

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122165.8 - phase: 1 /pseudo
(144 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
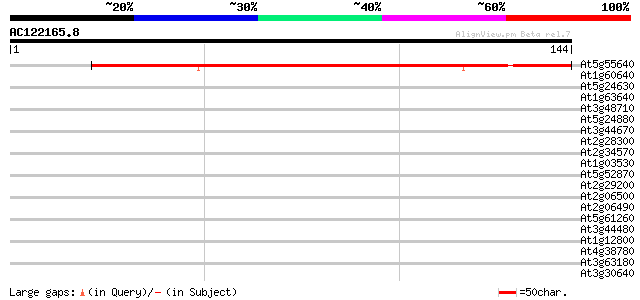
Score E
Sequences producing significant alignments: (bits) Value
At5g55640 Unknown protein (MDF20.8) 144 1e-35
At1g60640 unknown protein 33 0.052
At5g24630 unknown protein 32 0.15
At1g63640 kinesin-like protein 32 0.15
At3g48710 unknown protein 30 0.44
At5g24880 glutamic acid-rich protein 30 0.57
At3g44670 disease resistance protein homlog 29 0.75
At2g28300 unknown protein 29 0.75
At2g34570 unknown protein 29 0.98
At1g03530 unknown protein 29 0.98
At5g52870 unknown protein (At5g52870) 28 1.3
At2g29200 putative pumilio/Mpt5 family RNA-binding protein 28 1.3
At2g06500 Ac-like transposase 28 1.3
At2g06490 putative PttA-like transposon protein 28 1.3
At5g61260 putative protein 28 1.7
At3g44480 disease resistance protein -like 28 1.7
At1g12800 unknown protein 28 1.7
At4g38780 splicing factor - like protein 28 2.2
At3g63180 hypothetical protein 28 2.2
At3g30640 hypothetical protein 28 2.2
>At5g55640 Unknown protein (MDF20.8)
Length = 149
Score = 144 bits (364), Expect = 1e-35
Identities = 76/127 (59%), Positives = 94/127 (73%), Gaps = 5/127 (3%)
Query: 22 DTDGFVIPSLGIQDSDQSKNNAASVE-SPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIP 80
DTDGFVIPSL I++ + NN + VE S S+ K+K EE IYLGPHGAPPSQ +
Sbjct: 20 DTDGFVIPSLEIEEDKVNNNNDSEVETSKPSSPKSKAEENIYLGPHGAPPSQLQDGGSNT 79
Query: 81 SNRKQRFKQKLKEADKRISGTGRENKVDNLRELVG---GEKTSVGMAKGSSPKDWLDPHC 137
S+RKQRFKQ LKEAD+++SG+GRENK+ NLRELVG G + M KG S +DWLDPHC
Sbjct: 80 SSRKQRFKQMLKEADQKMSGSGRENKMANLRELVGGGAGAEKGTNMGKGVS-RDWLDPHC 138
Query: 138 HEAEFER 144
HE++FE+
Sbjct: 139 HESQFEK 145
>At1g60640 unknown protein
Length = 340
Score = 33.1 bits (74), Expect = 0.052
Identities = 21/84 (25%), Positives = 39/84 (46%), Gaps = 7/84 (8%)
Query: 33 IQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGA-----PPSQPKQQEVIPSNRKQRF 87
++ + Q K + S +++K+EK L +G P P Q++ I NR+ R
Sbjct: 162 LETASQEKEKKGNRRSQRQGKRSQKQEKDSLTKNGENEEVEDPETPSQEKQIKGNRRARR 221
Query: 88 KQKLKEADKRISGT--GRENKVDN 109
+ + + ++ S T G +VDN
Sbjct: 222 ELRRSQKQEKDSSTKHGENEEVDN 245
>At5g24630 unknown protein
Length = 531
Score = 31.6 bits (70), Expect = 0.15
Identities = 30/108 (27%), Positives = 49/108 (44%), Gaps = 18/108 (16%)
Query: 30 SLGIQDSDQSKNNAASVESPNSAAKTKKEE----KIYLGPHGAPPSQPKQQEVIPSNRKQ 85
SL ++DS K N +V+ ++K K + ++L + PS P +QEV S
Sbjct: 146 SLNLEDSCAGKENGNNVDCEKLSSKHKDAQGGADSVWLVSSDSEPSSPIKQEVTVSTE-- 203
Query: 86 RFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGS--SPKD 131
K+AD + T E V +R+ EK+ +K S +PK+
Sbjct: 204 ------KDADFVLEATEEEPAVKTVRK----EKSPKTKSKSSRKTPKE 241
>At1g63640 kinesin-like protein
Length = 1071
Score = 31.6 bits (70), Expect = 0.15
Identities = 17/59 (28%), Positives = 33/59 (55%), Gaps = 1/59 (1%)
Query: 74 KQQEVIPSNRKQRFKQKLKEADKRISGTGRENK-VDNLRELVGGEKTSVGMAKGSSPKD 131
K Q ++ R+++++ ++K + +GT +EN+ V N E + EKT + + S KD
Sbjct: 255 KNQNILFRVREEKYRSRIKVLESLAAGTTKENEIVTNCMEHIKLEKTRIEEKERSEEKD 313
>At3g48710 unknown protein
Length = 462
Score = 30.0 bits (66), Expect = 0.44
Identities = 20/96 (20%), Positives = 43/96 (43%), Gaps = 2/96 (2%)
Query: 34 QDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKE 93
+++++S++ E + KTK +K L ++E + ++ ++
Sbjct: 276 EENNKSEDTETEDEKDKAKEKTKSTDKKRLSKRTKKEKPAAEEEKSIKGSAKSSRKSFRQ 335
Query: 94 ADKRISGTGRENKV--DNLRELVGGEKTSVGMAKGS 127
DK + + ++ KV D+ + G +TS AKGS
Sbjct: 336 VDKSTTSSSKKQKVDKDDSSKEKGKTQTSKPQAKGS 371
>At5g24880 glutamic acid-rich protein
Length = 443
Score = 29.6 bits (65), Expect = 0.57
Identities = 22/89 (24%), Positives = 41/89 (45%), Gaps = 4/89 (4%)
Query: 46 VESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGREN 105
VES + K+EE G + ++++V ++K++ +++ KE ++ G +
Sbjct: 325 VESVEETTQEKEEEVKEEGKERVEEEEKEKEKVKEDDQKEKVEEEEKE---KVKGDEEKE 381
Query: 106 KVDNLRELVGGEKTSVGMAKGSSPKDWLD 134
KV E G+K V K SP + D
Sbjct: 382 KVKE-EESAEGKKKEVVKGKKESPSAYND 409
>At3g44670 disease resistance protein homlog
Length = 1199
Score = 29.3 bits (64), Expect = 0.75
Identities = 22/100 (22%), Positives = 45/100 (45%), Gaps = 6/100 (6%)
Query: 34 QDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKE 93
QD+ +K++A S+ SP ++ + ++ HGA + ++ S R++ +
Sbjct: 69 QDNKYTKSSALSLPSPPTSVSRIWKHHVFPSFHGADVRKTILSHILESFRRKGIDPFIDN 128
Query: 94 ADKRISGTGRENKVDNLRELVGGEKTS-VGMAKGSSPKDW 132
+R G E L+E + G K + V ++K + W
Sbjct: 129 NIERSKSIGHE-----LKEAIKGSKIAIVLLSKNYASSSW 163
>At2g28300 unknown protein
Length = 2218
Score = 29.3 bits (64), Expect = 0.75
Identities = 25/119 (21%), Positives = 45/119 (37%), Gaps = 16/119 (13%)
Query: 33 IQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQ------PKQQEVI------- 79
I+D DQS A E + +A+ E ++ L P P + E++
Sbjct: 1607 IEDKDQSHVETAGSELVDVSAECSTEPQVQLPPSSEPVGDMHVHLGASKSEIVAEGTDFS 1666
Query: 80 ---PSNRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDWLDP 135
P ++ K +L + + S T + +++ E G E V + P+ L P
Sbjct: 1667 SSLPKTEEENAKSQLADTEPSSSLTAVQKNIEDQVETAGCEFVVVSTGCSTEPQVQLPP 1725
>At2g34570 unknown protein
Length = 281
Score = 28.9 bits (63), Expect = 0.98
Identities = 21/79 (26%), Positives = 38/79 (47%), Gaps = 4/79 (5%)
Query: 15 FGLLLVVDTDGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPK 74
+G+ VV T LG++D Q K N A +P S K KKE + + K
Sbjct: 198 WGMPRVVSTKN----GLGVKDRPQFKRNRAKGPNPLSCMKKKKENPQSKSKADSNSNAQK 253
Query: 75 QQEVIPSNRKQRFKQKLKE 93
+++ S+ ++R +++ K+
Sbjct: 254 EKKEGGSDTQKRSRKRSKK 272
>At1g03530 unknown protein
Length = 801
Score = 28.9 bits (63), Expect = 0.98
Identities = 20/99 (20%), Positives = 47/99 (47%), Gaps = 10/99 (10%)
Query: 19 LVVDTDGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEV 78
L VD D + G DS +S++ +S + +S + + +EE+ S + +
Sbjct: 227 LAVDDDEKSDEAKGEMDSAESESETSSSSASSSDSSSSEEEE----------SDEDESDK 276
Query: 79 IPSNRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGE 117
+ ++++F+ + + ++G E +++NL E G +
Sbjct: 277 EENKKEEKFEHMVVGKEDDLAGELEEGEIENLDEENGDD 315
>At5g52870 unknown protein (At5g52870)
Length = 326
Score = 28.5 bits (62), Expect = 1.3
Identities = 18/76 (23%), Positives = 33/76 (42%), Gaps = 2/76 (2%)
Query: 35 DSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQ--KLK 92
D SK + ++ + + K+ P +P ++ KQ+ IPS + +Q K +
Sbjct: 169 DDSVSKRFFSLIKPLYTKSTKKQSSSTITSPTSSPATREKQRSNIPSGMRSVRRQLGKSR 228
Query: 93 EADKRISGTGRENKVD 108
A I G N++D
Sbjct: 229 SASAAIGGMSPANRID 244
>At2g29200 putative pumilio/Mpt5 family RNA-binding protein
Length = 968
Score = 28.5 bits (62), Expect = 1.3
Identities = 16/60 (26%), Positives = 26/60 (42%)
Query: 73 PKQQEVIPSNRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDW 132
P+ + S R Q+LK ++ + G G KV++ R L G G+S +W
Sbjct: 116 PRLPPPLMSREDLRVAQRLKGSNNVLGGVGDRRKVNDNRSLFSMPPGFEGEKTGASASEW 175
>At2g06500 Ac-like transposase
Length = 582
Score = 28.5 bits (62), Expect = 1.3
Identities = 20/59 (33%), Positives = 29/59 (48%), Gaps = 10/59 (16%)
Query: 56 KKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGRENKVDNLRELV 114
K++EK + + P P EVI S+ +++ E D ENKVDNL E+V
Sbjct: 9 KRDEKSW---YMKKPENPTISEVINSDGNNNQQERADEVDP-------ENKVDNLDEVV 57
>At2g06490 putative PttA-like transposon protein
Length = 491
Score = 28.5 bits (62), Expect = 1.3
Identities = 26/125 (20%), Positives = 49/125 (38%), Gaps = 17/125 (13%)
Query: 27 VIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQR 86
V+ S G + S S N PNSA ++ + P P + P Q + P+N +Q
Sbjct: 38 VLGSSGHRASSHSSN-------PNSATQSHRSRPSQYPPATVPQTTPPQTTLTPTNSRQP 90
Query: 87 F----------KQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDWLDPH 136
+ L +R + T ++ ++ + S+G+ + + +P+
Sbjct: 91 LLSGSQQPPPVHRNLPPTRQRQAPTSQQQPPTCQQKAPSRQHQSLGLQQQPRQTEQQNPN 150
Query: 137 CHEAE 141
HE E
Sbjct: 151 NHEEE 155
>At5g61260 putative protein
Length = 496
Score = 28.1 bits (61), Expect = 1.7
Identities = 32/110 (29%), Positives = 47/110 (42%), Gaps = 20/110 (18%)
Query: 30 SLGIQDSDQS---KNNAASVE-----SPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPS 81
S GI D S K +A +V+ S NS + +K+ +I G + S
Sbjct: 147 SPGITKPDSSVSAKRDALAVKKKPCASVNSESSSKEGSEIAKSVDGLS---------VKS 197
Query: 82 NRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKD 131
N + R K KE + +SG+ KV LR T VG +K +PK+
Sbjct: 198 NDRAR---KNKETESGLSGSAVVKKVPALRTDKSSTSTGVGSSKVCAPKN 244
>At3g44480 disease resistance protein -like
Length = 1220
Score = 28.1 bits (61), Expect = 1.7
Identities = 18/88 (20%), Positives = 40/88 (45%), Gaps = 5/88 (5%)
Query: 34 QDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKE 93
QD+ +S +++ S+ SP ++ + ++ HGA + ++ S R++ +
Sbjct: 73 QDNQESNSSSLSLPSPATSVSRNWKHDVFPSFHGADVRRTFLSHIMESFRRKGIDTFIDN 132
Query: 94 ADKRISGTGRENKVDNLRELVGGEKTSV 121
+R G E L+E + G K ++
Sbjct: 133 NIERSKSIGPE-----LKEAIKGSKIAI 155
>At1g12800 unknown protein
Length = 767
Score = 28.1 bits (61), Expect = 1.7
Identities = 21/103 (20%), Positives = 45/103 (43%), Gaps = 3/103 (2%)
Query: 32 GIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKL 91
G+ +S + +NN+ E ++ + +E+ I P PS+P Q ++ ++ + + Q+L
Sbjct: 313 GVLESSEIENNSIPTEMQLNSEMSSEEKTINSDPLERIPSKPISQTIVEASLQGK-PQRL 371
Query: 92 --KEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDW 132
A+ + G+ + V++ V E DW
Sbjct: 372 DPSSAEPSVPNIGKPSVVNHEGRQVSVELKGPPTRSSLEENDW 414
>At4g38780 splicing factor - like protein
Length = 2352
Score = 27.7 bits (60), Expect = 2.2
Identities = 28/98 (28%), Positives = 42/98 (42%), Gaps = 14/98 (14%)
Query: 36 SDQSKNNAASVESPNSAAKTKKEEK-IYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEA 94
SD +K N N++A T+ E + I LG PPSQ +QQ +++ KEA
Sbjct: 2008 SDYAKKNKV-----NTSALTQSEIRDIILGAEITPPSQQRQQIA-------EIEKQAKEA 2055
Query: 95 DKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDW 132
+ + T R V EL+ + + S DW
Sbjct: 2056 SQLTAVTTRTTNVHG-DELISTTISPYEQSAFGSKTDW 2092
>At3g63180 hypothetical protein
Length = 975
Score = 27.7 bits (60), Expect = 2.2
Identities = 17/71 (23%), Positives = 31/71 (42%)
Query: 39 SKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEADKRI 98
S +++ V SP+S+ + KE KI G P PK + + + + L K+
Sbjct: 124 SGSHSCEVVSPSSSTLSVKERKIVNGSKSRIPKPPKSSGTVEDDLEIEIAEVLSGLKKQP 183
Query: 99 SGTGRENKVDN 109
+ R + +N
Sbjct: 184 HSSKRGDDSEN 194
>At3g30640 hypothetical protein
Length = 661
Score = 27.7 bits (60), Expect = 2.2
Identities = 18/68 (26%), Positives = 33/68 (48%), Gaps = 2/68 (2%)
Query: 69 PPSQ--PKQQEVIPSNRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKG 126
PP++ PK + + + ++K KE + +G +NK + +E V G+ +SV K
Sbjct: 321 PPTKVPPKVLLKVTIKKPKSLEEKAKEKADCGTSSGPKNKPEEAKEKVDGDTSSVPKNKP 380
Query: 127 SSPKDWLD 134
K+ D
Sbjct: 381 KEAKEKAD 388
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,511,705
Number of Sequences: 26719
Number of extensions: 151607
Number of successful extensions: 701
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 45
Number of HSP's that attempted gapping in prelim test: 672
Number of HSP's gapped (non-prelim): 74
length of query: 144
length of database: 11,318,596
effective HSP length: 90
effective length of query: 54
effective length of database: 8,913,886
effective search space: 481349844
effective search space used: 481349844
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC122165.8


