

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.5 + phase: 1 /partial
(95 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
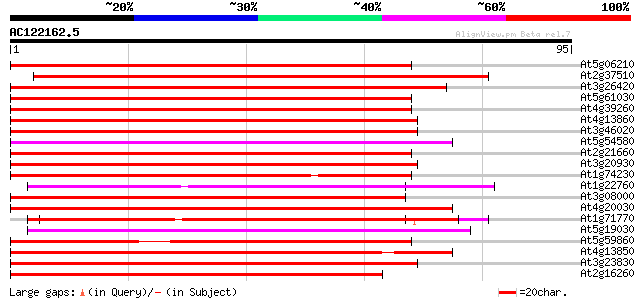
Score E
Sequences producing significant alignments: (bits) Value
At5g06210 unknown protein 76 3e-15
At2g37510 putative RNA-binding protein 70 2e-13
At3g26420 RNA-binding protein, putative 64 1e-11
At5g61030 RNA-binding protein - like 63 3e-11
At4g39260 glycine-rich protein (clone AtGRP8) 62 6e-11
At4g13860 RNA-binding protein like 62 6e-11
At3g46020 RNA binding protein -like 62 6e-11
At5g54580 unknown protein 60 2e-10
At2g21660 glycine-rich RNA binding protein 59 5e-10
At3g20930 unknown protein 58 9e-10
At1g74230 putative RNA-binding protein 58 9e-10
At1g22760 putative polyA-binding protein, PAB3 57 1e-09
At3g08000 RNA-binding protein like 57 2e-09
At4g20030 putative protein 56 4e-09
At1g71770 polyadenylate-binding protein 5 56 4e-09
At5g19030 unknown protein 55 8e-09
At5g59860 RNA-binding protein - like 54 1e-08
At4g13850 glycine-rich RNA-binding protein AtGRP2 - like 54 1e-08
At3g23830 glycine-rich RNA binding protein, putative 53 2e-08
At2g16260 putative glycine-rich RNA-binding protein 52 5e-08
>At5g06210 unknown protein
Length = 146
Score = 76.3 bits (186), Expect = 3e-15
Identities = 36/68 (52%), Positives = 52/68 (75%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+F TTE+ L+EAFS+ G VV+A IV+++ R KGFG+VTFA +EA+KA + NG+ L
Sbjct: 41 LSFCTTEQGLSEAFSKCGQVVEAQIVMDRVSDRSKGFGFVTFASADEAQKALMEFNGQQL 100
Query: 61 HGRVLYVD 68
+GR ++VD
Sbjct: 101 NGRTIFVD 108
>At2g37510 putative RNA-binding protein
Length = 142
Score = 70.1 bits (170), Expect = 2e-13
Identities = 33/77 (42%), Positives = 50/77 (64%)
Query: 5 TTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGRV 64
TT EKL +AF+ +G +V A ++ ++ R KGFG+VT+A E+A KA+ MN K L G V
Sbjct: 45 TTNEKLQDAFASFGQLVDARVITDRDSGRSKGFGFVTYATIEDAEKAKAEMNAKFLDGWV 104
Query: 65 LYVDMDPPNEQKNKIKQ 81
++VD P E + ++Q
Sbjct: 105 IFVDPARPREPRRPLQQ 121
>At3g26420 RNA-binding protein, putative
Length = 245
Score = 63.9 bits (154), Expect = 1e-11
Identities = 30/74 (40%), Positives = 50/74 (67%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T++ L +AF +YG++V+A +VL+K R +GFG++TF E++ +A MNG L
Sbjct: 14 LAWTTSDRGLRDAFEKYGHLVEAKVVLDKFSGRSRGFGFITFDEKKAMDEAIAAMNGMDL 73
Query: 61 HGRVLYVDMDPPNE 74
GR + VD P++
Sbjct: 74 DGRTITVDKAQPHQ 87
>At5g61030 RNA-binding protein - like
Length = 309
Score = 62.8 bits (151), Expect = 3e-11
Identities = 29/68 (42%), Positives = 45/68 (65%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
+A+S E+ L EAF++YG VV ++L++ R +GFG+VTF E A A ++G+ L
Sbjct: 47 MAYSMDEDSLREAFTKYGEVVDTRVILDRETGRSRGFGFVTFTSSEAASSAIQALDGRDL 106
Query: 61 HGRVLYVD 68
HGRV+ V+
Sbjct: 107 HGRVVKVN 114
>At4g39260 glycine-rich protein (clone AtGRP8)
Length = 169
Score = 61.6 bits (148), Expect = 6e-11
Identities = 30/68 (44%), Positives = 46/68 (67%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T +E L FSQ+G+V+ + I+ ++ R +GFG+VTF +E+ R A MNGK L
Sbjct: 13 LAWATNDEDLQRTFSQFGDVIDSKIINDRESGRSRGFGFVTFKDEKAMRDAIEEMNGKEL 72
Query: 61 HGRVLYVD 68
GRV+ V+
Sbjct: 73 DGRVITVN 80
>At4g13860 RNA-binding protein like
Length = 126
Score = 61.6 bits (148), Expect = 6e-11
Identities = 31/69 (44%), Positives = 47/69 (67%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+ +TT++ L EAFS YGNVV A ++ ++ R +GFG+VT++ EA A GM+GK L
Sbjct: 49 LSPTTTDDMLREAFSGYGNVVDAIVMRDRYTDRSRGFGFVTYSSHSEAEAAVSGMDGKEL 108
Query: 61 HGRVLYVDM 69
+GR + V +
Sbjct: 109 NGRRVSVKL 117
>At3g46020 RNA binding protein -like
Length = 102
Score = 61.6 bits (148), Expect = 6e-11
Identities = 26/69 (37%), Positives = 49/69 (70%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+ TT++ L + FS +G + +A ++ + +R KGFG++TF E++ARKA ++GKI+
Sbjct: 14 LSAYTTDQSLRQLFSPFGQIKEARLIRDSETQRPKGFGFITFDSEDDARKALKSLDGKIV 73
Query: 61 HGRVLYVDM 69
GR+++V++
Sbjct: 74 DGRLIFVEV 82
>At5g54580 unknown protein
Length = 156
Score = 59.7 bits (143), Expect = 2e-10
Identities = 30/75 (40%), Positives = 44/75 (58%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+ TT E L AF+Q+G V A +V ++ KGFG+V +A E++ K GM+GK L
Sbjct: 63 LSKRTTSEGLRTAFAQFGEVADAKVVTDRVSGYSKGFGFVRYATLEDSAKGIAGMDGKFL 122
Query: 61 HGRVLYVDMDPPNEQ 75
G V++ + P EQ
Sbjct: 123 DGWVIFAEYARPREQ 137
>At2g21660 glycine-rich RNA binding protein
Length = 176
Score = 58.5 bits (140), Expect = 5e-10
Identities = 28/68 (41%), Positives = 46/68 (67%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T + L AF+QYG+V+ + I+ ++ R +GFG+VTF +E+ + A GMNG+ L
Sbjct: 15 LAWATDDRALETAFAQYGDVIDSKIINDRETGRSRGFGFVTFKDEKAMKDAIEGMNGQDL 74
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 75 DGRSITVN 82
>At3g20930 unknown protein
Length = 374
Score = 57.8 bits (138), Expect = 9e-10
Identities = 27/69 (39%), Positives = 45/69 (65%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+F T+E+ L AF +G +V+ I+++K KR KG+ ++ + EE A A MNGKI+
Sbjct: 289 LSFYTSEKTLRAAFEGFGELVEVKIIMDKISKRSKGYAFLEYTTEEAAGTALKEMNGKII 348
Query: 61 HGRVLYVDM 69
+G ++ VD+
Sbjct: 349 NGWMIVVDV 357
>At1g74230 putative RNA-binding protein
Length = 289
Score = 57.8 bits (138), Expect = 9e-10
Identities = 30/68 (44%), Positives = 46/68 (67%), Gaps = 1/68 (1%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
+++ST E L EAFS+YG VV A I++++ R +GF +VTF EEA A + ++G+ L
Sbjct: 41 ISYSTDEFGLREAFSKYGEVVDAKIIVDRETGRSRGFAFVTFTSTEEASNA-MQLDGQDL 99
Query: 61 HGRVLYVD 68
HGR + V+
Sbjct: 100 HGRRIRVN 107
>At1g22760 putative polyA-binding protein, PAB3
Length = 660
Score = 57.4 bits (137), Expect = 1e-09
Identities = 30/79 (37%), Positives = 47/79 (58%), Gaps = 1/79 (1%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGR 63
S +EKL E FS+YGNV + ++LN + +GFG+V ++ EEA +A MNGK++ +
Sbjct: 342 SVDDEKLKEMFSEYGNVTSSKVMLNP-QGMSRGFGFVAYSNPEEALRALSEMNGKMIGRK 400
Query: 64 VLYVDMDPPNEQKNKIKQA 82
LY+ + E + QA
Sbjct: 401 PLYIALAQRKEDRRAHLQA 419
Score = 40.4 bits (93), Expect = 2e-04
Identities = 19/64 (29%), Positives = 37/64 (57%), Gaps = 1/64 (1%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGR 63
S + L E FS +G ++ + ++ R KG+G+V F +EE A+ A +NG +++ +
Sbjct: 146 SIDNKALFETFSSFGTILSCKVAMD-VTGRSKGYGFVQFEKEESAQAAIDKLNGMLMNDK 204
Query: 64 VLYV 67
++V
Sbjct: 205 QVFV 208
Score = 34.7 bits (78), Expect = 0.008
Identities = 21/61 (34%), Positives = 32/61 (52%), Gaps = 1/61 (1%)
Query: 7 EEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGRVLY 66
E++L + F ++G V+ + +V+ + FG+V F E A A MNG L VLY
Sbjct: 242 EDELRKTFGKFG-VISSAVVMRDQSGNSRCFGFVNFECTEAAASAVEKMNGISLGDDVLY 300
Query: 67 V 67
V
Sbjct: 301 V 301
>At3g08000 RNA-binding protein like
Length = 143
Score = 57.0 bits (136), Expect = 2e-09
Identities = 29/67 (43%), Positives = 42/67 (62%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L++S E+ L +AFS +G V + I +K R +GFG+V FAEE +A A+ M+GK L
Sbjct: 48 LSWSVDEQSLKDAFSSFGEVAEVRIAYDKGSGRSRGFGFVDFAEEGDALSAKDAMDGKGL 107
Query: 61 HGRVLYV 67
GR L +
Sbjct: 108 LGRPLRI 114
>At4g20030 putative protein
Length = 115
Score = 55.8 bits (133), Expect = 4e-09
Identities = 23/75 (30%), Positives = 48/75 (63%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L FST+E+ L FS +G + + ++ ++A KR KG+ ++ F +++A A M+ ++
Sbjct: 10 LPFSTSEDFLKREFSAFGEIAEVKLIKDEAMKRSKGYAFIQFTSQDDAFLAIETMDRRMY 69
Query: 61 HGRVLYVDMDPPNEQ 75
+GR++Y+D+ P ++
Sbjct: 70 NGRMIYIDIAKPGKR 84
>At1g71770 polyadenylate-binding protein 5
Length = 668
Score = 55.8 bits (133), Expect = 4e-09
Identities = 28/73 (38%), Positives = 45/73 (61%), Gaps = 1/73 (1%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGR 63
S +EKL E FS+YGNV +++N ++ +GFG+V ++ EEA A MNGK++ +
Sbjct: 338 SVNDEKLKEMFSEYGNVTSCKVMMN-SQGLSRGFGFVAYSNPEEALLAMKEMNGKMIGRK 396
Query: 64 VLYVDMDPPNEQK 76
LYV + E++
Sbjct: 397 PLYVALAQRKEER 409
Score = 40.4 bits (93), Expect = 2e-04
Identities = 20/64 (31%), Positives = 37/64 (57%), Gaps = 1/64 (1%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGR 63
S + L E FS +G ++ + ++ R KG+G+V F +EE A+ A +NG +L+ +
Sbjct: 142 SIDNKALYETFSSFGTILSCKVAMDVVG-RSKGYGFVQFEKEETAQAAIDKLNGMLLNDK 200
Query: 64 VLYV 67
++V
Sbjct: 201 QVFV 204
Score = 39.3 bits (90), Expect = 3e-04
Identities = 26/81 (32%), Positives = 39/81 (48%), Gaps = 6/81 (7%)
Query: 6 TEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGRVL 65
T+++L + F +YG++ A +V+ + FG+V F E A A MNG L VL
Sbjct: 237 TDDELKKTFGKYGDISSA-VVMKDQSGNSRSFGFVNFVSPEAAAVAVEKMNGISLGEDVL 295
Query: 66 YV-----DMDPPNEQKNKIKQ 81
YV D E + K +Q
Sbjct: 296 YVGRAQKKSDREEELRRKFEQ 316
Score = 30.4 bits (67), Expect = 0.16
Identities = 25/96 (26%), Positives = 40/96 (41%), Gaps = 6/96 (6%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHG- 62
S E L + F+Q V V R G+ YV FA E+A +A +N +
Sbjct: 55 SVNESHLLDLFNQVAPVHNLR-VCRDLTHRSLGYAYVNFANPEDASRAMESLNYAPIRDR 113
Query: 63 --RVLYVDMDPPNEQKNKIKQATKNTE--VDNEDVH 94
R++ + DP K KN + +DN+ ++
Sbjct: 114 PIRIMLSNRDPSTRLSGKGNVFIKNLDASIDNKALY 149
>At5g19030 unknown protein
Length = 172
Score = 54.7 bits (130), Expect = 8e-09
Identities = 25/75 (33%), Positives = 41/75 (54%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGR 63
S +E +L + FS++G V I+ N+ ++ G+GYV F +E+A+ A MNGK GR
Sbjct: 87 SVSEGRLKKVFSEFGQVTNVKIIANERTRQSLGYGYVWFNSKEDAQSAVEAMNGKFFDGR 146
Query: 64 VLYVDMDPPNEQKNK 78
+ V P + +
Sbjct: 147 FILVKFGQPGLSRRR 161
>At5g59860 RNA-binding protein - like
Length = 157
Score = 54.3 bits (129), Expect = 1e-08
Identities = 24/68 (35%), Positives = 47/68 (68%), Gaps = 5/68 (7%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+ TT++ L + F+ + ++K ++ +R KGFG++TF E++A+KA +NGKI+
Sbjct: 78 LSAYTTDQSLRQLFAPFARLIK-----DQQTQRPKGFGFITFESEDDAQKALKALNGKIV 132
Query: 61 HGRVLYVD 68
+GR+++V+
Sbjct: 133 NGRLIFVE 140
>At4g13850 glycine-rich RNA-binding protein AtGRP2 - like
Length = 158
Score = 53.9 bits (128), Expect = 1e-08
Identities = 26/75 (34%), Positives = 49/75 (64%), Gaps = 2/75 (2%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L++ T + L +AF+ +G+VV A +++++ R +GFG+V F +E A A M+GK L
Sbjct: 42 LSWGTDDASLRDAFAHFGDVVDAKVIVDRETGRSRGFGFVNFNDEGAATAAISEMDGKEL 101
Query: 61 HGRVLYVDMDPPNEQ 75
+GR ++ ++P N++
Sbjct: 102 NGR--HIRVNPANDR 114
>At3g23830 glycine-rich RNA binding protein, putative
Length = 136
Score = 53.1 bits (126), Expect = 2e-08
Identities = 24/69 (34%), Positives = 45/69 (64%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L++ T + L +AF+ +G V +A ++ ++ R +GFG+V+F+ E+ A A M+GK L
Sbjct: 42 LSWGTDDSSLKQAFTSFGEVTEATVIADRETGRSRGFGFVSFSCEDSANNAIKEMDGKEL 101
Query: 61 HGRVLYVDM 69
+GR + V++
Sbjct: 102 NGRQIRVNL 110
>At2g16260 putative glycine-rich RNA-binding protein
Length = 185
Score = 52.0 bits (123), Expect = 5e-08
Identities = 24/63 (38%), Positives = 40/63 (63%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T E+ + F+++G V + I++++ R KGF +VTF +E+ R A MNG+ L
Sbjct: 51 LAWATDEQSIERCFNEFGEVFDSKIIIDRETGRSKGFRFVTFKDEDSMRTAIDRMNGQEL 110
Query: 61 HGR 63
GR
Sbjct: 111 DGR 113
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.130 0.356
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,152,573
Number of Sequences: 26719
Number of extensions: 80756
Number of successful extensions: 422
Number of sequences better than 10.0: 170
Number of HSP's better than 10.0 without gapping: 152
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 173
Number of HSP's gapped (non-prelim): 255
length of query: 95
length of database: 11,318,596
effective HSP length: 71
effective length of query: 24
effective length of database: 9,421,547
effective search space: 226117128
effective search space used: 226117128
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC122162.5


