

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121241.7 + phase: 0 /pseudo
(69 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
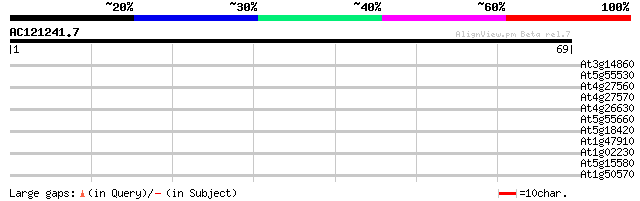
Score E
Sequences producing significant alignments: (bits) Value
At3g14860 unknown protein 38 0.001
At5g55530 unknown protein 35 0.009
At4g27560 UDP rhamnose-anthocyanidin-3-glucoside rhamnosyltransf... 30 0.29
At4g27570 UDP rhamnose--anthocyanidin-3-glucoside rhamnosyltrans... 28 0.84
At4g26630 unknown protein 27 1.4
At5g55660 putative protein 27 1.9
At5g18420 unknown protein 25 5.4
At1g47910 reverse transcriptase, putative 25 5.4
At1g02230 hypothetical protein 25 5.4
At5g15580 putative protein 25 9.3
At1g50570 unknown protein 25 9.3
>At3g14860 unknown protein
Length = 493
Score = 37.7 bits (86), Expect = 0.001
Identities = 20/39 (51%), Positives = 28/39 (71%), Gaps = 3/39 (7%)
Query: 26 TKEETGWPSFRQVV-DLLKLSLEEFTSSIL--KFMSKPH 61
TKEE GWPSF Q++ DL KL+LE TS ++ +F + P+
Sbjct: 308 TKEEPGWPSFGQLLTDLCKLALEFITSHLVPARFQTNPN 346
>At5g55530 unknown protein
Length = 405
Score = 34.7 bits (78), Expect = 0.009
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 6/72 (8%)
Query: 4 QPSERDFKGEASSDKSKSTLERTKEETGWPSFRQ-VVDLLKLSLEEFTSSILKF-----M 57
Q SE + GE S +K+ ++ K E +Q +VD+ S+++FT S+ K +
Sbjct: 311 QESESEASGETSEEKTVKSVLTVKVEPESKVVQQDIVDMYMKSMQQFTDSLAKMKLPLDI 370
Query: 58 SKPHTTERSSSD 69
P +E SSSD
Sbjct: 371 DSPTKSENSSSD 382
>At4g27560 UDP rhamnose-anthocyanidin-3-glucoside
rhamnosyltransferase - like protein
Length = 455
Score = 29.6 bits (65), Expect = 0.29
Identities = 18/52 (34%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Query: 16 SDKSKSTLERTKEETGWPSFRQVVDLLKLSLEEFTSSILKFMSKPHTTERSS 67
SD+ K ++E +EETGW S + D + S+ + S I + K HT R +
Sbjct: 375 SDELKVSVEVAREETGWFSKESLFDAIN-SVMKRDSEIGNLVKKNHTKWRET 425
>At4g27570 UDP rhamnose--anthocyanidin-3-glucoside
rhamnosyltransferase - like protein
Length = 453
Score = 28.1 bits (61), Expect = 0.84
Identities = 17/52 (32%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Query: 16 SDKSKSTLERTKEETGWPSFRQVVDLLKLSLEEFTSSILKFMSKPHTTERSS 67
SD+ K ++E +EETGW S + D + S+ + S + + K HT R +
Sbjct: 375 SDELKVSVEVAREETGWFSKESLCDAVN-SVMKRDSELGNLVRKNHTKWRET 425
>At4g26630 unknown protein
Length = 763
Score = 27.3 bits (59), Expect = 1.4
Identities = 16/59 (27%), Positives = 28/59 (47%), Gaps = 5/59 (8%)
Query: 15 SSDKSKSTLERTKEETGWPSFRQVVDL----LKLSLEEFTSSILKFMSKPHTTERSSSD 69
+ +K K LE+ +E W F V+D+ E+ + + +F+ KPH T + D
Sbjct: 407 AKEKVKEKLEKCTKEKLW-EFCDVLDIHITKATTKKEDIITKLFEFLEKPHVTGDVTGD 464
>At5g55660 putative protein
Length = 759
Score = 26.9 bits (58), Expect = 1.9
Identities = 13/61 (21%), Positives = 29/61 (47%), Gaps = 7/61 (11%)
Query: 10 FKGEASSDKSKSTLERTKEETGWPSFRQVVDLLKLSL-------EEFTSSILKFMSKPHT 62
+K + +K+K ++ E+ + DL +S+ E+ + +++F+ KPH
Sbjct: 392 YKWQGDEEKAKLKVKEKFEKINKEKLLEFCDLFDISVAKATTKKEDIVTKLVEFLEKPHA 451
Query: 63 T 63
T
Sbjct: 452 T 452
>At5g18420 unknown protein
Length = 441
Score = 25.4 bits (54), Expect = 5.4
Identities = 16/55 (29%), Positives = 25/55 (45%), Gaps = 5/55 (9%)
Query: 3 QQPSERDFKGEASSDKSKSTLERTKEE-----TGWPSFRQVVDLLKLSLEEFTSS 52
Q+PS F E S LE+++ W S+ V ++LKLS ++ S
Sbjct: 80 QKPSFNPFLSEMISAACNEQLEKSERAFLLHLLQWNSYNNVKEILKLSAVDYIRS 134
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 25.4 bits (54), Expect = 5.4
Identities = 11/30 (36%), Positives = 18/30 (59%)
Query: 30 TGWPSFRQVVDLLKLSLEEFTSSILKFMSK 59
T WP+F ++ L+ EEFT+ L +S+
Sbjct: 1081 TEWPAFSVYLEELQSDREEFTNFSLSLISR 1110
>At1g02230 hypothetical protein
Length = 192
Score = 25.4 bits (54), Expect = 5.4
Identities = 11/30 (36%), Positives = 18/30 (59%)
Query: 30 TGWPSFRQVVDLLKLSLEEFTSSILKFMSK 59
T WP+F ++ L+ EEFT+ L +S+
Sbjct: 131 TEWPAFSVYLEELQSDREEFTNFSLSLISR 160
>At5g15580 putative protein
Length = 927
Score = 24.6 bits (52), Expect = 9.3
Identities = 15/58 (25%), Positives = 28/58 (47%)
Query: 12 GEASSDKSKSTLERTKEETGWPSFRQVVDLLKLSLEEFTSSILKFMSKPHTTERSSSD 69
G+AS + + + K+ET ++ + + +SS L F S P ++ SS+D
Sbjct: 51 GKASDNVGDTNISADKKETEKSKKKKTAKEKQRGVSSESSSRLSFSSSPCSSSFSSAD 108
>At1g50570 unknown protein
Length = 388
Score = 24.6 bits (52), Expect = 9.3
Identities = 13/38 (34%), Positives = 21/38 (55%), Gaps = 5/38 (13%)
Query: 36 RQVVDLLKLSLEEFTSSILKF-----MSKPHTTERSSS 68
+ +VD+ SL++FT S+ K + P +E SSS
Sbjct: 331 QDIVDMYTKSLQQFTESLAKMKLPLDIDSPTQSENSSS 368
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.306 0.122 0.322
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,365,686
Number of Sequences: 26719
Number of extensions: 41660
Number of successful extensions: 119
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 112
Number of HSP's gapped (non-prelim): 12
length of query: 69
length of database: 11,318,596
effective HSP length: 45
effective length of query: 24
effective length of database: 10,116,241
effective search space: 242789784
effective search space used: 242789784
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC121241.7


