

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.2 - phase: 0 /pseudo
(547 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
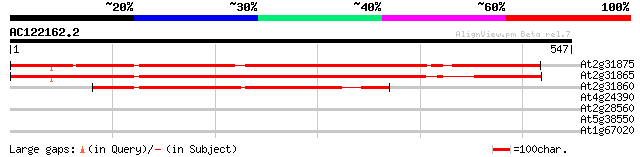
At2g31875 poly(ADP-ribose) like glycohydrolase 535 e-152
At2g31865 poly(ADP-ribose) like glycohydrolase 493 e-139
At2g31860 putative poly(ADP-ribose) glycohydrolase 293 2e-79
At4g24390 transport inhibitor response-like protein 32 0.69
At2g28560 putative RAD51B-like DNA repair protein 30 4.5
At5g38550 myrosinase binding protein-like; similar to jasmonate ... 29 5.9
At1g67020 hypothetical protein 29 5.9
>At2g31875 poly(ADP-ribose) like glycohydrolase
Length = 548
Score = 535 bits (1377), Expect = e-152
Identities = 287/521 (55%), Positives = 363/521 (69%), Gaps = 29/521 (5%)
Query: 1 MEKREDWRSILPYLPVVMRSSSLFWPSQVVGALRELGCG----RVDSGQLLFIFITELRN 56
ME RED SILPYLP+V+RSSSL+WP +VV AL+ + G +VDSG++L I ++R
Sbjct: 1 MENREDLNSILPYLPLVIRSSSLYWPPRVVEALKAMSEGPSHSQVDSGEVLRQAIFDMRR 60
Query: 57 SLSLSSEPLAPSAAHGYALFFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENAD 116
SLS S+ L PSA++GYA FDELI +E ++WF+E++P+L LLL+ PSLLE H++NAD
Sbjct: 61 SLSFST--LEPSASNGYAFLFDELIDEKESKRWFDEIIPALASLLLQFPSLLEVHFQNAD 118
Query: 117 MVIDGKGATVRTGLRMLDSQEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFD 176
++ G ++TGLR+L+SQ+AGIVFL+QELI ALLACSF CLFP +R K L +NFD
Sbjct: 119 NIVSG----IKTGLRLLNSQQAGIVFLSQELIGALLACSFFCLFPDDNRGAKHLPVINFD 174
Query: 177 ELFASLYNDYTQKQEDKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASF 236
LFASLY Y+Q QE KI CI+HYF+R S +P G+VSFERK+ + P+A F
Sbjct: 175 HLFASLYISYSQSQESKIRCIMHYFERFCSCVPIGIVSFERKIT---------AAPDADF 225
Query: 237 WSTSVKPLCRFEVKSSGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIA 296
WS S LC F+V S GLIED +EVDFAN+YLGGG+L RGCVQEEIRFMI+PELIA
Sbjct: 226 WSKSDVSLCAFKVHSFGLIEDQPDNALEVDFANKYLGGGSLSRGCVQEEIRFMINPELIA 285
Query: 297 GMIFLPSMADNEAIDIVGVERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDAL 356
GM+FLP M DNEAI+IVG ERFS YTGYASSFRF+G+Y+D K +DPF RR+TRIVAIDAL
Sbjct: 286 GMLFLPRMDDNEAIEIVGAERFSCYTGYASSFRFAGEYIDKKAMDPFKRRRTRIVAIDAL 345
Query: 357 CGPGMRQYREKFLLREINKAFCGFLQQSQYQRDQKIPQENVCSSLLVVFSMTF*HQYGLL 416
C P MR +++ LLREINKA CGFL S+ Q I + + + +V + GLL
Sbjct: 346 CTPKMRHFKDICLLREINKALCGFLNCSKAWEHQNIFMDEGDNEIQLVRNG---RDSGLL 402
Query: 417 SASLVVQLSVLNIFRFS*FDAMETSEGKYSYQEIRNSQNDYDMMENSNDIGVATGNWGCG 476
+ + +E + K + IR+ + E+ D GVATGNWGCG
Sbjct: 403 RTETTAS-------HRTPLNDVEMNREKPANNLIRDFYVEGVDNEDHEDDGVATGNWGCG 455
Query: 477 AFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGALQNLDKI 517
FGGDPE+K IQWLAASQ RPFI+YYTFG AL+NLD++
Sbjct: 456 VFGGDPELKATIQWLAASQTRRPFISYYTFGVEALRNLDQV 496
>At2g31865 poly(ADP-ribose) like glycohydrolase
Length = 522
Score = 493 bits (1269), Expect = e-139
Identities = 271/522 (51%), Positives = 341/522 (64%), Gaps = 47/522 (9%)
Query: 1 MEKREDWRSILPYLPVVMRSSSLFWPSQVVGALRELGCG----RVDSGQLLFIFITELRN 56
ME R D RSIL YLP+V +SSSL WP V L+ + G V+SG+ L + IT +R
Sbjct: 1 MELRADLRSILQYLPLVAQSSSLVWPPSVEEELQTISRGPSESMVNSGEALALHITNMRK 60
Query: 57 SLSLSSEPLAPSAAHGYALFFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENAD 116
SLSL++ LAP A GY LFFD+ ISREE +F EV+P+L LLL+LPS+LE HY+ AD
Sbjct: 61 SLSLNASDLAPYALQGYGLFFDKKISREESANFFGEVVPALCRLLLQLPSMLEKHYQKAD 120
Query: 117 MVIDGKGATVRTGLRMLDSQEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFD 176
V+DG V++GLR+L QEAGIV L+QELIAALLACSF CLFP DR K LQ +NF
Sbjct: 121 HVLDG----VKSGLRLLGPQEAGIVLLSQELIAALLACSFFCLFPEVDRSLKNLQGINFS 176
Query: 177 ELFASLYNDYTQKQEDKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASF 236
LF+ Y + KQE+KI C+IHYF RI MP G VSFERK+LP E +SYP A
Sbjct: 177 GLFSFPYMRHCTKQENKIKCLIHYFGRICRWMPTGFVSFERKILPLEYHPHFVSYPKADS 236
Query: 237 WSTSVKPLCRFEVKSSGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIA 296
W+ SV PLC E+ +SG IED E +EVDFA+EY GG L +QEEIRF+I+PELIA
Sbjct: 237 WANSVTPLCSIEIHTSGAIEDQPCEALEVDFADEYFGGLTLSYDTLQEEIRFVINPELIA 296
Query: 297 GMIFLPSMADNEAIDIVGVERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDAL 356
GMIFLP M NEAI+IVGVERFS YTGY SF+++GDY D+K++D F RRKTR++AIDA+
Sbjct: 297 GMIFLPRMDANEAIEIVGVERFSGYTGYGPSFQYAGDYTDNKDLDIFRRRKTRVIAIDAM 356
Query: 357 CGPGMRQYREKFLLREINKAFCGFLQQSQYQRDQKIPQENVCSSLLVVFSMTF*HQYGLL 416
PGM QY+ L+RE+NKAF G++ Q +Y D K E S + +
Sbjct: 357 PDPGMGQYKLDALIREVNKAFSGYMHQCKYNIDVKHDPEASSSHVPLTSD---------- 406
Query: 417 SASLVVQLSVLNIFRFS*FDAMETSEGKYSYQEIRNSQNDYDMMENSNDIGVATGNWGCG 476
SAS V++ S + + + IGVATGNWGCG
Sbjct: 407 SASQVIE-----------------------------SSHRWCIDHEEKKIGVATGNWGCG 437
Query: 477 AFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGALQNLDKIL 518
FGGDPE+K ++QWLA SQ+GRPF++YYTFG ALQNL++++
Sbjct: 438 VFGGDPELKIMLQWLAISQSGRPFMSYYTFGLQALQNLNQVI 479
>At2g31860 putative poly(ADP-ribose) glycohydrolase
Length = 364
Score = 293 bits (749), Expect = 2e-79
Identities = 162/291 (55%), Positives = 200/291 (68%), Gaps = 26/291 (8%)
Query: 81 ISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDSQEAGI 140
+S+EE +WF E LP++ LLLR PSLLE+HY N+D +I+G +TGLR+L +AGI
Sbjct: 1 MSKEESSRWFNEFLPAMACLLLRFPSLLESHYLNSDNLING----TKTGLRVLVPNKAGI 56
Query: 141 VFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYND-YTQKQEDKIWCIIH 199
VFL+QELI ALL+CSF CLFPV DR L +NFD+LF SL N + QE+KI CIIH
Sbjct: 57 VFLSQELIGALLSCSFFCLFPVDDRGSNHLPIINFDKLFGSLINTGRNEHQENKIKCIIH 116
Query: 200 YFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLIEDHS 259
YFQR++S++ G VSFERK+L E D S + FW S LC EV++SGLIED S
Sbjct: 117 YFQRLSSSISPGFVSFERKILSLEQDS---STLDEGFWGKSTVNLCPVEVRTSGLIEDQS 173
Query: 260 SETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGVERFS 319
E +EVDFAN+ LGGGALR+GCVQEEIRFMI+PELI GM+FLP+M EAI++VG ERFS
Sbjct: 174 VEALEVDFANKNLGGGALRKGCVQEEIRFMINPELIVGMLFLPTMEVTEAIEVVGAERFS 233
Query: 320 SYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKFLL 370
YTG F + KTRIVAIDAL PG+ QY+ + LL
Sbjct: 234 LYTGC------------------FRKAKTRIVAIDALRHPGVSQYKLESLL 266
Score = 37.7 bits (86), Expect = 0.017
Identities = 17/34 (50%), Positives = 25/34 (73%), Gaps = 3/34 (8%)
Query: 487 IIQWLA---ASQAGRPFIAYYTFGSGALQNLDKI 517
++ WL + QA RPF++YYTFG ALQNL+++
Sbjct: 331 VVVWLTTLHSFQARRPFMSYYTFGFEALQNLNQV 364
>At4g24390 transport inhibitor response-like protein
Length = 623
Score = 32.3 bits (72), Expect = 0.69
Identities = 43/175 (24%), Positives = 66/175 (37%), Gaps = 15/175 (8%)
Query: 281 CVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGVERFSSYTGYASSF---RFSGDYVDD 337
CV+ I F EL+ FL + N + + + R +S FS D V
Sbjct: 238 CVESPINFKALEELVVRSPFLKKLRTNRFVSLEELHRLMVRAPQLTSLGTGSFSPDNVPQ 297
Query: 338 KEVDPFGRRKTRIVAIDALCGPGMRQYREKFLLREINKAFCGFLQQSQYQRDQKIPQE-- 395
E P R +C G R++R ++LL + C L + P
Sbjct: 298 GEQQPDYAAAFR-ACKSIVCLSGFREFRPEYLL--AISSVCANLTSLNFSYANISPHMLK 354
Query: 396 ---NVCSSLLVVFSMTF*HQYGLLS-ASLVVQLSVLNIFRFS*FDAMETSEGKYS 446
+ C ++ V +++ GL + A+ +L L IF FD E SEG S
Sbjct: 355 PIISNCHNIRVFWALDSIRDEGLQAVAATCKELRELRIFP---FDPREDSEGPVS 406
>At2g28560 putative RAD51B-like DNA repair protein
Length = 353
Score = 29.6 bits (65), Expect = 4.5
Identities = 20/81 (24%), Positives = 36/81 (43%)
Query: 21 SSLFWPSQVVGALRELGCGRVDSGQLLFIFITELRNSLSLSSEPLAPSAAHGYALFFDEL 80
S++F ++ A L + +LL + + E+R+++S SE +P +LFF +
Sbjct: 16 SNIFAARNIITAKDALSMTEFELMELLDVGMKEIRSAISFISEATSPPCQSVSSLFFFKK 75
Query: 81 ISREECRKWFEEVLPSLGDLL 101
+ E L L D L
Sbjct: 76 VENEHLSGHLPTHLKGLDDTL 96
>At5g38550 myrosinase binding protein-like; similar to jasmonate
induced protein
Length = 594
Score = 29.3 bits (64), Expect = 5.9
Identities = 25/80 (31%), Positives = 35/80 (43%), Gaps = 9/80 (11%)
Query: 200 YFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLIE--- 256
YF T N K +RK LPW+D C + S S RFE + G +E
Sbjct: 290 YFSTTTPN--KLECQGDRKGLPWDDGCNYDGVKKVYVDSISDIDSVRFEYDNGGKVEKTP 347
Query: 257 ---DHSSETVEV-DFANEYL 272
D ++E V D+ NE++
Sbjct: 348 YRRDVTNEKEFVLDYPNEFI 367
>At1g67020 hypothetical protein
Length = 659
Score = 29.3 bits (64), Expect = 5.9
Identities = 28/92 (30%), Positives = 40/92 (43%), Gaps = 14/92 (15%)
Query: 180 ASLYNDYTQKQEDKIWCII--HYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASF- 236
A+L N +++ K W I HY R+ N+ G+ +R PW C+ N
Sbjct: 424 ATLMNSALKRKCSKSWHFIYKHYKVRM-QNLHTGIYKTQR---PWH--CLVHEANNTDSH 477
Query: 237 ----WSTSVKPLCRF-EVKSSGLIEDHSSETV 263
W T V+ L E S GL+ HSSE +
Sbjct: 478 LWHRWKTKVETLQEISEFLSWGLVHSHSSEVI 509
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,134,971
Number of Sequences: 26719
Number of extensions: 522180
Number of successful extensions: 1494
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 1475
Number of HSP's gapped (non-prelim): 11
length of query: 547
length of database: 11,318,596
effective HSP length: 104
effective length of query: 443
effective length of database: 8,539,820
effective search space: 3783140260
effective search space used: 3783140260
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC122162.2


