

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122163.4 - phase: 0
(431 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
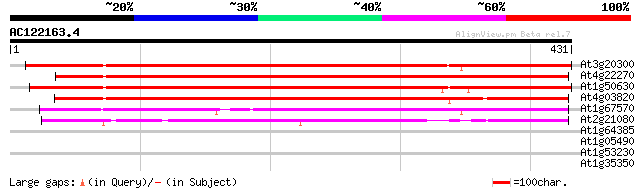
Score E
Sequences producing significant alignments: (bits) Value
At3g20300 unknown protein 458 e-129
At4g22270 unknown protein 443 e-124
At1g50630 unknown protein 428 e-120
At4g03820 unknown protein 405 e-113
At1g67570 unknown protein 208 4e-54
At2g21080 unknown protein 204 6e-53
At1g64385 unknown protein 30 3.4
At1g05490 hypothetical protein 29 5.7
At1g53230 flower development cycloidea like protein 28 9.8
At1g35350 hypothetical protein 28 9.8
>At3g20300 unknown protein
Length = 452
Score = 458 bits (1178), Expect = e-129
Identities = 235/424 (55%), Positives = 302/424 (70%), Gaps = 7/424 (1%)
Query: 13 NKENSGIYHDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCST 72
NK + H EL FR YLRW+ VDQS+ A +SWS+F VVP SHF+L CS
Sbjct: 30 NKFTRSVSHAQDELHSFRKYLRWMCVDQSSPWTAVLSWSMFVVFTLVVPATSHFMLACSD 89
Query: 73 TCDADHRRPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAI 132
CD+ H RPY VQ+SLS FA LSF+ LS++ KYG +FLF DK+ DES ++ GY
Sbjct: 90 -CDSHHSRPYDSVVQLSLSSFAALSFLCLSRFVSKYGLRRFLFFDKLWDESETVRLGYTN 148
Query: 133 QMKRTMKLILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRT 192
Q+ R++K++ + PCF+ YKIWWY SGASQIP+ G V +S + C +ELCSW YRT
Sbjct: 149 QLNRSLKILSYFVSPCFLAMSSYKIWWYASGASQIPFLGNVILSDTVACLMELCSWLYRT 208
Query: 193 SIFFLVCVLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFIL 252
++ FLVCVLFRLI +LQI RL +FA VFQ +++VG+IL EHLRIRR+LR+ISHR+R FIL
Sbjct: 209 TVIFLVCVLFRLICHLQILRLQDFAQVFQMDSDVGSILSEHLRIRRHLRIISHRYRTFIL 268
Query: 253 SSLLLVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSL 312
SL+LVT SQ LL+ K +A+++I + GEL L S+TLV+ L ILLRSA+KITHKAQ++
Sbjct: 269 LSLILVTGSQFYSLLITTKAYAELNIYRAGELALCSMTLVTALLILLRSASKITHKAQAV 328
Query: 313 TGLASKWHICATINSFDNIDGETPTALIASAQA---MAADISWGSSEDES-GDEEDELNN 368
T LA+KWH+CATI SF+ +DGETP L+ A D G S+ E GDEED+ +N
Sbjct: 329 TCLAAKWHVCATIESFETVDGETP-RLVDRASGHGYYPTDDDNGESDSEDYGDEEDDFDN 387
Query: 369 TKMMPIQT-RTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNK 427
++P TISF KRQALV Y ENNR+GITV+GF LDR+ LH+IFGI+++L LWLL K
Sbjct: 388 NNLIPAYAYSTISFQKRQALVNYFENNRSGITVFGFTLDRSTLHTIFGIEMSLVLWLLGK 447
Query: 428 TVGI 431
T+GI
Sbjct: 448 TIGI 451
>At4g22270 unknown protein
Length = 437
Score = 443 bits (1139), Expect = e-124
Identities = 216/395 (54%), Positives = 298/395 (74%), Gaps = 2/395 (0%)
Query: 36 VYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYHVPVQISLSVFAT 95
++ DQSN A +SWSVFF L +VP++SHFLL CS CD HRRPY V VQ+SLS+FA
Sbjct: 40 LWFDQSNFGTALLSWSVFFLLVVIVPLISHFLLVCSD-CDFHHRRPYDVIVQLSLSIFAG 98
Query: 96 LSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVY 155
+SF+SLS W RK+G +FLFLDK+ D S K++ Y +++R++K ++++ LP + Y
Sbjct: 99 ISFVSLSIWSRKFGMRRFLFLDKLWDVSDKVRIEYEAEIQRSLKRLMIFVLPSLTLEATY 158
Query: 156 KIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQRLDE 215
+IWWYISG +QIPY +S ++ CTL+L SW YR S+F +VC+L+++ +LQ RLD+
Sbjct: 159 RIWWYISGFNQIPYIINPILSHVVACTLQLSSWLYRNSLFIIVCILYKITCHLQTLRLDD 218
Query: 216 FAPVFQRE-TEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTASQLIFLLMVIKPHA 274
FA F E T+V + L EH +IRRNLR++SHRFR FIL SL+LVTA+Q + LL +
Sbjct: 219 FARCFASEITDVRSALGEHQKIRRNLRIVSHRFRRFILLSLILVTATQFMALLTTTRASV 278
Query: 275 DVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATINSFDNIDGE 334
V+I + GEL L S++LV+G++I LRSATKITHKAQS+T LA+KW++CAT++SFD++DGE
Sbjct: 279 AVNIYEVGELALCSLSLVTGVFICLRSATKITHKAQSVTSLAAKWNVCATVDSFDHLDGE 338
Query: 335 TPTALIASAQAMAADISWGSSEDESGDEEDELNNTKMMPIQTRTISFHKRQALVTYMENN 394
TPT I +Q + +S+DE G+ +D+L+NTK+ PI TIS+ KRQALVTY+ENN
Sbjct: 339 TPTGSIIESQVSLRGNAIETSDDEEGEGDDDLDNTKIHPIYANTISYQKRQALVTYLENN 398
Query: 395 RAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTV 429
+AGITVYGF++DR+WL++IFGI+LAL LWLLNKT+
Sbjct: 399 KAGITVYGFLVDRSWLNTIFGIELALLLWLLNKTI 433
>At1g50630 unknown protein
Length = 453
Score = 428 bits (1100), Expect = e-120
Identities = 222/428 (51%), Positives = 295/428 (68%), Gaps = 14/428 (3%)
Query: 16 NSGIYHDGKELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCD 75
N + H EL FR YLRW+ VD S+ A +SW++F VVP +SHFLL C+ CD
Sbjct: 27 NRCVSHQQDELHSFRKYLRWMCVDHSSPWTAILSWTMFIVFTLVVPAISHFLLACAD-CD 85
Query: 76 ADHRRPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMK 135
+ H RPY VQ+SLS AT+SF+ L+++ KYG +FLF DK+ DES ++R Y Q+
Sbjct: 86 SYHSRPYDSVVQLSLSSVATVSFLCLTRFVSKYGLRRFLFFDKLWDESETVRRNYTNQLN 145
Query: 136 RTMKLILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIF 195
++ ++ + +PCF YKIWWY SG S+IP+ G +S + C +ELCSW YRT++
Sbjct: 146 TSLHIVSYFVIPCFSAMSAYKIWWYASGGSRIPFLGNAVLSDTVACIMELCSWLYRTTVI 205
Query: 196 FLVCVLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSL 255
FLVCVLFRLI +LQI RL +FA +FQ +++VG+IL EHLRIRR+LR+ISHR+R+FIL L
Sbjct: 206 FLVCVLFRLICHLQILRLQDFAKLFQIDSDVGSILSEHLRIRRHLRIISHRYRSFILCLL 265
Query: 256 LLVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGL 315
+LVT SQ LL+ K + +V+I + GEL L S+TLV+ L ILLRSA+KITHKAQ++T L
Sbjct: 266 ILVTGSQFSSLLITTKAYTEVNIYRAGELALCSMTLVTALLILLRSASKITHKAQAVTCL 325
Query: 316 ASKWHICATINSFDNI------DGETPTALIASAQAMAADIS-----WGSSEDESGDEED 364
A+KWH+CAT+ SFD ETPT L+A ++ S DE GDEED
Sbjct: 326 AAKWHVCATLESFDQTVESFDQTVETPT-LVARNNNDNNNVHDVVTLTESDSDEYGDEED 384
Query: 365 ELNNTKMMPIQT-RTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLW 423
+L+N ++P+ T+SF KRQALV+Y ENN AGITVYGF LDR LH+IFG++L+L LW
Sbjct: 385 DLDNNDIIPVYAFSTMSFQKRQALVSYFENNSAGITVYGFTLDRGTLHTIFGLELSLVLW 444
Query: 424 LLNKTVGI 431
LL KT+GI
Sbjct: 445 LLGKTIGI 452
>At4g03820 unknown protein
Length = 437
Score = 405 bits (1040), Expect = e-113
Identities = 204/401 (50%), Positives = 286/401 (70%), Gaps = 9/401 (2%)
Query: 35 WVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYHVPVQISLSVFA 94
+++ DQSN K +SWS+FF LA +VP++SHF+L C+ CD HRRPY VQ+SLS+FA
Sbjct: 36 FLWFDQSNRIKTLLSWSIFFLLAVIVPMISHFVLICAD-CDFKHRRPYDGLVQLSLSIFA 94
Query: 95 TLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCV 154
+SF+SLS W +KYG +FLF DK+ D S K++ GY +++R+MKL+ ++ LP Q +
Sbjct: 95 GISFVSLSDWSKKYGIRRFLFFDKLKDVSDKVRIGYEAKIQRSMKLLAIFVLPSTTLQAI 154
Query: 155 YKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQRLD 214
Y+IWWY SG +QIPY +S ++ CTL+L SW YRTS+F + C+L++ I +LQ+ RLD
Sbjct: 155 YRIWWYASGFNQIPYIINPTLSHVLACTLQLSSWLYRTSLFIIACILYQNICHLQVLRLD 214
Query: 215 EFAPVFQRE-TEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTASQLIFLLMVIKPH 273
EFA F E + +IL EHL+IRR L+++SHRFR FIL SL VTA+Q + LL I+
Sbjct: 215 EFARCFASEIKDFSSILAEHLKIRRELKIVSHRFRRFILLSLFFVTATQFMALLTTIRAS 274
Query: 274 ADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATINSFDNI-D 332
+I + GEL L S +LVSGL+I L+SAT++THKAQS+T +A+KW++CA++++FD + D
Sbjct: 275 VPFNIYEVGELALCSTSLVSGLFICLKSATQMTHKAQSVTSIATKWNVCASLDTFDVLYD 334
Query: 333 GETP----TALIASAQAMAADISWGSSEDESGDEEDELNNTKMMPIQTRTISFHKRQALV 388
GETP T + + ++ S +DE G+ +D N+ ++ PI R IS KRQALV
Sbjct: 335 GETPKCPTTTQHSQILSRRRNVVQSSDDDEEGEGDD--NDLEIHPIFARAISSQKRQALV 392
Query: 389 TYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTV 429
TY+ENNRAGITVYGF++D+TWL IF I+LAL LWLL KT+
Sbjct: 393 TYLENNRAGITVYGFLVDKTWLRMIFSIELALLLWLLKKTI 433
>At1g67570 unknown protein
Length = 456
Score = 208 bits (530), Expect = 4e-54
Identities = 125/412 (30%), Positives = 225/412 (54%), Gaps = 15/412 (3%)
Query: 24 KELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYH 83
+ L +L + +QS+ +SW VF ++ V+P+ L C C+ + +
Sbjct: 49 RTLEWLETFLTLLGFNQSSKQSLVLSWIVFLSIGLVLPVTVLELGHC-LGCERYQYKSFE 107
Query: 84 VPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILL 143
+ + +S ++ A +S + +S RK+G KFLF+D++S +++ Y Q+ +++L+ +
Sbjct: 108 LNIVVSQALLAGVSLLCVSHNLRKHGIRKFLFVDQLSGRMDRLKAQYIQQILNSVRLLAV 167
Query: 144 WGLPCFICQCVYKI--WWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVL 201
W LPCF + V +I +Y+ + + ++S IL ++ L SW Y ++IF +
Sbjct: 168 WSLPCFALKGVREIIRMYYVP-------HDQPWLSVAILLSMIL-SWTYLSTIFLAASAM 219
Query: 202 FRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTAS 261
F L+ LQ+ +++A + + E+E+ + EH+R+R L ISHRFR F+L L+VTAS
Sbjct: 220 FHLVCNLQVIHFEDYAKLLEGESEISLFIYEHMRLRHYLSKISHRFRIFLLLQFLVVTAS 279
Query: 262 QLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHI 321
Q L + + GG+ + ++ V G+ + L +ATKI+H+AQ++ +AS+WH
Sbjct: 280 QFTTLFQTTAYSGRITYINGGDFAVSAVVQVVGIILCLHAATKISHRAQAIASVASRWHA 339
Query: 322 CATINSFDNID-GETPTALIASAQA---MAADISWGSSEDESGDEEDELNNTKMMPIQTR 377
+ +S D+ +P+ + A ++ IS S+ ES D + T P
Sbjct: 340 MMSCSSTDSTQIRASPSGVHLEATTNPPISFPISRSDSDVESMDHYMRMPVTNQFPSYMS 399
Query: 378 TISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTV 429
S+HKRQA V Y++ N GIT++G+ +DR +++IF I+L+L ++L KTV
Sbjct: 400 MSSYHKRQAFVLYLQMNPGGITIFGWTVDRHLINTIFFIELSLVTFVLGKTV 451
>At2g21080 unknown protein
Length = 414
Score = 204 bits (520), Expect = 6e-53
Identities = 124/413 (30%), Positives = 215/413 (52%), Gaps = 40/413 (9%)
Query: 25 ELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAYVVPILSHFLLDC-----STTCDADHR 79
+LR FR L+W +D S+ C +S+ +F +VP++S + S DA+
Sbjct: 31 DLRNFRLLLKWCALDHSSSCGKAVSYMMFVVFTLLVPLISCLFIKTPRNRPSAVMDANS- 89
Query: 80 RPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMK 139
++V VQ S A + F++L + R Y +K LFLD +S ++ GY+ ++ + ++
Sbjct: 90 --FNVLVQFPESGLAVIGFLTLICFFRIYSLTKLLFLD----DSTLVRLGYSRELDKALR 143
Query: 140 LILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVS-GIILCTLELCSWFYRTSIFFLV 198
+ +P F+ + V+K ++ S P+ + ++ L L SW YRT +F LV
Sbjct: 144 YLAYILVPSFLVELVHKSIFFYSAEVSFPFIKSSCAALNFVMFFLVLFSWVYRTGVFLLV 203
Query: 199 CVLFRLIGYLQIQRLDEFAPVFQR--ETEVGTILLEHLRIRRNLRVISHRFRAFILSSLL 256
C+LFRL LQI R +F R + + EH+RI++ L SHR+R FI+++ +
Sbjct: 204 CILFRLTCELQILRFRGLHKLFDRCGSDTIEDVCKEHVRIKKQLSATSHRYRFFIITAFV 263
Query: 257 LVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLA 316
+++ SQ + LL+V+ ++ L G+L + S +SG ++ L A +ITH+AQ + +A
Sbjct: 264 VISTSQFVALLLVLASKSEKSFLSSGDLVVCSAVQLSGFFLCLLGAARITHRAQGVVCIA 323
Query: 317 SKWHICATINSFDNIDGETPTALIASAQAMAADISWGSSEDESGDEEDELNNTKMMPIQT 376
++WH+ AL +++A+ S + D D + + P
Sbjct: 324 TRWHM----------------ALTCASEAV--------SPESDTDSSDNI-YINVSPSLD 358
Query: 377 RTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTV 429
+ F RQALV Y+ +N GIT+YG+ LDR LH++F + +L +W+L+K V
Sbjct: 359 LSSFFQARQALVEYLRHNNKGITLYGYALDRGLLHTLFAFEFSLVMWILSKVV 411
>At1g64385 unknown protein
Length = 351
Score = 29.6 bits (65), Expect = 3.4
Identities = 15/57 (26%), Positives = 29/57 (50%), Gaps = 8/57 (14%)
Query: 326 NSFDNIDGETPT---ALIASAQAMAADISWGSS-----EDESGDEEDELNNTKMMPI 374
N + +D E P AL+ + + D W ++ +DE+G ++E NT ++P+
Sbjct: 274 NKYQRLDMELPVSNPALVTKSDQESGDDGWNNNWGDDWDDENGGGDEEQPNTPVLPL 330
>At1g05490 hypothetical protein
Length = 731
Score = 28.9 bits (63), Expect = 5.7
Identities = 18/68 (26%), Positives = 37/68 (53%), Gaps = 7/68 (10%)
Query: 329 DNIDGETPTALIASAQAMAADISWGSSEDES------GDEEDELNNTKMMPIQTRTISFH 382
+++ E P++ +S+ + ++ S SS+DES GD D+ ++ + + +S
Sbjct: 276 ESLSSEDPSSSSSSSSSSSSSSSSSSSDDESYVKEVVGDNRDD-DDLRKASSPIKRVSLV 334
Query: 383 KRQALVTY 390
+R+ALV Y
Sbjct: 335 ERKALVRY 342
>At1g53230 flower development cycloidea like protein
Length = 391
Score = 28.1 bits (61), Expect = 9.8
Identities = 12/29 (41%), Positives = 15/29 (51%)
Query: 72 TTCDADHRRPYHVPVQISLSVFATLSFIS 100
TT D H PYH+P I S ++F S
Sbjct: 328 TTDDLHHHHPYHIPPGIHQSAIPGIAFAS 356
>At1g35350 hypothetical protein
Length = 747
Score = 28.1 bits (61), Expect = 9.8
Identities = 15/39 (38%), Positives = 21/39 (53%), Gaps = 3/39 (7%)
Query: 167 IPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLI 205
+P + V I +C + FYR+S FF + VLFR I
Sbjct: 471 VPLFVVALVIAISVCPFNI---FYRSSRFFFLMVLFRCI 506
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,389,257
Number of Sequences: 26719
Number of extensions: 383315
Number of successful extensions: 1212
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 1184
Number of HSP's gapped (non-prelim): 13
length of query: 431
length of database: 11,318,596
effective HSP length: 102
effective length of query: 329
effective length of database: 8,593,258
effective search space: 2827181882
effective search space used: 2827181882
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC122163.4


