

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121245.9 + phase: 2 /pseudo/partial
(223 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
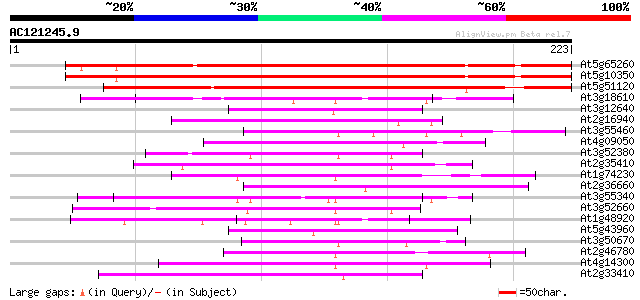
Score E
Sequences producing significant alignments: (bits) Value
At5g65260 poly(A)-binding protein II-like 293 4e-80
At5g10350 RNA binding protein - like 286 8e-78
At5g51120 putative polyA-binding protein II 258 2e-69
At3g18610 unknown protein 60 1e-09
At3g12640 hypothetical protein 59 2e-09
At2g16940 splicing factor like protein 57 1e-08
At3g55460 RNA binding protein like 55 4e-08
At4g09050 unknown protein 52 2e-07
At3g52380 RNA-binding protein cp33 precursor 51 4e-07
At2g35410 putative chloroplast RNA binding protein precursor 50 9e-07
At1g74230 putative RNA-binding protein 50 9e-07
At2g36660 putative poly(A) binding protein 50 1e-06
At3g55340 putative protein 49 2e-06
At3g52660 putative RNA binding protein 49 3e-06
At1g48920 unknown protein 49 3e-06
At5g43960 unknown protein 48 5e-06
At3g50670 U1 snRNP 70K protein 48 5e-06
At2g46780 putative RNA-binding protein 48 5e-06
At4g14300 ribonucleoprotein like protein 47 8e-06
At2g33410 putative RNA-binding protein 47 8e-06
>At5g65260 poly(A)-binding protein II-like
Length = 220
Score = 293 bits (751), Expect = 4e-80
Identities = 152/204 (74%), Positives = 173/204 (84%), Gaps = 7/204 (3%)
Query: 23 D*DME--NDDVDMIGADNNEA-DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPAN 79
D DME N D+DM AD++ +LD+MKKRLKEMEDEAAAL+EMQAKVEKEMG+ QDPA+
Sbjct: 20 DGDMEALNPDLDMAAADDDAVKELDEMKKRLKEMEDEAAALREMQAKVEKEMGA-QDPAS 78
Query: 80 ASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVE 139
+A+Q +EEVDARS+FVGNVDYACTPE+VQQHFQ+CGTV+RVTI TDKFGQPKG+AYVE
Sbjct: 79 IAANQAGKEEVDARSVFVGNVDYACTPEEVQQHFQTCGTVHRVTILTDKFGQPKGFAYVE 138
Query: 140 FVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPF 199
FVEVEAVQEAL LNESELHGRQLKV KRTNVPG+KQFR RR NPYMG+R R P+ P+
Sbjct: 139 FVEVEAVQEALQLNESELHGRQLKVLQKRTNVPGLKQFRGRR-FNPYMGYRFRRPFMSPY 197
Query: 200 AYAPAPYGYGKIPRFRMGMRYSPY 223
Y +PYGYGK PRFR MRY PY
Sbjct: 198 MY--SPYGYGKAPRFRRPMRYMPY 219
>At5g10350 RNA binding protein - like
Length = 217
Score = 286 bits (731), Expect = 8e-78
Identities = 143/202 (70%), Positives = 168/202 (82%), Gaps = 4/202 (1%)
Query: 23 D*DMENDDVDMIGADNNEA-DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANAS 81
D D+ + D+DM AD + +L +MK+RLKEME+EAAAL+EMQAKVEKEMG+ QDPA+ +
Sbjct: 18 DTDVPDPDIDMSAADEDAVTELAEMKRRLKEMEEEAAALREMQAKVEKEMGATQDPASMA 77
Query: 82 ASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFV 141
A+Q +EEVDARS++VGNVDYACTPE+VQ HFQ+CGTVNRVTI DKFGQPKG+AYVEFV
Sbjct: 78 ANQEGKEEVDARSVYVGNVDYACTPEEVQLHFQTCGTVNRVTILMDKFGQPKGFAYVEFV 137
Query: 142 EVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPFAY 201
EVEAVQEAL LNESELHGRQLKV+ KRTNVPGMKQ+ P R NP MG+R R P+ PP+ Y
Sbjct: 138 EVEAVQEALQLNESELHGRQLKVSPKRTNVPGMKQYHPGR-FNPSMGYRFRRPFVPPYFY 196
Query: 202 APAPYGYGKIPRFRMGMRYSPY 223
+PYGYGK PRFR MRY PY
Sbjct: 197 --SPYGYGKAPRFRRPMRYMPY 216
>At5g51120 putative polyA-binding protein II
Length = 227
Score = 258 bits (659), Expect = 2e-69
Identities = 128/187 (68%), Positives = 151/187 (80%), Gaps = 9/187 (4%)
Query: 38 NNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFV 97
++ DL+DMKKR+KE+E+EA AL+EMQAK EK+MG+ QDP+ S +EEVD+RSI+V
Sbjct: 49 SSSRDLEDMKKRIKEIEEEAGALREMQAKAEKDMGASQDPSGG-VSAAEKEEVDSRSIYV 107
Query: 98 GNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEALLLNESEL 157
GNVDYACTPE+VQQHFQSCGTVNRVTI TDKFGQPKG+AYVEFVEVEAVQ +L+LNESEL
Sbjct: 108 GNVDYACTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAYVEFVEVEAVQNSLILNESEL 167
Query: 158 HGRQLKVTAKRTNVPGMKQFRPR-RPSNPYMGFRGRTPYAPPFAYAPAPYGYGKIPRFRM 216
HGRQ+KV+AKRTNVPGM+QFR R RP P GF P+ P PY YG++PRFR
Sbjct: 168 HGRQIKVSAKRTNVPGMRQFRGRGRPFRPMRGFMPGVPFYP-------PYAYGRVPRFRR 220
Query: 217 GMRYSPY 223
MRY PY
Sbjct: 221 PMRYRPY 227
>At3g18610 unknown protein
Length = 636
Score = 59.7 bits (143), Expect = 1e-09
Identities = 40/151 (26%), Positives = 67/151 (43%), Gaps = 6/151 (3%)
Query: 51 KEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQ 110
+E +DE K+ + VE + A + N+ + ++++F GN+ Y D++
Sbjct: 342 EESKDEKVTPKKKDSDVEMVDAEQKSNAKQPKTPTNQTQGGSKTLFAGNLSYQIARSDIE 401
Query: 111 QHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEALLLNESELHGRQLKV-TAKRT 169
F+ G V V + + G KGY ++EF E Q+AL +N L GR +++ A
Sbjct: 402 NFFKEAGEVVDVRLSSFDDGSFKGYGHIEFASPEEAQKALEMNGKLLLGRDVRLDLANER 461
Query: 170 NVPGMKQFRPRRPSNPYMGFRGRTPYAPPFA 200
P R P G + RT Y F+
Sbjct: 462 GTP-----RNSNPGRKGEGSQSRTIYVRGFS 487
Score = 44.7 bits (104), Expect = 4e-05
Identities = 36/145 (24%), Positives = 68/145 (46%), Gaps = 11/145 (7%)
Query: 29 DDVDMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINRE 88
DD G + E + ++ EM + ++++ + E G+ P N++ + E
Sbjct: 419 DDGSFKGYGHIEFASPEEAQKALEMNGKLLLGRDVRLDLANERGT---PRNSNPGR-KGE 474
Query: 89 EVDARSIFVGNVDYACTPEDVQQ----HFQSCGTVNRVTIRTDK-FGQPKGYAYVEFVEV 143
+R+I+V + +++++ HF CG V RV + TD+ G +G+AY++
Sbjct: 475 GSQSRTIYVRGFSSSLGEDEIKKELRSHFSKCGEVTRVHVPTDRETGASRGFAYIDL--T 532
Query: 144 EAVQEALLLNESELHGRQLKVTAKR 168
EAL L+ SE+ G + V R
Sbjct: 533 SGFDEALQLSGSEIGGGNIHVEESR 557
>At3g12640 hypothetical protein
Length = 674
Score = 58.9 bits (141), Expect = 2e-09
Identities = 31/78 (39%), Positives = 44/78 (55%), Gaps = 1/78 (1%)
Query: 88 EEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTD-KFGQPKGYAYVEFVEVEAV 146
E+ +R+IFV NV + T + + +HF G V + I TD GQP G AY+EF EA
Sbjct: 511 EDASSRTIFVANVHFGATKDSLSRHFNKFGEVLKAFIVTDPATGQPSGSAYIEFTRKEAA 570
Query: 147 QEALLLNESELHGRQLKV 164
+ AL L+ + R LK+
Sbjct: 571 ENALSLDGTSFMSRILKI 588
>At2g16940 splicing factor like protein
Length = 561
Score = 56.6 bits (135), Expect = 1e-08
Identities = 34/111 (30%), Positives = 61/111 (54%), Gaps = 3/111 (2%)
Query: 65 AKVEKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTI 124
++ EK + A + + AR ++VGN+ + +D+++ F+S G+V V +
Sbjct: 257 SEAEKNLVQSTTAAAGAGGMLGPYSGGARRLYVGNLHINMSEDDLRKVFESFGSVELVQV 316
Query: 125 RTDKFGQPKGYAYVEFVEVEAVQEALLLN-ESELHGRQLKVTA--KRTNVP 172
D+ G KG+ +V+F +E + AL LN + E+ GR +KV+A +T VP
Sbjct: 317 PRDETGLCKGFGFVQFARLEDARNALNLNGQLEIAGRAIKVSAVTDQTEVP 367
>At3g55460 RNA binding protein like
Length = 262
Score = 54.7 bits (130), Expect = 4e-08
Identities = 46/161 (28%), Positives = 69/161 (42%), Gaps = 40/161 (24%)
Query: 94 SIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEV-EAVQEALL 151
S+ V N+ C PE++++ F+ G V V I D + GQP+G+A+VEFV+ +A +
Sbjct: 48 SLLVRNIPLDCRPEELREPFERFGPVRDVYIPRDYYSGQPRGFAFVEFVDAYDAGEAQRS 107
Query: 152 LNESELHGRQLKV----------------TAKRTNVPGMKQFR---------------PR 180
+N GR++ V T R+ P + R PR
Sbjct: 108 MNRRSFAGREITVVVASESRKRPEEMRVKTRTRSREPSGSRDRSHGRSRSRSISRSRSPR 167
Query: 181 RPSNPYMGFRGRTPYAPPFAYAPAPYGYGKIPRFRMGMRYS 221
RPS+ +R R +Y+PAP G PR YS
Sbjct: 168 RPSDSRSRYRSR-------SYSPAPRRRGGPPRGEEDENYS 201
>At4g09050 unknown protein
Length = 304
Score = 52.4 bits (124), Expect = 2e-07
Identities = 33/113 (29%), Positives = 55/113 (48%), Gaps = 3/113 (2%)
Query: 78 ANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAY 137
+++SA ++ EE+ + NV + TPED++ F+ G+V + + K + +G +
Sbjct: 79 SSSSAPEVVEEEISKTRLIAQNVPWTSTPEDIRSLFEKYGSVIDIEMSMHKKERNRGLVF 138
Query: 138 VEFVEVEAVQEALLLNES-ELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGF 189
+E E AL ES E GR+LKV +T K + PR +P F
Sbjct: 139 IEMASPEEAATALKSLESCEYEGRRLKVDYAKTK--KKKTYAPRETPSPVPTF 189
>At3g52380 RNA-binding protein cp33 precursor
Length = 329
Score = 51.2 bits (121), Expect = 4e-07
Identities = 34/118 (28%), Positives = 63/118 (52%), Gaps = 9/118 (7%)
Query: 55 DEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARS------IFVGNVDYACTPED 108
+ ++A E+QA VE+E V++ + ++ E+ ++ ++VGN+ Y T +
Sbjct: 73 EASSADDEIQASVEEEE-EVEEEGDEGEEEVEEEKQTTQASGEEGRLYVGNLPYTITSSE 131
Query: 109 VQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEAL-LLNESELHGRQLKV 164
+ Q F GTV V I DK + +G+ +V +E +EA+ + N S++ GR +KV
Sbjct: 132 LSQIFGEAGTVVDVQIVYDKVTDRSRGFGFVTMGSIEEAKEAMQMFNSSQIGGRTVKV 189
>At2g35410 putative chloroplast RNA binding protein precursor
Length = 308
Score = 50.1 bits (118), Expect = 9e-07
Identities = 35/137 (25%), Positives = 66/137 (47%), Gaps = 6/137 (4%)
Query: 50 LKEMEDEAAALKEMQAKV-EKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPED 108
L+ D + L + V EKE + ++ + ++ + R +FV N+ ++ + D
Sbjct: 51 LRSGSDNSRRLVSVLCSVAEKETSADEETSQEEKTEETQNSNLKRKLFVFNLPWSMSVND 110
Query: 109 VQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL-LLNESELHGRQLKVTAK 167
+ + F CGTVN V I K G+ +G+A+V E Q A+ + ++ GR + V+
Sbjct: 111 ISELFGQCGTVNNVEIIRQKDGKNRGFAFVTMASGEEAQAAIDKFDTFQVSGRIISVSFA 170
Query: 168 RTNVPGMKQFRPRRPSN 184
R K+ P+ P++
Sbjct: 171 RR----FKKPTPKSPND 183
>At1g74230 putative RNA-binding protein
Length = 289
Score = 50.1 bits (118), Expect = 9e-07
Identities = 39/147 (26%), Positives = 63/147 (42%), Gaps = 17/147 (11%)
Query: 65 AKVEKEMGSVQDPANASASQINREE-VDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVT 123
+KV + AS+S + + + IFVG + Y+ +++ F G V
Sbjct: 5 SKVGRLFSQTSSHVTASSSMLQSIRCMSSSKIFVGGISYSTDEFGLREAFSKYGEVVDAK 64
Query: 124 IRTDK-FGQPKGYAYVEFVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRP 182
I D+ G+ +G+A+V F E A+ L+ +LHGR+++V + R
Sbjct: 65 IIVDRETGRSRGFAFVTFTSTEEASNAMQLDGQDLHGRRIRV-----------NYATERG 113
Query: 183 SNPYMGFRGRTPYAPPFAYAPAPYGYG 209
S GF GR P Y + GYG
Sbjct: 114 S----GFGGRGFGGPGGGYGASDGGYG 136
>At2g36660 putative poly(A) binding protein
Length = 609
Score = 49.7 bits (117), Expect = 1e-06
Identities = 30/114 (26%), Positives = 51/114 (44%), Gaps = 1/114 (0%)
Query: 94 SIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEF-VEVEAVQEALLL 152
+I+V NV+ A T E++++HF CGT+ + D+ G+ KG+ +V F EA+
Sbjct: 305 NIYVKNVNVAVTEEELRKHFSQCGTITSTKLMCDEKGKSKGFGFVCFSTPEEAIDAVKTF 364
Query: 153 NESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPFAYAPAPY 206
+ HG+ L V + Q + + + + P YAP Y
Sbjct: 365 HGQMFHGKPLYVAIAQKKEDRKMQLQVQFGNRVEARKSSSSASVNPGTYAPLYY 418
Score = 34.3 bits (77), Expect = 0.054
Identities = 24/80 (30%), Positives = 40/80 (50%), Gaps = 6/80 (7%)
Query: 91 DARSIFVGNVDYACTPEDV-----QQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEA 145
DAR VGNV PE V Q F+ G + + T + G+ +GY +V+F + +A
Sbjct: 105 DARRNGVGNVFVKNLPESVTNAVLQDMFKKFGNIVSCKVATLEDGKSRGYGFVQFEQEDA 164
Query: 146 VQEAL-LLNESELHGRQLKV 164
A+ LN + + +++ V
Sbjct: 165 AHAAIQTLNSTIVADKEIYV 184
Score = 27.3 bits (59), Expect = 6.6
Identities = 26/115 (22%), Positives = 51/115 (43%), Gaps = 7/115 (6%)
Query: 41 ADLDDMKKR-----LKEMEDEA-AALKEMQAKVEKEMGSVQDPANASASQINREEVDARS 94
A L+D K R E ED A AA++ + + + + ++ EE +
Sbjct: 144 ATLEDGKSRGYGFVQFEQEDAAHAAIQTLNSTIVADKEIYVGKFMKKTDRVKPEE-KYTN 202
Query: 95 IFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEA 149
+++ N+D + + +++ F G + + I D+ +GYA+V F E + A
Sbjct: 203 LYMKNLDADVSEDLLREKFAEFGKIVSLAIAKDENRLCRGYAFVNFDNPEDARRA 257
>At3g55340 putative protein
Length = 597
Score = 49.3 bits (116), Expect = 2e-06
Identities = 33/164 (20%), Positives = 77/164 (46%), Gaps = 11/164 (6%)
Query: 28 NDDVDMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINR 87
+D V++ + +E ++ K + K+ + + K + + E G+V++ ++N
Sbjct: 91 DDAVEVDELEGDEGTKEEQKPQKKKNKKKKKKRKVNKTPKKAEEGNVEEKVKVEEIEVNT 150
Query: 88 EE-----VDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIR-TDKFGQPKGYAYVEFV 141
+ V ++VG + Y T ++++ +F+SCG + +V + + G G A++ F
Sbjct: 151 DNKEEDGVVPNKLYVGGIPYQSTEDEIRSYFRSCGVIIKVDCKMRPEDGAFSGIAFITFD 210
Query: 142 EVEAVQEALLLNESELHGRQLKVTA-KRTNVPGMKQFRPRRPSN 184
+ + AL + + + R L + +T P + PRR ++
Sbjct: 211 TEDGAKRALAFDRAAMGDRYLTIQQYVKTTTPSI----PRRKTS 250
Score = 46.6 bits (109), Expect = 1e-05
Identities = 33/125 (26%), Positives = 64/125 (50%), Gaps = 3/125 (2%)
Query: 42 DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARS-IFVGNV 100
D +D KR + A + + + + + P ++S E VD + +++GN+
Sbjct: 210 DTEDGAKRALAFDRAAMGDRYLTIQQYVKTTTPSIPRRKTSSGFAPEMVDGYNRVYIGNL 269
Query: 101 DYACTPEDVQQHFQSCGTVNRVTIRTDK-FGQPKGYAYVEFVEVEAVQEALLLNESELHG 159
+ T D+++ F C +N V + +K G+ KGYA+V+F + +V AL L++ + G
Sbjct: 270 AWDTTERDIRKLFSDC-VINSVRLGKNKETGEFKGYAHVDFKDSVSVAIALKLDQQVICG 328
Query: 160 RQLKV 164
R +K+
Sbjct: 329 RPVKI 333
>At3g52660 putative RNA binding protein
Length = 471
Score = 48.5 bits (114), Expect = 3e-06
Identities = 36/152 (23%), Positives = 73/152 (47%), Gaps = 16/152 (10%)
Query: 26 MENDDVDMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQI 85
ME+++ + DN+ ++ + + +E+E+E ++E++ ++E+E+ ++ A
Sbjct: 15 MESEERVDLDGDNDPEEILEEEVEYEEVEEE--EIEEIEEEIEEEVEVEEEEEEEDAVAT 72
Query: 86 NREEVDAR------------SIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDK-FGQP 132
EE R +++G + T D++ S G V V I +K G
Sbjct: 73 EEEEEKKRHVELLALPPHGSEVYLGGIPTDATEGDLKGFCGSIGEVTEVRIMREKDSGDG 132
Query: 133 KGYAYVEFVEVEAVQEAL-LLNESELHGRQLK 163
KGYA+V F + EA+ LN ++ G+++K
Sbjct: 133 KGYAFVTFRSKDLAAEAIDTLNNTDFRGKRIK 164
>At1g48920 unknown protein
Length = 557
Score = 48.5 bits (114), Expect = 3e-06
Identities = 45/172 (26%), Positives = 73/172 (42%), Gaps = 18/172 (10%)
Query: 25 DMENDDVDMIGADNNEADLD-DMKKRLKEMEDEAAALKEMQAKVEKEMGSVQ-------D 76
D ++D D D A D K K DE++ +E +++ E+E + D
Sbjct: 217 DSSDEDSDEESEDEKPAQKKADTKASKKSSSDESSESEEDESEDEEETPKKKSSDVEMVD 276
Query: 77 PANASASQINREEVDA----RSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQ 131
+SA Q A +++F N+ + DV+ F+ G V V T++ G
Sbjct: 277 AEKSSAKQPKTPSTPAAGGSKTLFAANLSFNIERADVENFFKEAGEVVDVRFSTNRDDGS 336
Query: 132 PKGYAYVEFVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPS 183
+G+ +VEF E Q+AL E HGR L R ++ + R RP+
Sbjct: 337 FRGFGHVEFASSEEAQKAL-----EFHGRPLLGREIRLDIAQERGERGERPA 383
Score = 46.6 bits (109), Expect = 1e-05
Identities = 27/74 (36%), Positives = 43/74 (57%), Gaps = 7/74 (9%)
Query: 91 DARSIFVGNVDYACTPEDVQ----QHFQSCGTVNRVTIRTDK-FGQPKGYAYVEFVEVEA 145
D + IFV D + + +D++ +HF SCG + V++ D+ G KG AY+EF E
Sbjct: 399 DEKKIFVKGFDASLSEDDIKNTLREHFSSCGEIKNVSVPIDRDTGNSKGIAYLEF--SEG 456
Query: 146 VQEALLLNESELHG 159
++AL LN S++ G
Sbjct: 457 KEKALELNGSDMGG 470
>At5g43960 unknown protein
Length = 450
Score = 47.8 bits (112), Expect = 5e-06
Identities = 26/93 (27%), Positives = 54/93 (57%), Gaps = 2/93 (2%)
Query: 88 EEVDARSIFVGNVDYACTPEDVQQHFQSCGTV--NRVTIRTDKFGQPKGYAYVEFVEVEA 145
E+ + +S++V N+ + ++++ F++ GT+ + V +RT K YA+VEF ++ +
Sbjct: 311 EDFEFKSVYVRNLPSDISASEIEEEFKNFGTIKPDGVFLRTRKDVMGVCYAFVEFEDMTS 370
Query: 146 VQEALLLNESELHGRQLKVTAKRTNVPGMKQFR 178
V+ A+ + L GRQ+ + +R N G++ R
Sbjct: 371 VENAIKASPIYLGGRQVYIEERRPNPAGVRGAR 403
>At3g50670 U1 snRNP 70K protein
Length = 427
Score = 47.8 bits (112), Expect = 5e-06
Identities = 25/91 (27%), Positives = 53/91 (57%), Gaps = 4/91 (4%)
Query: 93 RSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALL 151
+++FV ++Y + +++ F+S G + RV + TD+ +PKGYA++E++ ++ A
Sbjct: 138 KTLFVSRLNYESSESKIKREFESYGPIKRVHLVTDQLTNKPKGYAFIEYMHTRDMKAAYK 197
Query: 152 LNESE-LHGRQLKVTAKRTNVPGMKQFRPRR 181
+ + + GR++ V +R + +RPRR
Sbjct: 198 QADGQKIDGRRVLVDVERGRT--VPNWRPRR 226
>At2g46780 putative RNA-binding protein
Length = 304
Score = 47.8 bits (112), Expect = 5e-06
Identities = 34/122 (27%), Positives = 57/122 (45%), Gaps = 7/122 (5%)
Query: 86 NREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDK-FGQPKGYAYVEFVEVE 144
N + IFVG + + + ++++F+ G + + TDK G+ KGY +V F E E
Sbjct: 15 NTTDTKLTKIFVGGLAWETQRDTMRRYFEQFGEIVEAVVITDKNTGRSKGYGFVTFKEAE 74
Query: 145 AVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGF-RGRTPYAPPFAYAP 203
A A + GR+ N+ + +PR P++P G R R+P + AP
Sbjct: 75 AAMRACQNMNPVIDGRR-----ANCNLACLGAQKPRPPTSPRHGTGRFRSPGSGVGLVAP 129
Query: 204 AP 205
+P
Sbjct: 130 SP 131
>At4g14300 ribonucleoprotein like protein
Length = 406
Score = 47.0 bits (110), Expect = 8e-06
Identities = 36/136 (26%), Positives = 60/136 (43%), Gaps = 4/136 (2%)
Query: 60 LKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTV 119
+K ++ E+++ N S S + IFVG + T E+ +Q+F+ G V
Sbjct: 72 VKRAMSREEQQVSGRTGNLNTSRSSGGDAYNKTKKIFVGGLPPTLTDEEFRQYFEVYGPV 131
Query: 120 NRVTIRTDK-FGQPKGYAYVEFVEVEAVQEALLLNESELHGRQLKV---TAKRTNVPGMK 175
V I D+ +P+G+ +V F +AV L +L G+Q++V K N G
Sbjct: 132 TDVAIMYDQATNRPRGFGFVSFDSEDAVDSVLHKTFHDLSGKQVEVKRALPKDANPGGGG 191
Query: 176 QFRPRRPSNPYMGFRG 191
+ S Y G+ G
Sbjct: 192 RSMGGGGSGGYQGYGG 207
Score = 34.3 bits (77), Expect = 0.054
Identities = 21/93 (22%), Positives = 43/93 (45%), Gaps = 13/93 (13%)
Query: 91 DARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYA------------Y 137
D +FVG + + + +++HF + G V++ + DK G+P+G+ Y
Sbjct: 4 DQGKLFVGGISWETDEDKLREHFTNYGEVSQAIVMRDKLTGRPRGFGIRKNRCIYFLLRY 63
Query: 138 VEFVEVEAVQEALLLNESELHGRQLKVTAKRTN 170
V + V+ A+ E ++ GR + R++
Sbjct: 64 VLDLGKVDVKRAMSREEQQVSGRTGNLNTSRSS 96
>At2g33410 putative RNA-binding protein
Length = 404
Score = 47.0 bits (110), Expect = 8e-06
Identities = 31/130 (23%), Positives = 65/130 (49%), Gaps = 1/130 (0%)
Query: 36 ADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSI 95
A ++ A +D + + +++ +K ++ E+ NAS + + V + I
Sbjct: 53 AFSDPAVIDRVLQDKHHIDNRDVDVKRAMSREEQSPAGRSGTFNASRNFDSGANVRTKKI 112
Query: 96 FVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQ-PKGYAYVEFVEVEAVQEALLLNE 154
FVG + A T ++ + +F++ G V+ I D+ Q P+G+ +V F ++V L
Sbjct: 113 FVGGLPPALTSDEFRAYFETYGPVSDAVIMIDQTTQRPRGFGFVSFDSEDSVDLVLHKTF 172
Query: 155 SELHGRQLKV 164
+L+G+Q++V
Sbjct: 173 HDLNGKQVEV 182
Score = 37.0 bits (84), Expect = 0.008
Identities = 20/77 (25%), Positives = 43/77 (54%), Gaps = 2/77 (2%)
Query: 89 EVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDK-FGQPKGYAYVEFVEVEAVQ 147
E D +F+G + + ++++F + G V +VT+ +K G+P+G+ +V F + AV
Sbjct: 2 ESDQGKLFIGGISWDTDENLLREYFSNFGEVLQVTVMREKATGRPRGFGFVAFSD-PAVI 60
Query: 148 EALLLNESELHGRQLKV 164
+ +L ++ + R + V
Sbjct: 61 DRVLQDKHHIDNRDVDV 77
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.137 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,109,973
Number of Sequences: 26719
Number of extensions: 218866
Number of successful extensions: 1504
Number of sequences better than 10.0: 222
Number of HSP's better than 10.0 without gapping: 99
Number of HSP's successfully gapped in prelim test: 123
Number of HSP's that attempted gapping in prelim test: 1260
Number of HSP's gapped (non-prelim): 310
length of query: 223
length of database: 11,318,596
effective HSP length: 96
effective length of query: 127
effective length of database: 8,753,572
effective search space: 1111703644
effective search space used: 1111703644
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC121245.9


