

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121245.1 - phase: 0 /pseudo
(660 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
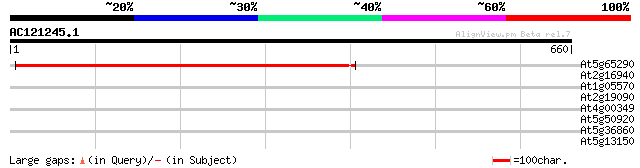
Score E
Sequences producing significant alignments: (bits) Value
At5g65290 unknown protein 591 e-169
At2g16940 splicing factor like protein 33 0.66
At1g05570 putative glucan synthase 32 1.1
At2g19090 hypothetical protein 30 3.3
At4g00349 unknown protein 30 4.3
At5g50920 ATP-dependent Clp protease, ATP-binding subunit 30 5.6
At5g36860 putative protein 29 7.3
At5g13150 putative protein 29 9.5
>At5g65290 unknown protein
Length = 733
Score = 591 bits (1523), Expect = e-169
Identities = 284/401 (70%), Positives = 342/401 (84%), Gaps = 2/401 (0%)
Query: 7 NVTISFLWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVAL 66
N ISFLWS SYWSTFLLTWAVVPL+QG+EDAGDFTV RL+TS+H NLVFYL LG + L
Sbjct: 67 NGAISFLWSWSYWSTFLLTWAVVPLIQGFEDAGDFTVSERLKTSVHVNLVFYLVLGFIGL 126
Query: 67 FGLILLITLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKF 126
GLILLI +++ W+GS+ G+AMACSNTFGLVTGAFLLGFG+SEIPK +W NADW +QK
Sbjct: 127 LGLILLIMMHRNWTGSILGYAMACSNTFGLVTGAFLLGFGLSEIPKTLWKNADWTTRQKV 186
Query: 127 LSHKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSF 186
LSHK+AK+AVKLD+AHQ+ SNAIV+ QATS QMSKRD +RPYMN+ID ML +M EDPSF
Sbjct: 187 LSHKIAKIAVKLDNAHQELSNAIVVAQATSTQMSKRDPMRPYMNVIDAMLAKMFREDPSF 246
Query: 187 KPQGGRLGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVKNYD 246
KPQGG+LGE+DMDYDTDEKSMA+LRR LR A+++YYRY+SEY +V EAL LEDT+KNY+
Sbjct: 247 KPQGGQLGENDMDYDTDEKSMATLRRHLRNAKDEYYRYKSEYLTYVTEALVLEDTMKNYE 306
Query: 247 RRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAILLAEA 306
RRD+TGW+YIS R R GK+G +LD++EF+WRCIL+KQ++ LAV++G MS AILLAEA
Sbjct: 307 RRDATGWKYISSFRATRTGKMGNLLDSLEFMWRCILKKQIQMVLAVVMGIMSAAILLAEA 366
Query: 307 TILPSGVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYSLTPRQ 366
T+L S +DLSLFSIL+ + E+LVQ AFVPL+YMCVCTYYSLFK+GM M YSLTPRQ
Sbjct: 367 TLLLSKLDLSLFSILISSVKSDELLVQAFAFVPLVYMCVCTYYSLFKIGMLMIYSLTPRQ 426
Query: 367 TSSVSLLMICSMVARYAAPISYNFLNLINIGGDRKTVFEKR 407
TSSV+LLMICSM+ARYA PISYNF+NLI + + T+FEK+
Sbjct: 427 TSSVNLLMICSMIARYAPPISYNFINLIQLHSE--TIFEKK 465
>At2g16940 splicing factor like protein
Length = 561
Score = 32.7 bits (73), Expect = 0.66
Identities = 21/66 (31%), Positives = 32/66 (47%)
Query: 182 EDPSFKPQGGRLGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDT 241
+D + + G+ E D D D D K + R RR+R + R+RS+ + LE E
Sbjct: 93 KDVDKEERNGKDRERDRDKDRDSKGRDHEKDRSRRSRSRSERHRSQEREKSLEIEPKERE 152
Query: 242 VKNYDR 247
K+ DR
Sbjct: 153 TKDRDR 158
>At1g05570 putative glucan synthase
Length = 1858
Score = 32.0 bits (71), Expect = 1.1
Identities = 20/72 (27%), Positives = 37/72 (50%), Gaps = 2/72 (2%)
Query: 24 LTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILLITLNKFWSGSV 83
+T A++ L Q D + KAR SL+ L + + +G+ A++ +++ +T W +
Sbjct: 521 ITAAILKLAQAVLDIA-LSWKARHSMSLYVKLRYVMKVGAAAVWVVVMAVTYAYSWK-NA 578
Query: 84 RGFAMACSNTFG 95
GF+ N FG
Sbjct: 579 SGFSQTIKNWFG 590
>At2g19090 hypothetical protein
Length = 844
Score = 30.4 bits (67), Expect = 3.3
Identities = 14/35 (40%), Positives = 20/35 (57%)
Query: 477 SIWSNNFAERALSSSARAQSWRTCFPFSQKFQHEY 511
+I S A+ AL+ A+ ++WRTCF F Q Y
Sbjct: 650 AINSQRLAQSALNLEAQLRNWRTCFEFWITSQRSY 684
>At4g00349 unknown protein
Length = 514
Score = 30.0 bits (66), Expect = 4.3
Identities = 26/92 (28%), Positives = 44/92 (47%), Gaps = 4/92 (4%)
Query: 27 AVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSV--ALFGLILLITLNKFWSGSVR 84
A+V L+ + A FT +R ++ +L + L + + +L +I + + W V
Sbjct: 377 AIVILITRDDFAVIFTESEEMRKAV-ADLAYLLGITMILNSLQPVISGVAVGGGWQAPVA 435
Query: 85 GFAMACSNTFGLVTGAFLLGFGMSEIPKGIWL 116
+ C FGL G FLLG+ S +GIW+
Sbjct: 436 YINLFCYYAFGLPLG-FLLGYKTSLGVQGIWI 466
>At5g50920 ATP-dependent Clp protease, ATP-binding subunit
Length = 929
Score = 29.6 bits (65), Expect = 5.6
Identities = 19/87 (21%), Positives = 37/87 (41%), Gaps = 3/87 (3%)
Query: 137 KLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKPQGGRLGES 196
+++ AH D N ++ + + + N + L+ N S +GGR
Sbjct: 718 EIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTL---LIMTSNVGSSVIEKGGRRIGF 774
Query: 197 DMDYDTDEKSMASLRRRLRRAREQYYR 223
D+DYD + S ++ + +QY+R
Sbjct: 775 DLDYDEKDSSYNRIKSLVTEELKQYFR 801
>At5g36860 putative protein
Length = 1204
Score = 29.3 bits (64), Expect = 7.3
Identities = 20/87 (22%), Positives = 37/87 (41%), Gaps = 21/87 (24%)
Query: 184 PSFKPQGGRLGESDMDYDTDEKS------------MASLRRRLRRAREQYYRYRSEYTKF 231
PS K + +S+MDYD +E + + +RR L+ +Q Y+ + F
Sbjct: 428 PSPKEPAMKNQKSEMDYDKEETAEDCFGEPVPERFIVEMRRSLKELEDQMYQMHEDMKDF 487
Query: 232 VLEALEL---------EDTVKNYDRRD 249
V + + E T+ ++D R+
Sbjct: 488 VRDQIRAALDPKGKRPEQTIPSHDSRE 514
>At5g13150 putative protein
Length = 653
Score = 28.9 bits (63), Expect = 9.5
Identities = 31/116 (26%), Positives = 53/116 (44%), Gaps = 11/116 (9%)
Query: 144 DFSNAIVITQATSKQMSK----RDSLRPYMNIIDKMLVQMLNEDPSFKPQGGRLGESDMD 199
DFS A+V+T+ +S+++ K ++LR + +++ ++ E K RLGE+ +
Sbjct: 324 DFSGAVVLTKRSSEKLFKFLDMYETLRDLIPAVEQSDSDLIQE---IKLAQTRLGEAAVT 380
Query: 200 -YDTDEKSMASLRRRL---RRAREQYYRYRSEYTKFVLEALELEDTVKNYDRRDST 251
+ EKS+ S R A RY Y K+ E E D V + + T
Sbjct: 381 IFGELEKSIKSDNGRTPVPSGAVHPLTRYTMNYLKYACEYKETLDQVFQHYEANQT 436
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.348 0.152 0.521
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,094,341
Number of Sequences: 26719
Number of extensions: 516635
Number of successful extensions: 2316
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 2310
Number of HSP's gapped (non-prelim): 10
length of query: 660
length of database: 11,318,596
effective HSP length: 106
effective length of query: 554
effective length of database: 8,486,382
effective search space: 4701455628
effective search space used: 4701455628
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC121245.1


