Query: 137 IDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMI 196
+ L+ Y + A + + G++ + L+ L NK + + M+
Sbjct: 158 LSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTITFTTLIHGLFLHNKASEAVALVDRMV 217
Query: 197 KRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAG 256
+R NL T+ + +NGLC+ G ++ A + + M+A I NVV Y+T++D CK
Sbjct: 218 QRGCQPNLVTYGVVVNGLCKRGDIDLAFNLLNKMEAAKIEANVVIYSTVIDSLCKYRHED 277
Query: 257 KMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYN 316
A EM + PN +T+++LI C E + A + +M ++ + PN+VT+N
Sbjct: 278 D---ALNLFTEMENKGVRPNVITYSSLISCLCNYERWSDASRLLSDMIERKINPNVVTFN 334
Query: 317 SLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQ 376
+LI+ GKL EA L+D+M+ + P+I TY++LINGFC + EA +F+ + +
Sbjct: 335 ALIDAFVKEGKLVEAEKLYDEMIKRSIDPDIFTYSSLINGFCMHDRLDEAKHMFELMISK 394
Query: 377 ELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAA 436
+ PNV+T+NT+I+ +CK ++EG L M G++ N TY LI G + +D A
Sbjct: 395 DCFPNVVTYNTLINGFCKAKRIDEGVELFREMSQRGLVGNTVTYTTLIHGFFQARDCDNA 454
Query: 437 KELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDG 496
+ + +M + G+ +++TYN L+DGLCKN K A + + ++P TYN +++G
Sbjct: 455 QMVFKQMVSDGVHPNIMTYNTLLDGLCKNGKLEKAMVVFEYLQRSKMEPTIYTYNIMIEG 514
Query: 497 YCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPN 556
C GK++ ++ + + +P+V+ YN +I G+C+ E A+ L +M E G P+
Sbjct: 515 MCKAGKVEDGWDLFCSLSLKGVKPDVIIYNTMISGFCRKGLKEEADALFRKMREDGPLPD 574
Query: 557 RTT 559
T
Sbjct: 575 SGT 577
Score = 194 bits (492), Expect = 1e-49
Identities = 102/323 (31%), Positives = 182/323 (55%), Gaps = 3/323 (0%)
Query: 125 LDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENK 184
++ + A +I ++ + K A FT ++ G + ++ + + L+S L +
Sbjct: 251 MEAAKIEANVVIYSTVIDSLCKYRHEDDALNLFTEMENKGVRPNVITYSSLISCLCNYER 310
Query: 185 IGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNT 244
D + +MI+R+I+ N+ TFN I+ + GKL +AE ++M I P++ TY++
Sbjct: 311 WSDASRLLSDMIERKINPNVVTFNALIDAFVKEGKLVEAEKLYDEMIKRSIDPDIFTYSS 370
Query: 245 LVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQ 304
L++G+C ++ +A+ + M++ PN VT+NTLI+GFCK + + + F EM
Sbjct: 371 LINGFCMH---DRLDEAKHMFELMISKDCFPNVVTYNTLINGFCKAKRIDEGVELFREMS 427
Query: 305 KQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMK 364
++GL N VTY +LI+G + A ++ +MV G+ PNI+TYN L++G CK ++
Sbjct: 428 QRGLVGNTVTYTTLIHGFFQARDCDNAQMVFKQMVSDGVHPNIMTYNTLLDGLCKNGKLE 487
Query: 365 EATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLI 424
+A VF+ + + ++ P + T+N MI+ CK G +E+G+ L S+ +G+ P+V YN +I
Sbjct: 488 KAMVVFEYLQRSKMEPTIYTYNIMIEGMCKAGKVEDGWDLFCSLSLKGVKPDVIIYNTMI 547
Query: 425 AGLCRKQDLQAAKELLNEMENKG 447
+G CRK + A L +M G
Sbjct: 548 SGFCRKGLKEEADALFRKMREDG 570
Score = 193 bits (490), Expect = 3e-49
Identities = 104/325 (32%), Positives = 177/325 (54%), Gaps = 3/325 (0%)
Query: 237 PNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAA 296
P++ +N L+ K + M+ + I N T+N LI+ FC+ ++ A
Sbjct: 83 PSIFEFNKLLSAIAKMKKFDLVISLGEKMQRL---GISHNLYTYNILINCFCRRSQISLA 139
Query: 297 KKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALING 356
+M K G +P+IVT +SL+NG C+ ++ +A+ L D+MV +G +P+ +T+ LI+G
Sbjct: 140 LALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTITFTTLIHG 199
Query: 357 FCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPN 416
EA + D + ++ PN++T+ +++ CK G ++ F+L + M I N
Sbjct: 200 LFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDIDLAFNLLNKMEAAKIEAN 259
Query: 417 VSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLN 476
V Y+ +I LC+ + A L EMENKG++ +V+TY+ LI LC ++ +A +LL+
Sbjct: 260 VVIYSTVIDSLCKYRHEDDALNLFTEMENKGVRPNVITYSSLISCLCNYERWSDASRLLS 319
Query: 477 EMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKIN 536
+M + PN VT+N L+D + EGKL A + M K P++ TY+ LI G+C +
Sbjct: 320 DMIERKINPNVVTFNALIDAFVKEGKLVEAEKLYDEMIKRSIDPDIFTYSSLINGFCMHD 379
Query: 537 KLEAANGLLNEMLEKGLNPNRTTYD 561
+L+ A + M+ K PN TY+
Sbjct: 380 RLDEAKHMFELMISKDCFPNVVTYN 404
Score = 184 bits (468), Expect = 9e-47
Identities = 99/293 (33%), Positives = 161/293 (54%)
Query: 268 MLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGK 327
M+ ++ P+ FN L+ K + E+MQ+ G+ N+ TYN LIN C +
Sbjct: 76 MVKSRPLPSIFEFNKLLSAIAKMKKFDLVISLGEKMQRLGISHNLYTYNILINCFCRRSQ 135
Query: 328 LEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNT 387
+ A+ L KM+ LG +P+IVT ++L+NG+C K + +A + D + + P+ ITF T
Sbjct: 136 ISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTITFTT 195
Query: 388 MIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKG 447
+I E +L M+ G PN+ TY ++ GLC++ D+ A LLN+ME
Sbjct: 196 LIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDIDLAFNLLNKMEAAK 255
Query: 448 LKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAAL 507
++ +VV Y+ +ID LCK +A L EM N G++PN +TY++L+ C + A
Sbjct: 256 IEANVVIYSTVIDSLCKYRHEDDALNLFTEMENKGVRPNVITYSSLISCLCNYERWSDAS 315
Query: 508 NVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTY 560
+ + M + + PNVVT+N LI + K KL A L +EM+++ ++P+ TY
Sbjct: 316 RLLSDMIERKINPNVVTFNALIDAFVKEGKLVEAEKLYDEMIKRSIDPDIFTY 368
Score = 155 bits (393), Expect = 5e-38
Identities = 84/272 (30%), Positives = 147/272 (53%)
Query: 300 FEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCK 359
F M K P+I +N L++ + K + I L +KM LG+ N+ TYN LIN FC+
Sbjct: 73 FGGMVKSRPLPSIFEFNKLLSAIAKMKKFDLVISLGEKMQRLGISHNLYTYNILINCFCR 132
Query: 360 KKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVST 419
+ + A + + K P+++T +++++ YC + + +L M++ G P+ T
Sbjct: 133 RSQISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTIT 192
Query: 420 YNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMF 479
+ LI GL A L++ M +G + ++VTY ++++GLCK A LLN+M
Sbjct: 193 FTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDIDLAFNLLNKME 252
Query: 480 NLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLE 539
++ N V Y+T++D C ALN+ T ME + +PNV+TY+ LI C +
Sbjct: 253 AAKIEANVVIYSTVIDSLCKYRHEDDALNLFTEMENKGVRPNVITYSSLISCLCNYERWS 312
Query: 540 AANGLLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
A+ LL++M+E+ +NPN T++ + +++G
Sbjct: 313 DASRLLSDMIERKINPNVVTFNALIDAFVKEG 344
Score = 133 bits (335), Expect = 2e-31
Identities = 71/246 (28%), Positives = 126/246 (50%)
Query: 315 YNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVS 374
Y ++ ++ KL++AI L+ MV P+I +N L++ K K + + +
Sbjct: 53 YREILRNGLHSMKLDDAIGLFGGMVKSRPLPSIFEFNKLLSAIAKMKKFDLVISLGEKMQ 112
Query: 375 KQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQ 434
+ + N+ T+N +I+ +C+ + +L M+ G P++ T + L+ G C + +
Sbjct: 113 RLGISHNLYTYNILINCFCRRSQISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRIS 172
Query: 435 AAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLM 494
A L+++M G + D +T+ LI GL ++K+ A L++ M G +PN VTY ++
Sbjct: 173 DAVALVDQMVEMGYRPDTITFTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVV 232
Query: 495 DGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLN 554
+G C G + A N+ +ME + + NVV Y+ +I CK + A L EM KG+
Sbjct: 233 NGLCKRGDIDLAFNLLNKMEAAKIEANVVIYSTVIDSLCKYRHEDDALNLFTEMENKGVR 292
Query: 555 PNRTTY 560
PN TY
Sbjct: 293 PNVITY 298
Score = 112 bits (281), Expect = 4e-25
Identities = 63/214 (29%), Positives = 106/214 (49%)
Query: 350 YNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSML 409
Y ++ + +A +F + K +P++ FN ++ A K + SL M
Sbjct: 53 YREILRNGLHSMKLDDAIGLFGGMVKSRPLPSIFEFNKLLSAIAKMKKFDLVISLGEKMQ 112
Query: 410 DEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSR 469
GI N+ TYN LI CR+ + A LL +M G + +VT + L++G C +
Sbjct: 113 RLGISHNLYTYNILINCFCRRSQISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRIS 172
Query: 470 NAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLI 529
+A L+++M +G +P+ +T+ TL+ G + K A+ + RM + QPN+VTY V++
Sbjct: 173 DAVALVDQMVEMGYRPDTITFTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVV 232
Query: 530 KGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIV 563
G CK ++ A LLN+M + N Y V
Sbjct: 233 NGLCKRGDIDLAFNLLNKMEAAKIEANVVIYSTV 266
Score = 92.8 bits (229), Expect = 5e-19
Identities = 48/174 (27%), Positives = 94/174 (53%)
Query: 398 MEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNI 457
+++ L M+ LP++ +N L++ + + + L +M+ G+ ++ TYNI
Sbjct: 66 LDDAIGLFGGMVKSRPLPSIFEFNKLLSAIAKMKKFDLVISLGEKMQRLGISHNLYTYNI 125
Query: 458 LIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKER 517
LI+ C+ + A LL +M LG +P+ VT ++L++GYC ++ A+ + +M +
Sbjct: 126 LINCFCRRSQISLALALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMG 185
Query: 518 KQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
+P+ +T+ LI G NK A L++ M+++G PN TY +V + ++G
Sbjct: 186 YRPDTITFTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRG 239
>At1g63330 unknown protein
Length = 558
Score = 265 bits (678), Expect = 4e-71
Identities = 140/420 (33%), Positives = 235/420 (55%), Gaps = 3/420 (0%)
Query: 142 LAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIH 201
+A +K +L + + G++ S+ + + LL+ +I D + +M++
Sbjct: 52 IAKMKKFDLVISLALLGKMMKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMGYR 111
Query: 202 TNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKA 261
+ TF I+GL K ++A ++ M G PN+VTY +V+G CKRG +
Sbjct: 112 PDTITFTTLIHGLFLHNKASEAVALVDRMVQRGCQPNLVTYGVVVNGLCKRGDIDLAFN- 170
Query: 262 EAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLING 321
+ +M A KI + V FNT+ID CK +V A F+EM+ +G++PN+VTY+SLI+
Sbjct: 171 --LLNKMEAAKIEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKGIRPNVVTYSSLISC 228
Query: 322 LCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPN 381
LC+ G+ +A L M+ + PN+VT+NALI+ F K+ EA K+ DD+ K+ + P+
Sbjct: 229 LCSYGRWSDASQLLSDMIEKKINPNLVTFNALIDAFVKEGKFVEAEKLHDDMIKRSIDPD 288
Query: 382 VITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLN 441
+ T+N++I+ +C +++ + M+ + P++ TYN LI G C+ + ++ EL
Sbjct: 289 IFTYNSLINGFCMHDRLDKAKQMFEFMVSKDCFPDLDTYNTLIKGFCKSKRVEDGTELFR 348
Query: 442 EMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEG 501
EM ++GL GD VTY LI GL + NA+K+ +M + G+ P+ +TY+ L+DG C G
Sbjct: 349 EMSHRGLVGDTVTYTTLIQGLFHDGDCDNAQKVFKQMVSDGVPPDIMTYSILLDGLCNNG 408
Query: 502 KLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYD 561
KL+ AL V M+K + ++ Y +I+G CK K++ L + KG+ PN TY+
Sbjct: 409 KLEKALEVFDYMQKSEIKLDIYIYTTMIEGMCKAGKVDDGWDLFCSLSLKGVKPNVVTYN 468
Score = 239 bits (610), Expect = 3e-63
Identities = 144/479 (30%), Positives = 249/479 (51%), Gaps = 32/479 (6%)
Query: 78 LSYGLEPT----SKVLHFLANSKRYSKVRSFLDSFV----KNEKHTVSSVFHSLLLDGGR 129
+ G EP+ S +L+ + KR S + +D V + + T +++ H L L
Sbjct: 71 MKLGYEPSIVTLSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTITFTTLIHGLFLHNKA 130
Query: 130 PGATALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVE 189
A AL+ M+ G + +L + +++ L K I
Sbjct: 131 SEAVALVDRMV---------------------QRGCQPNLVTYGVVVNGLCKRGDIDLAF 169
Query: 190 YVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGY 249
+ +M +I ++ FN I+ LC+ ++ A + ++M+ GI PNVVTY++L+
Sbjct: 170 NLLNKMEAAKIEADVVIFNTIIDSLCKYRHVDDALNLFKEMETKGIRPNVVTYSSLISCL 229
Query: 250 CKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLK 309
C S G+ A + +M+ KI PN VTFN LID F K+ A+K ++M K+ +
Sbjct: 230 C---SYGRWSDASQLLSDMIEKKINPNLVTFNALIDAFVKEGKFVEAEKLHDDMIKRSID 286
Query: 310 PNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKV 369
P+I TYNSLING C + +L++A +++ MV P++ TYN LI GFCK K +++ T++
Sbjct: 287 PDIFTYNSLINGFCMHDRLDKAKQMFEFMVSKDCFPDLDTYNTLIKGFCKSKRVEDGTEL 346
Query: 370 FDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCR 429
F ++S + LV + +T+ T+I +G + + M+ +G+ P++ TY+ L+ GLC
Sbjct: 347 FREMSHRGLVGDTVTYTTLIQGLFHDGDCDNAQKVFKQMVSDGVPPDIMTYSILLDGLCN 406
Query: 430 KQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVT 489
L+ A E+ + M+ +K D+ Y +I+G+CK K + L + G+KPN VT
Sbjct: 407 NGKLEKALEVFDYMQKSEIKLDIYIYTTMIEGMCKAGKVDDGWDLFCSLSLKGVKPNVVT 466
Query: 490 YNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEM 548
YNT++ G C + L+ A + +M+++ P+ TYN LI+ + + A+ L+ EM
Sbjct: 467 YNTMISGLCSKRLLQEAYALLKKMKEDGPLPDSGTYNTLIRAHLRDGDKAASAELIREM 525
Score = 199 bits (505), Expect = 5e-51
Identities = 111/305 (36%), Positives = 176/305 (57%), Gaps = 9/305 (2%)
Query: 168 SLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAI 227
+L + N L+ A VKE K + E ++ +MIKR I ++ T+N ING C +L+KA+
Sbjct: 253 NLVTFNALIDAFVKEGKFVEAEKLHDDMIKRSIDPDIFTYNSLINGFCMHDRLDKAKQMF 312
Query: 228 EDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGF 287
E M + P++ TYNTL+ G+CK + ++ +EM + + VT+ TLI G
Sbjct: 313 EFMVSKDCFPDLDTYNTLIKGFCK---SKRVEDGTELFREMSHRGLVGDTVTYTTLIQGL 369
Query: 288 CKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNI 347
D + A+K F++M G+ P+I+TY+ L++GLCNNGKLE+A++++D M +K +I
Sbjct: 370 FHDGDCDNAQKVFKQMVSDGVPPDIMTYSILLDGLCNNGKLEKALEVFDYMQKSEIKLDI 429
Query: 348 VTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSS 407
Y +I G CK + + +F +S + + PNV+T+NTMI C + +++E ++L
Sbjct: 430 YIYTTMIEGMCKAGKVDDGWDLFCSLSLKGVKPNVVTYNTMISGLCSKRLLQEAYALLKK 489
Query: 408 MLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTY----NILIDGLC 463
M ++G LP+ TYN LI R D A+ EL+ EM + GD T N+L DG
Sbjct: 490 MKEDGPLPDSGTYNTLIRAHLRDGDKAASAELIREMRSCRFVGDASTIGLVANMLHDG-- 547
Query: 464 KNDKS 468
+ DKS
Sbjct: 548 RLDKS 552
Score = 127 bits (318), Expect = 2e-29
Identities = 73/247 (29%), Positives = 133/247 (53%), Gaps = 2/247 (0%)
Query: 327 KLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKM--MKEATKVFDDVSKQELVPNVIT 384
KL++AI L+ MV P+I +N L++ K K + + + + K P+++T
Sbjct: 22 KLDDAIGLFGGMVKSRPLPSIFEFNKLLSAIAKMKKFDLVISLALLGKMMKLGYEPSIVT 81
Query: 385 FNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEME 444
+++++ YC + + +L M++ G P+ T+ LI GL A L++ M
Sbjct: 82 LSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTITFTTLIHGLFLHNKASEAVALVDRMV 141
Query: 445 NKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLK 504
+G + ++VTY ++++GLCK A LLN+M ++ + V +NT++D C +
Sbjct: 142 QRGCQPNLVTYGVVVNGLCKRGDIDLAFNLLNKMEAAKIEADVVIFNTIIDSLCKYRHVD 201
Query: 505 AALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVR 564
ALN+ ME + +PNVVTY+ LI C + A+ LL++M+EK +NPN T++ +
Sbjct: 202 DALNLFKEMETKGIRPNVVTYSSLISCLCSYGRWSDASQLLSDMIEKKINPNLVTFNALI 261
Query: 565 LEMLEKG 571
+++G
Sbjct: 262 DAFVKEG 268
Score = 111 bits (277), Expect = 1e-24
Identities = 61/204 (29%), Positives = 104/204 (50%), Gaps = 2/204 (0%)
Query: 359 KKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLC--SSMLDEGILPN 416
K + +A +F + K +P++ FN ++ A K + SL M+ G P+
Sbjct: 19 KDLKLDDAIGLFGGMVKSRPLPSIFEFNKLLSAIAKMKKFDLVISLALLGKMMKLGYEPS 78
Query: 417 VSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLN 476
+ T + L+ G C + + A L+++M G + D +T+ LI GL ++K+ A L++
Sbjct: 79 IVTLSSLLNGYCHGKRISDAVALVDQMVEMGYRPDTITFTTLIHGLFLHNKASEAVALVD 138
Query: 477 EMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKIN 536
M G +PN VTY +++G C G + A N+ +ME + + +VV +N +I CK
Sbjct: 139 RMVQRGCQPNLVTYGVVVNGLCKRGDIDLAFNLLNKMEAAKIEADVVIFNTIIDSLCKYR 198
Query: 537 KLEAANGLLNEMLEKGLNPNRTTY 560
++ A L EM KG+ PN TY
Sbjct: 199 HVDDALNLFKEMETKGIRPNVVTY 222
>At5g01110 unknown protein (At5g01110)
Length = 729
Score = 264 bits (675), Expect = 9e-71
Identities = 166/563 (29%), Positives = 279/563 (49%), Gaps = 26/563 (4%)
Query: 33 KQHWSELKPHLRVTKPATFLDQLLNAGVDSELVLRFFKWSQKEY-RLSYGLEPTSKVLHF 91
KQ + ++ HL P ++ L D L RF + + S ++H
Sbjct: 63 KQGNNNVRNHLIRLNPLAVVEVLYRCRNDLTLGQRFVDQLGFHFPNFKHTSLSLSAMIHI 122
Query: 92 LANSKRYSKVRSFLDSFVKNEKHTVSSVFHSLLLDGGRPGATALIIDMLVLAYVKNLELH 151
L S R S +S L ++ + + +SL G+ + D+L+ YV+ +L
Sbjct: 123 LVRSGRLSDAQSCLLRMIRRSGVSRLEIVNSLDSTFSNCGSNDSVFDLLIRTYVQARKLR 182
Query: 152 CAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEY---VYKEMIKRRIHTNLNTFN 208
A+EAFT + GF +S+ +CN L+ +LV+ IG VE VY+E+ + + N+ T N
Sbjct: 183 EAHEAFTLLRSKGFTVSIDACNALIGSLVR---IGWVELAWGVYQEISRSGVGINVYTLN 239
Query: 209 IFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEM 268
I +N LC+ GK+ K + ++ G+ P++VTYNTL+ Y S G M +A M M
Sbjct: 240 IMVNALCKDGKMEKVGTFLSQVQEKGVYPDIVTYNTLISAY---SSKGLMEEAFELMNAM 296
Query: 269 LANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKL 328
P T+NT+I+G CK AK+ F EM + GL P+ TY SL+ C G +
Sbjct: 297 PGKGFSPGVYTYNTVINGLCKHGKYERAKEVFAEMLRSGLSPDSTTYRSLLMEACKKGDV 356
Query: 329 EEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTM 388
E ++ M + P++V ++++++ F + + +A F+ V + L+P+ + + +
Sbjct: 357 VETEKVFSDMRSRDVVPDLVCFSSMMSLFTRSGNLDKALMYFNSVKEAGLIPDNVIYTIL 416
Query: 389 IDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGL 448
I YC++GM+ +L + ML +G +V TYN ++ GLC+++ L A +L NEM + L
Sbjct: 417 IQGYCRKGMISVAMNLRNEMLQQGCAMDVVTYNTILHGLCKRKMLGEADKLFNEMTERAL 476
Query: 449 KGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALN 508
D T ILIDG CK +NA +L +M ++ + VTYNTL+DG+ G + A
Sbjct: 477 FPDSYTLTILIDGHCKLGNLQNAMELFQKMKEKRIRLDVVTYNTLLDGFGKVGDIDTAKE 536
Query: 509 VRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNP------------- 555
+ M + P ++Y++L+ C L A + +EM+ K + P
Sbjct: 537 IWADMVSKEILPTPISYSILVNALCSKGHLAEAFRVWDEMISKNIKPTVMICNSMIKGYC 596
Query: 556 ---NRTTYDIVRLEMLEKGFSPD 575
N + + +M+ +GF PD
Sbjct: 597 RSGNASDGESFLEKMISEGFVPD 619
Score = 208 bits (530), Expect = 6e-54
Identities = 121/406 (29%), Positives = 211/406 (51%), Gaps = 5/406 (1%)
Query: 153 AYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFIN 212
A E F G T+ LL K+ + + E V+ +M R + +L F+ ++
Sbjct: 324 AKEVFAEMLRSGLSPDSTTYRSLLMEACKKGDVVETEKVFSDMRSRDVVPDLVCFSSMMS 383
Query: 213 GLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANK 272
R+G L+KA +K G+ P+ V Y L+ GYC++G A EML
Sbjct: 384 LFTRSGNLDKALMYFNSVKEAGLIPDNVIYTILIQGYCRKGMISV---AMNLRNEMLQQG 440
Query: 273 ICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAI 332
+ VT+NT++ G CK + + A K F EM ++ L P+ T LI+G C G L+ A+
Sbjct: 441 CAMDVVTYNTILHGLCKRKMLGEADKLFNEMTERALFPDSYTLTILIDGHCKLGNLQNAM 500
Query: 333 DLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAY 392
+L+ KM ++ ++VTYN L++GF K + A +++ D+ +E++P I+++ +++A
Sbjct: 501 ELFQKMKEKRIRLDVVTYNTLLDGFGKVGDIDTAKEIWADMVSKEILPTPISYSILVNAL 560
Query: 393 CKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDV 452
C +G + E F + M+ + I P V N +I G CR + + L +M ++G D
Sbjct: 561 CSKGHLAEAFRVWDEMISKNIKPTVMICNSMIKGYCRSGNASDGESFLEKMISEGFVPDC 620
Query: 453 VTYNILIDGLCKNDKSRNAEKLLNEMFNL--GLKPNHVTYNTLMDGYCMEGKLKAALNVR 510
++YN LI G + + A L+ +M GL P+ TYN+++ G+C + ++K A V
Sbjct: 621 ISYNTLIYGFVREENMSKAFGLVKKMEEEQGGLVPDVFTYNSILHGFCRQNQMKEAEVVL 680
Query: 511 TRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPN 556
+M + P+ TY +I G+ + L A + +EML++G +P+
Sbjct: 681 RKMIERGVNPDRSTYTCMINGFVSQDNLTEAFRIHDEMLQRGFSPD 726
Score = 204 bits (520), Expect = 8e-53
Identities = 126/484 (26%), Positives = 239/484 (49%), Gaps = 13/484 (2%)
Query: 44 RVTKPATFLDQLLNAGVDSELVLRFFKWSQKEYRLSYGLEPTSKVLHFLANSKRYSKVRS 103
++ K TFL Q+ GV ++V + Y +E ++++ + V +
Sbjct: 250 KMEKVGTFLSQVQEKGVYPDIVT--YNTLISAYSSKGLMEEAFELMNAMPGKGFSPGVYT 307
Query: 104 F---LDSFVKNEKHT-VSSVFHSLLLDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTR 159
+ ++ K+ K+ VF +L G P +T L++ K ++ + F+
Sbjct: 308 YNTVINGLCKHGKYERAKEVFAEMLRSGLSPDSTTY--RSLLMEACKKGDVVETEKVFSD 365
Query: 160 AKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGK 219
+ L + ++S + + + + + + + + I I G CR G
Sbjct: 366 MRSRDVVPDLVCFSSMMSLFTRSGNLDKALMYFNSVKEAGLIPDNVIYTILIQGYCRKGM 425
Query: 220 LNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVT 279
++ A + +M G + +VVTYNT++ G CKR G+ A+ EM + P+ T
Sbjct: 426 ISVAMNLRNEMLQQGCAMDVVTYNTILHGLCKRKMLGE---ADKLFNEMTERALFPDSYT 482
Query: 280 FNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMV 339
LIDG CK N+ A + F++M+++ ++ ++VTYN+L++G G ++ A ++W MV
Sbjct: 483 LTILIDGHCKLGNLQNAMELFQKMKEKRIRLDVVTYNTLLDGFGKVGDIDTAKEIWADMV 542
Query: 340 GLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMME 399
+ P ++Y+ L+N C K + EA +V+D++ + + P V+ N+MI YC+ G
Sbjct: 543 SKEILPTPISYSILVNALCSKGHLAEAFRVWDEMISKNIKPTVMICNSMIKGYCRSGNAS 602
Query: 400 EGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENK--GLKGDVVTYNI 457
+G S M+ EG +P+ +YN LI G R++++ A L+ +ME + GL DV TYN
Sbjct: 603 DGESFLEKMISEGFVPDCISYNTLIYGFVREENMSKAFGLVKKMEEEQGGLVPDVFTYNS 662
Query: 458 LIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKER 517
++ G C+ ++ + AE +L +M G+ P+ TY +++G+ + L A + M +
Sbjct: 663 ILHGFCRQNQMKEAEVVLRKMIERGVNPDRSTYTCMINGFVSQDNLTEAFRIHDEMLQRG 722
Query: 518 KQPN 521
P+
Sbjct: 723 FSPD 726
>At5g12100 unknown protein (At5g12100)
Length = 816
Score = 261 bits (667), Expect = 8e-70
Identities = 160/458 (34%), Positives = 245/458 (52%), Gaps = 6/458 (1%)
Query: 105 LDSFVKNEKHTVS-SVFHSLLLDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDY 163
LD VK ++ V+ +VF ++L RP + + + A VK ++ E F R K
Sbjct: 151 LDHLVKTKQFRVTINVFLNILESDFRP--SKFMYGKAIQAAVKLSDVGKGLELFNRMKHD 208
Query: 164 GFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKA 223
S+ N L+ L K ++ D E ++ EM+ RR+ +L T+N I+G C+AG K+
Sbjct: 209 RIYPSVFIYNVLIDGLCKGKRMNDAEQLFDEMLARRLLPSLITYNTLIDGYCKAGNPEKS 268
Query: 224 EDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTL 283
E MKA I P+++T+NTL+ G K AG + AE +KEM P+ TF+ L
Sbjct: 269 FKVRERMKADHIEPSLITFNTLLKGLFK---AGMVEDAENVLKEMKDLGFVPDAFTFSIL 325
Query: 284 IDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGL 343
DG+ +E AA +E G+K N T + L+N LC GK+E+A ++ + + GL
Sbjct: 326 FDGYSSNEKAEAALGVYETAVDSGVKMNAYTCSILLNALCKEGKIEKAEEILGREMAKGL 385
Query: 344 KPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFS 403
PN V YN +I+G+C+K + A + + KQ + P+ + +N +I +C+ G ME
Sbjct: 386 VPNEVIYNTMIDGYCRKGDLVGARMKIEAMEKQGMKPDHLAYNCLIRRFCELGEMENAEK 445
Query: 404 LCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLC 463
+ M +G+ P+V TYN LI G RK + ++L EME+ G +VV+Y LI+ LC
Sbjct: 446 EVNKMKLKGVSPSVETYNILIGGYGRKYEFDKCFDILKEMEDNGTMPNVVSYGTLINCLC 505
Query: 464 KNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVV 523
K K A+ + +M + G+ P YN L+DG C +GK++ A M K+ + N+V
Sbjct: 506 KGSKLLEAQIVKRDMEDRGVSPKVRIYNMLIDGCCSKGKIEDAFRFSKEMLKKGIELNLV 565
Query: 524 TYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYD 561
TYN LI G KL A LL E+ KGL P+ TY+
Sbjct: 566 TYNTLIDGLSMTGKLSEAEDLLLEISRKGLKPDVFTYN 603
Score = 253 bits (645), Expect = 3e-67
Identities = 157/489 (32%), Positives = 252/489 (51%), Gaps = 24/489 (4%)
Query: 107 SFVKNEKHTVSS---VFHSLLLDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAKDY 163
S + NE +S +F +L +G P + +L + L+ VK + F +
Sbjct: 116 SVLLNESKMISEAADLFFALRNEGIYPSSDSLTL--LLDHLVKTKQFRVTINVFLNILES 173
Query: 164 GFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKA 223
F+ S + A VK + +G ++ M RI+ ++ +N+ I+GLC+ ++N A
Sbjct: 174 DFRPSKFMYGKAIQAAVKLSDVGKGLELFNRMKHDRIYPSVFIYNVLIDGLCKGKRMNDA 233
Query: 224 EDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTL 283
E ++M A + P+++TYNTL+DGYCK G+ K +K MK A+ I P+ +TFNTL
Sbjct: 234 EQLFDEMLARRLLPSLITYNTLIDGYCKAGNPEKSFKVRERMK---ADHIEPSLITFNTL 290
Query: 284 IDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGL 343
+ G K V A+ +EM+ G P+ T++ L +G +N K E A+ +++ V G+
Sbjct: 291 LKGLFKAGMVEDAENVLKEMKDLGFVPDAFTFSILFDGYSSNEKAEAALGVYETAVDSGV 350
Query: 344 KPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFS 403
K N T + L+N CK+ +++A ++ + LVPN + +NTMID YC++G +
Sbjct: 351 KMNAYTCSILLNALCKEGKIEKAEEILGREMAKGLVPNEVIYNTMIDGYCRKGDLVGARM 410
Query: 404 LCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLC 463
+M +G+ P+ YNCLI C +++ A++ +N+M+ KG+ V TYNILI G
Sbjct: 411 KIEAMEKQGMKPDHLAYNCLIRRFCELGEMENAEKEVNKMKLKGVSPSVETYNILIGGYG 470
Query: 464 KNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVV 523
+ + +L EM + G PN V+Y TL++ C KL A V+ ME P V
Sbjct: 471 RKYEFDKCFDILKEMEDNGTMPNVVSYGTLINCLCKGSKLLEAQIVKRDMEDRGVSPKVR 530
Query: 524 TYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDI----------------VRLEM 567
YN+LI G C K+E A EML+KG+ N TY+ + LE+
Sbjct: 531 IYNMLIDGCCSKGKIEDAFRFSKEMLKKGIELNLVTYNTLIDGLSMTGKLSEAEDLLLEI 590
Query: 568 LEKGFSPDI 576
KG PD+
Sbjct: 591 SRKGLKPDV 599
Score = 224 bits (570), Expect = 1e-58
Identities = 147/467 (31%), Positives = 221/467 (46%), Gaps = 58/467 (12%)
Query: 161 KDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKL 220
KD GF + + L K VY+ + + N T +I +N LC+ GK+
Sbjct: 311 KDLGFVPDAFTFSILFDGYSSNEKAEAALGVYETAVDSGVKMNAYTCSILLNALCKEGKI 370
Query: 221 NKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGS-AGKMYKAEAFMKEMLANKICPNEVT 279
KAE+ + A G+ PN V YNT++DGYC++G G K EA K+ + P+ +
Sbjct: 371 EKAEEILGREMAKGLVPNEVIYNTMIDGYCRKGDLVGARMKIEAMEKQGMK----PDHLA 426
Query: 280 FNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMV 339
+N LI FC+ + A+K +M+ +G+ P++ TYN LI G + ++ D+ +M
Sbjct: 427 YNCLIRRFCELGEMENAEKEVNKMKLKGVSPSVETYNILIGGYGRKYEFDKCFDILKEME 486
Query: 340 GLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMME 399
G PN+V+Y LIN CK + EA V D+ + + P V +N +ID C +G +E
Sbjct: 487 DNGTMPNVVSYGTLINCLCKGSKLLEAQIVKRDMEDRGVSPKVRIYNMLIDGCCSKGKIE 546
Query: 400 EGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILI 459
+ F ML +GI N+ TYN LI GL L A++LL E+ KGLK DV TYN LI
Sbjct: 547 DAFRFSKEMLKKGIELNLVTYNTLIDGLSMTGKLSEAEDLLLEISRKGLKPDVFTYNSLI 606
Query: 460 DG----------------------------------LCKNDKSRNAEKLLNEMFNLGLKP 485
G LC + E+L EM LKP
Sbjct: 607 SGYGFAGNVQRCIALYEEMKRSGIKPTLKTYHLLISLCTKEGIELTERLFGEM---SLKP 663
Query: 486 NHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLL 545
+ + YN ++ Y + G ++ A N++ +M ++ + TYN LI G K+ KL L+
Sbjct: 664 DLLVYNGVLHCYAVHGDMEKAFNLQKQMIEKSIGLDKTTYNSLILGQLKVGKLCEVRSLI 723
Query: 546 NEMLEKGLNPNRTTYDIV----------------RLEMLEKGFSPDI 576
+EM + + P TY+I+ EM EKGF D+
Sbjct: 724 DEMNAREMEPEADTYNIIVKGHCEVKDYMSAYVWYREMQEKGFLLDV 770
Score = 211 bits (536), Expect = 1e-54
Identities = 134/454 (29%), Positives = 216/454 (47%), Gaps = 42/454 (9%)
Query: 134 ALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYK 193
A +L Y N + A + A D G K++ +C+ LL+AL KE KI E +
Sbjct: 319 AFTFSILFDGYSSNEKAEAALGVYETAVDSGVKMNAYTCSILLNALCKEGKIEKAEEILG 378
Query: 194 EMIKRRIHTNLNTFNIFINGLCRAGKL--------------------------------- 220
+ + + N +N I+G CR G L
Sbjct: 379 REMAKGLVPNEVIYNTMIDGYCRKGDLVGARMKIEAMEKQGMKPDHLAYNCLIRRFCELG 438
Query: 221 --NKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEV 278
AE + MK G+SP+V TYN L+ GY ++ K + +KEM N PN V
Sbjct: 439 EMENAEKEVNKMKLKGVSPSVETYNILIGGYGRKYEFDKCFD---ILKEMEDNGTMPNVV 495
Query: 279 TFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKM 338
++ TLI+ CK + A+ +M+ +G+ P + YN LI+G C+ GK+E+A +M
Sbjct: 496 SYGTLINCLCKGSKLLEAQIVKRDMEDRGVSPKVRIYNMLIDGCCSKGKIEDAFRFSKEM 555
Query: 339 VGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMM 398
+ G++ N+VTYN LI+G + EA + ++S++ L P+V T+N++I Y G +
Sbjct: 556 LKKGIELNLVTYNTLIDGLSMTGKLSEAEDLLLEISRKGLKPDVFTYNSLISGYGFAGNV 615
Query: 399 EEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNIL 458
+ +L M GI P + TY+ LI+ LC K+ ++ + L EM LK D++ YN +
Sbjct: 616 QRCIALYEEMKRSGIKPTLKTYHLLIS-LCTKEGIELTERLFGEMS---LKPDLLVYNGV 671
Query: 459 IDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERK 518
+ + A L +M + + TYN+L+ G GKL ++ M
Sbjct: 672 LHCYAVHGDMEKAFNLQKQMIEKSIGLDKTTYNSLILGQLKVGKLCEVRSLIDEMNAREM 731
Query: 519 QPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKG 552
+P TYN+++KG+C++ +A EM EKG
Sbjct: 732 EPEADTYNIIVKGHCEVKDYMSAYVWYREMQEKG 765
Score = 175 bits (443), Expect = 7e-44
Identities = 118/379 (31%), Positives = 197/379 (51%), Gaps = 7/379 (1%)
Query: 135 LIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKE 194
L + L+ + + E+ A + + K G S+ + N L+ ++ + + KE
Sbjct: 425 LAYNCLIRRFCELGEMENAEKEVNKMKLKGVSPSVETYNILIGGYGRKYEFDKCFDILKE 484
Query: 195 MIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGS 254
M N+ ++ IN LC+ KL +A+ DM+ G+SP V YN L+DG C S
Sbjct: 485 MEDNGTMPNVVSYGTLINCLCKGSKLLEAQIVKRDMEDRGVSPKVRIYNMLIDGCC---S 541
Query: 255 AGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVT 314
GK+ A F KEML I N VT+NTLIDG ++ A+ E+ ++GLKP++ T
Sbjct: 542 KGKIEDAFRFSKEMLKKGIELNLVTYNTLIDGLSMTGKLSEAEDLLLEISRKGLKPDVFT 601
Query: 315 YNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVS 374
YNSLI+G G ++ I L+++M G+KP + TY+ LI+ C K+ ++ ++F ++S
Sbjct: 602 YNSLISGYGFAGNVQRCIALYEEMKRSGIKPTLKTYHLLIS-LCTKEGIELTERLFGEMS 660
Query: 375 KQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQ 434
L P+++ +N ++ Y G ME+ F+L M+++ I + +TYN LI G + L
Sbjct: 661 ---LKPDLLVYNGVLHCYAVHGDMEKAFNLQKQMIEKSIGLDKTTYNSLILGQLKVGKLC 717
Query: 435 AAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLM 494
+ L++EM + ++ + TYNI++ G C+ +A EM G + N L+
Sbjct: 718 EVRSLIDEMNAREMEPEADTYNIIVKGHCEVKDYMSAYVWYREMQEKGFLLDVCIGNELV 777
Query: 495 DGYCMEGKLKAALNVRTRM 513
G E + K A V + M
Sbjct: 778 SGLKEEWRSKEAEIVISEM 796
Score = 140 bits (354), Expect = 2e-33
Identities = 89/292 (30%), Positives = 155/292 (52%), Gaps = 8/292 (2%)
Query: 161 KDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKL 220
+D G + N L+ + KI D KEM+K+ I NL T+N I+GL GKL
Sbjct: 521 EDRGVSPKVRIYNMLIDGCCSKGKIEDAFRFSKEMLKKGIELNLVTYNTLIDGLSMTGKL 580
Query: 221 NKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTF 280
++AED + ++ G+ P+V TYN+L+ GY G AG + + A +EM + I P T+
Sbjct: 581 SEAEDLLLEISRKGLKPDVFTYNSLISGY---GFAGNVQRCIALYEEMKRSGIKPTLKTY 637
Query: 281 NTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVG 340
+ LI C E + ++ F EM LKP+++ YN +++ +G +E+A +L +M+
Sbjct: 638 HLLIS-LCTKEGIELTERLFGEMS---LKPDLLVYNGVLHCYAVHGDMEKAFNLQKQMIE 693
Query: 341 LGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEE 400
+ + TYN+LI G K + E + D+++ +E+ P T+N ++ +C+
Sbjct: 694 KSIGLDKTTYNSLILGQLKVGKLCEVRSLIDEMNAREMEPEADTYNIIVKGHCEVKDYMS 753
Query: 401 GFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDV 452
+ M ++G L +V N L++GL + + A+ +++EM + L GDV
Sbjct: 754 AYVWYREMQEKGFLLDVCIGNELVSGLKEEWRSKEAEIVISEMNGRML-GDV 804
Score = 64.7 bits (156), Expect = 1e-10
Identities = 43/190 (22%), Positives = 78/190 (40%)
Query: 397 MMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYN 456
M+ E L ++ +EGI P+ + L+ L + + + + + + Y
Sbjct: 124 MISEAADLFFALRNEGIYPSSDSLTLLLDHLVKTKQFRVTINVFLNILESDFRPSKFMYG 183
Query: 457 ILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKE 516
I K +L N M + + P+ YN L+DG C ++ A + M
Sbjct: 184 KAIQAAVKLSDVGKGLELFNRMKHDRIYPSVFIYNVLIDGLCKGKRMNDAEQLFDEMLAR 243
Query: 517 RKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKGFSPDI 576
R P+++TYN LI GYCK E + + M + P+ T++ + + + G D
Sbjct: 244 RLLPSLITYNTLIDGYCKAGNPEKSFKVRERMKADHIEPSLITFNTLLKGLFKAGMVEDA 303
Query: 577 EGHLYNISSM 586
E L + +
Sbjct: 304 ENVLKEMKDL 313
Score = 57.4 bits (137), Expect = 2e-08
Identities = 42/160 (26%), Positives = 69/160 (42%), Gaps = 5/160 (3%)
Query: 402 FSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDG 461
FSL S L +Y L L + + A +L + N+G+ + +L+D
Sbjct: 99 FSLSSPSLKHDF-----SYLLLSVLLNESKMISEAADLFFALRNEGIYPSSDSLTLLLDH 153
Query: 462 LCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPN 521
L K + R + + +P+ Y + + L + RM+ +R P+
Sbjct: 154 LVKTKQFRVTINVFLNILESDFRPSKFMYGKAIQAAVKLSDVGKGLELFNRMKHDRIYPS 213
Query: 522 VVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYD 561
V YNVLI G CK ++ A L +EML + L P+ TY+
Sbjct: 214 VFIYNVLIDGLCKGKRMNDAEQLFDEMLARRLLPSLITYN 253
>At3g22470 hypothetical protein
Length = 619
Score = 260 bits (664), Expect = 2e-69
Identities = 132/433 (30%), Positives = 247/433 (56%), Gaps = 3/433 (0%)
Query: 139 MLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKR 198
+++ Y + +L A+ RA G++ + + L++ E ++ + + M++
Sbjct: 110 IMINCYCRKKKLLFAFSVLGRAWKLGYEPDTITFSTLVNGFCLEGRVSEAVALVDRMVEM 169
Query: 199 RIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKM 258
+ +L T + INGLC G++++A I+ M +G P+ VTY +++ CK G++
Sbjct: 170 KQRPDLVTVSTLINGLCLKGRVSEALVLIDRMVEYGFQPDEVTYGPVLNRLCKSGNSAL- 228
Query: 259 YKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSL 318
A ++M I + V ++ +ID CKD + A F EM+ +G+K ++VTY+SL
Sbjct: 229 --ALDLFRKMEERNIKASVVQYSIVIDSLCKDGSFDDALSLFNEMEMKGIKADVVTYSSL 286
Query: 319 INGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQEL 378
I GLCN+GK ++ + +M+G + P++VT++ALI+ F K+ + EA ++++++ + +
Sbjct: 287 IGGLCNDGKWDDGAKMLREMIGRNIIPDVVTFSALIDVFVKEGKLLEAKELYNEMITRGI 346
Query: 379 VPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKE 438
P+ IT+N++ID +CKE + E + M+ +G P++ TY+ LI C+ + +
Sbjct: 347 APDTITYNSLIDGFCKENCLHEANQMFDLMVSKGCEPDIVTYSILINSYCKAKRVDDGMR 406
Query: 439 LLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYC 498
L E+ +KGL + +TYN L+ G C++ K A++L EM + G+ P+ VTY L+DG C
Sbjct: 407 LFREISSKGLIPNTITYNTLVLGFCQSGKLNAAKELFQEMVSRGVPPSVVTYGILLDGLC 466
Query: 499 MEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRT 558
G+L AL + +M+K R + YN++I G C +K++ A L + +KG+ P+
Sbjct: 467 DNGELNKALEIFEKMQKSRMTLGIGIYNIIIHGMCNASKVDDAWSLFCSLSDKGVKPDVV 526
Query: 559 TYDIVRLEMLEKG 571
TY+++ + +KG
Sbjct: 527 TYNVMIGGLCKKG 539
Score = 258 bits (659), Expect = 6e-69
Identities = 154/496 (31%), Positives = 264/496 (53%), Gaps = 33/496 (6%)
Query: 81 GLEPTSKVLHFLANS----KRYSKVRSFLDSFVKNEKH----TVSSVFHSLLLDGGRPGA 132
G EP + L N R S+ + +D V+ ++ TVS++ + L L G R
Sbjct: 135 GYEPDTITFSTLVNGFCLEGRVSEAVALVDRMVEMKQRPDLVTVSTLINGLCLKG-RVSE 193
Query: 133 TALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVY 192
++ID +V +YGF+ + P+L+ L K ++
Sbjct: 194 ALVLIDRMV--------------------EYGFQPDEVTYGPVLNRLCKSGNSALALDLF 233
Query: 193 KEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKR 252
++M +R I ++ ++I I+ LC+ G + A +M+ GI +VVTY++L+ G C
Sbjct: 234 RKMEERNIKASVVQYSIVIDSLCKDGSFDDALSLFNEMEMKGIKADVVTYSSLIGGLC-- 291
Query: 253 GSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNI 312
+ GK ++EM+ I P+ VTF+ LID F K+ + AK+ + EM +G+ P+
Sbjct: 292 -NDGKWDDGAKMLREMIGRNIIPDVVTFSALIDVFVKEGKLLEAKELYNEMITRGIAPDT 350
Query: 313 VTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDD 372
+TYNSLI+G C L EA ++D MV G +P+IVTY+ LIN +CK K + + ++F +
Sbjct: 351 ITYNSLIDGFCKENCLHEANQMFDLMVSKGCEPDIVTYSILINSYCKAKRVDDGMRLFRE 410
Query: 373 VSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQD 432
+S + L+PN IT+NT++ +C+ G + L M+ G+ P+V TY L+ GLC +
Sbjct: 411 ISSKGLIPNTITYNTLVLGFCQSGKLNAAKELFQEMVSRGVPPSVVTYGILLDGLCDNGE 470
Query: 433 LQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNT 492
L A E+ +M+ + + YNI+I G+C K +A L + + G+KP+ VTYN
Sbjct: 471 LNKALEIFEKMQKSRMTLGIGIYNIIIHGMCNASKVDDAWSLFCSLSDKGVKPDVVTYNV 530
Query: 493 LMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKG 552
++ G C +G L A + +M+++ P+ TYN+LI+ + + L ++ L+ EM G
Sbjct: 531 MIGGLCKKGSLSEADMLFRKMKEDGCTPDDFTYNILIRAHLGGSGLISSVELIEEMKVCG 590
Query: 553 LNPNRTTYDIVRLEML 568
+ + +T +V ++ML
Sbjct: 591 FSADSSTIKMV-IDML 605
Score = 229 bits (583), Expect = 4e-60
Identities = 126/399 (31%), Positives = 216/399 (53%), Gaps = 3/399 (0%)
Query: 173 NPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKA 232
N L SA+ + + V K M I ++ T I IN CR KL A +
Sbjct: 74 NRLCSAVARTKQYDLVLGFCKGMELNGIEHDMYTMTIMINCYCRKKKLLFAFSVLGRAWK 133
Query: 233 WGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDEN 292
G P+ +T++TLV+G+C G++ +A A + M+ K P+ VT +TLI+G C
Sbjct: 134 LGYEPDTITFSTLVNGFCLE---GRVSEAVALVDRMVEMKQRPDLVTVSTLINGLCLKGR 190
Query: 293 VAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNA 352
V+ A + M + G +P+ VTY ++N LC +G A+DL+ KM +K ++V Y+
Sbjct: 191 VSEALVLIDRMVEYGFQPDEVTYGPVLNRLCKSGNSALALDLFRKMEERNIKASVVQYSI 250
Query: 353 LINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEG 412
+I+ CK +A +F+++ + + +V+T++++I C +G ++G + M+
Sbjct: 251 VIDSLCKDGSFDDALSLFNEMEMKGIKADVVTYSSLIGGLCNDGKWDDGAKMLREMIGRN 310
Query: 413 ILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAE 472
I+P+V T++ LI ++ L AKEL NEM +G+ D +TYN LIDG CK + A
Sbjct: 311 IIPDVVTFSALIDVFVKEGKLLEAKELYNEMITRGIAPDTITYNSLIDGFCKENCLHEAN 370
Query: 473 KLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGY 532
++ + M + G +P+ VTY+ L++ YC ++ + + + + PN +TYN L+ G+
Sbjct: 371 QMFDLMVSKGCEPDIVTYSILINSYCKAKRVDDGMRLFREISSKGLIPNTITYNTLVLGF 430
Query: 533 CKINKLEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
C+ KL AA L EM+ +G+ P+ TY I+ + + G
Sbjct: 431 CQSGKLNAAKELFQEMVSRGVPPSVVTYGILLDGLCDNG 469
Score = 197 bits (502), Expect = 1e-50
Identities = 116/346 (33%), Positives = 178/346 (50%), Gaps = 5/346 (1%)
Query: 219 KLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAE-AFMKEMLANKICPNE 277
K+N A D E M P + +N L C + K Y F K M N I +
Sbjct: 50 KVNDAIDLFESMIQSRPLPTPIDFNRL----CSAVARTKQYDLVLGFCKGMELNGIEHDM 105
Query: 278 VTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDK 337
T +I+ +C+ + + A K G +P+ +T+++L+NG C G++ EA+ L D+
Sbjct: 106 YTMTIMINCYCRKKKLLFAFSVLGRAWKLGYEPDTITFSTLVNGFCLEGRVSEAVALVDR 165
Query: 338 MVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGM 397
MV + +P++VT + LING C K + EA + D + + P+ +T+ +++ CK G
Sbjct: 166 MVEMKQRPDLVTVSTLINGLCLKGRVSEALVLIDRMVEYGFQPDEVTYGPVLNRLCKSGN 225
Query: 398 MEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNI 457
L M + I +V Y+ +I LC+ A L NEME KG+K DVVTY+
Sbjct: 226 SALALDLFRKMEERNIKASVVQYSIVIDSLCKDGSFDDALSLFNEMEMKGIKADVVTYSS 285
Query: 458 LIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKER 517
LI GLC + K + K+L EM + P+ VT++ L+D + EGKL A + M
Sbjct: 286 LIGGLCNDGKWDDGAKMLREMIGRNIIPDVVTFSALIDVFVKEGKLLEAKELYNEMITRG 345
Query: 518 KQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIV 563
P+ +TYN LI G+CK N L AN + + M+ KG P+ TY I+
Sbjct: 346 IAPDTITYNSLIDGFCKENCLHEANQMFDLMVSKGCEPDIVTYSIL 391
Score = 129 bits (323), Expect = 6e-30
Identities = 87/307 (28%), Positives = 141/307 (45%), Gaps = 35/307 (11%)
Query: 290 DENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVT 349
D V A FE M + P + +N L + + + + + M G++ ++ T
Sbjct: 48 DIKVNDAIDLFESMIQSRPLPTPIDFNRLCSAVARTKQYDLVLGFCKGMELNGIEHDMYT 107
Query: 350 YNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSML 409
+IN +C+KK + A V K P+ ITF+T+++ +C EG + E +L M+
Sbjct: 108 MTIMINCYCRKKKLLFAFSVLGRAWKLGYEPDTITFSTLVNGFCLEGRVSEAVALVDRMV 167
Query: 410 DEGILPNVSTYNCLIAGLCRKQDLQ----------------------------------- 434
+ P++ T + LI GLC K +
Sbjct: 168 EMKQRPDLVTVSTLINGLCLKGRVSEALVLIDRMVEYGFQPDEVTYGPVLNRLCKSGNSA 227
Query: 435 AAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLM 494
A +L +ME + +K VV Y+I+ID LCK+ +A L NEM G+K + VTY++L+
Sbjct: 228 LALDLFRKMEERNIKASVVQYSIVIDSLCKDGSFDDALSLFNEMEMKGIKADVVTYSSLI 287
Query: 495 DGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLN 554
G C +GK + M P+VVT++ LI + K KL A L NEM+ +G+
Sbjct: 288 GGLCNDGKWDDGAKMLREMIGRNIIPDVVTFSALIDVFVKEGKLLEAKELYNEMITRGIA 347
Query: 555 PNRTTYD 561
P+ TY+
Sbjct: 348 PDTITYN 354
Score = 98.6 bits (244), Expect = 9e-21
Identities = 65/263 (24%), Positives = 123/263 (46%), Gaps = 6/263 (2%)
Query: 103 SFLDSFVK-NEKHTVSSVFHSLLLDGGRPGATALIIDMLVLAYVKNLELHCAYEAFTRAK 161
S +D F K N H + +F ++ G P I L+ +Y K + F
Sbjct: 355 SLIDGFCKENCLHEANQMFDLMVSKGCEPDIVTYSI--LINSYCKAKRVDDGMRLFREIS 412
Query: 162 DYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLN 221
G + + N L+ + K+ + +++EM+ R + ++ T+ I ++GLC G+LN
Sbjct: 413 SKGLIPNTITYNTLVLGFCQSGKLNAAKELFQEMVSRGVPPSVVTYGILLDGLCDNGELN 472
Query: 222 KAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFN 281
KA + E M+ ++ + YN ++ G C +A K+ A + + + P+ VT+N
Sbjct: 473 KALEIFEKMQKSRMTLGIGIYNIIIHGMC---NASKVDDAWSLFCSLSDKGVKPDVVTYN 529
Query: 282 TLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGL 341
+I G CK +++ A F +M++ G P+ TYN LI L +++L ++M
Sbjct: 530 VMIGGLCKKGSLSEADMLFRKMKEDGCTPDDFTYNILIRAHLGGSGLISSVELIEEMKVC 589
Query: 342 GLKPNIVTYNALINGFCKKKMMK 364
G + T +I+ +++ K
Sbjct: 590 GFSADSSTIKMVIDMLSDRRLDK 612
Score = 85.5 bits (210), Expect = 7e-17
Identities = 63/257 (24%), Positives = 116/257 (44%), Gaps = 16/257 (6%)
Query: 78 LSYGLEPTSKVLHFLANSKRYSKVRSFLDSFVKNEKHTVSSVFHSLLLDGGRPGATALII 137
+S G EP L NS Y K + D +F + G P +
Sbjct: 377 VSKGCEPDIVTYSILINS--YCKAKRVDDGM---------RLFREISSKGLIPNT--ITY 423
Query: 138 DMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIK 197
+ LVL + ++ +L+ A E F G S+ + LL L ++ ++++M K
Sbjct: 424 NTLVLGFCQSGKLNAAKELFQEMVSRGVPPSVVTYGILLDGLCDNGELNKALEIFEKMQK 483
Query: 198 RRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAGK 257
R+ + +NI I+G+C A K++ A + G+ P+VVTYN ++ G CK+GS
Sbjct: 484 SRMTLGIGIYNIIIHGMCNASKVDDAWSLFCSLSDKGVKPDVVTYNVMIGGLCKKGS--- 540
Query: 258 MYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNS 317
+ +A+ ++M + P++ T+N LI + ++ + EEM+ G + T
Sbjct: 541 LSEADMLFRKMKEDGCTPDDFTYNILIRAHLGGSGLISSVELIEEMKVCGFSADSSTIKM 600
Query: 318 LINGLCNNGKLEEAIDL 334
+I+ L + + +D+
Sbjct: 601 VIDMLSDRRLDKSFLDM 617
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,068,794
Number of Sequences: 26719
Number of extensions: 570472
Number of successful extensions: 15236
Number of sequences better than 10.0: 476
Number of HSP's better than 10.0 without gapping: 457
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 1466
Number of HSP's gapped (non-prelim): 2981
length of query: 587
length of database: 11,318,596
effective HSP length: 105
effective length of query: 482
effective length of database: 8,513,101
effective search space: 4103314682
effective search space used: 4103314682
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC122726.13


