

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121241.10 + phase: 0 /pseudo
(330 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
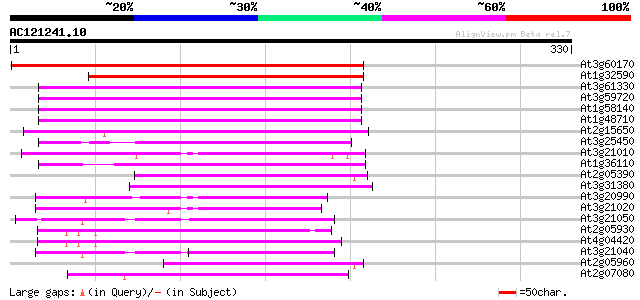
Score E
Sequences producing significant alignments: (bits) Value
At3g60170 putative protein 233 1e-61
At1g32590 hypothetical protein, 5' partial 153 1e-37
At3g61330 copia-type polyprotein 125 2e-29
At3g59720 copia-type reverse transcriptase-like protein 125 2e-29
At1g58140 hypothetical protein 125 2e-29
At1g48710 hypothetical protein 125 2e-29
At2g15650 putative retroelement pol polyprotein 104 8e-23
At3g25450 hypothetical protein 74 1e-13
At3g21010 unknown protein 72 4e-13
At1g36110 hypothetical protein 70 1e-12
At2g05390 putative retroelement pol polyprotein 65 4e-11
At3g31380 hypothetical protein 65 5e-11
At3g20990 unknown protein 61 1e-09
At3g21020 unknown protein 59 4e-09
At3g21050 hypothetical protein 57 1e-08
At2g05930 copia-like retroelement pol polyprotein 57 2e-08
At4g04420 putative transposon protein 55 5e-08
At3g21040 hypothetical protein 54 2e-07
At2g05960 putative retroelement pol polyprotein 54 2e-07
At2g07080 putative gag-protease polyprotein 52 3e-07
>At3g60170 putative protein
Length = 1339
Score = 233 bits (594), Expect = 1e-61
Identities = 118/209 (56%), Positives = 153/209 (72%), Gaps = 2/209 (0%)
Query: 2 NDSSNFVHSIIPKFDGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQ-TEAQRKLI 60
+ S FV IP+FDG+YD W+M MENFLRS+E W L+E I + G +EAQR +
Sbjct: 2 SSSEKFVQPAIPRFDGYYDFWSMTMENFLRSRELWRLVEEGIPAIVVGTTPVSEAQRSAV 61
Query: 61 KEQKLMNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRD 120
+E KL +LK KN+LFQAI ++LETIL+K T K IW+SMK + GS +V RAQLQALR++
Sbjct: 62 EEAKLKDLKVKNFLFQAIDREILETILDKSTSKAIWESMKKKYQGSTKVKRAQLQALRKE 121
Query: 121 FEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEES 180
FE+L MKEGE I+T+ RTLT+ NKMK + E+M Q+ I KILRS+ K +YVVCSIEES
Sbjct: 122 FELLAMKEGEKIDTFLGRTLTVVNKMKTNGEVMEQSTIVSKILRSLTPKFNYVVCSIEES 181
Query: 181 NNLDTMTIDELQSS-LVHEQRIKSNGEEE 208
N+L T++IDEL S LVHEQR+ + +EE
Sbjct: 182 NDLSTLSIDELHGSLLVHEQRLNGHVQEE 210
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 153 bits (387), Expect = 1e-37
Identities = 83/163 (50%), Positives = 110/163 (66%), Gaps = 1/163 (0%)
Query: 47 EAGVEQTEAQRKLIKEQKLMNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GS 106
E V T AQR + E+ + + K KNYLF +I +L+TIL K+T K++W+SMK + G+
Sbjct: 5 ERNVILTGAQRTELAEKTVKDHKVKNYLFASIDKTILKTILQKETSKDLWESMKRKYQGN 64
Query: 107 NRVIRAQLQALRRDFEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSM 166
+RV AQLQ LRR FE+L MK GETI YFSR + I N M+ E M + + EKILR++
Sbjct: 65 DRVQSAQLQRLRRSFEVLEMKIGETITGYFSRVMEITNDMRNLGEDMPDSKVVEKILRTL 124
Query: 167 VSKIDYVVCSIEESNNLDTMTIDELQSSL-VHEQRIKSNGEEE 208
V K YVVC+IEESNN+ +T+D LQSSL VHEQ + + EE
Sbjct: 125 VEKFTYVVCAIEESNNIKELTVDGLQSSLMVHEQNLSRHDVEE 167
>At3g61330 copia-type polyprotein
Length = 1352
Score = 125 bits (315), Expect = 2e-29
Identities = 59/190 (31%), Positives = 112/190 (58%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+YD+W++ M+ L + + W+++E + E ++ Q+ +++ + + K ++Q
Sbjct: 17 NYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQG 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ D E ++ + K W+ ++T + G+++V + +LQ LR +FE L MKEGE ++ YFS
Sbjct: 77 LDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFS 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVH 197
R LT+ N +K + E + I EK+LRS+ K +++V IEE+ +L+ MTI++L SL
Sbjct: 137 RVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQA 196
Query: 198 EQRIKSNGEE 207
+ K E+
Sbjct: 197 YEEKKKKKED 206
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 125 bits (315), Expect = 2e-29
Identities = 59/190 (31%), Positives = 112/190 (58%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+YD+W++ M+ L + + W+++E + E ++ Q+ +++ + + K ++Q
Sbjct: 17 NYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQG 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ D E ++ + K W+ ++T + G+++V + +LQ LR +FE L MKEGE ++ YFS
Sbjct: 77 LDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFS 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVH 197
R LT+ N +K + E + I EK+LRS+ K +++V IEE+ +L+ MTI++L SL
Sbjct: 137 RVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQA 196
Query: 198 EQRIKSNGEE 207
+ K E+
Sbjct: 197 YEEKKKKKED 206
>At1g58140 hypothetical protein
Length = 1320
Score = 125 bits (315), Expect = 2e-29
Identities = 59/190 (31%), Positives = 112/190 (58%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+YD+W++ M+ L + + W+++E + E ++ Q+ +++ + + K ++Q
Sbjct: 17 NYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQG 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ D E ++ + K W+ ++T + G+++V + +LQ LR +FE L MKEGE ++ YFS
Sbjct: 77 LDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFS 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVH 197
R LT+ N +K + E + I EK+LRS+ K +++V IEE+ +L+ MTI++L SL
Sbjct: 137 RVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQA 196
Query: 198 EQRIKSNGEE 207
+ K E+
Sbjct: 197 YEEKKKKKED 206
>At1g48710 hypothetical protein
Length = 1352
Score = 125 bits (315), Expect = 2e-29
Identities = 59/190 (31%), Positives = 112/190 (58%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+YD+W++ M+ L + + W+++E + E ++ Q+ +++ + + K ++Q
Sbjct: 17 NYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQG 76
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
+ D E ++ + K W+ ++T + G+++V + +LQ LR +FE L MKEGE ++ YFS
Sbjct: 77 LDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFS 136
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVH 197
R LT+ N +K + E + I EK+LRS+ K +++V IEE+ +L+ MTI++L SL
Sbjct: 137 RVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQA 196
Query: 198 EQRIKSNGEE 207
+ K E+
Sbjct: 197 YEEKKKKKED 206
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 104 bits (259), Expect = 8e-23
Identities = 61/208 (29%), Positives = 110/208 (52%), Gaps = 5/208 (2%)
Query: 9 HSIIPKFDGH-YDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTE--AQRKLIKEQKL 65
H +IP FDG YD W++ M R+++ W ++E + V E+T A+ K ++E+ +
Sbjct: 6 HQVIPIFDGEKYDFWSIKMATIFRTRKLWSVVEEGVPVEPVQAEETPETARAKTLREEAV 65
Query: 66 MNLKTKNYLFQ-AISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEIL 124
N + Q A++ + I + K WD +K + GS +V +LQ+LRR++E L
Sbjct: 66 TNDTMALQILQTAVTDQIFSRIAAASSSKEAWDVLKDEYQGSPQVRLVKLQSLRREYENL 125
Query: 125 HMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLD 184
M + + I T+ + + + ++ H E + T + +KIL S+ +K D +V +E++ +LD
Sbjct: 126 KMYDNDNIKTFTDKLIVLEIQLTYHGEKKTNTQLIQKILISLPAKFDSIVSVLEQTRDLD 185
Query: 185 TMTIDELQSSL-VHEQRIKSNGEEENGG 211
+T+ EL L E R+ + E G
Sbjct: 186 ALTMSELLGILKAQEARVTAREESTKEG 213
>At3g25450 hypothetical protein
Length = 1343
Score = 73.6 bits (179), Expect = 1e-13
Identities = 53/186 (28%), Positives = 91/186 (48%), Gaps = 20/186 (10%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+Y W M ME LR + W IE A E+ + R L LFQ+
Sbjct: 29 NYTVWTMRMEAVLRVHKLWGTIEPG----SADEEKNDMARAL--------------LFQS 70
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
I ++ + + T +W+++K+ G+ RV A+LQ L +F+ L MK+ ETI+ Y
Sbjct: 71 IPESLILQVGKQKTSSAVWEAIKSRNLGAERVKEARLQTLMAEFDKLKMKDSETIDDYVG 130
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSMV-SKIDYVVCSIEESNNLDTMTIDELQSSL- 195
R I K A E + ++ I +K L+S+ K ++V ++E+ +L T T +++ +
Sbjct: 131 RISEITTKAAALGEDIEESKIVKKFLKSLPRKKYIHIVAALEQVLDLKTTTFEDIAGRIK 190
Query: 196 VHEQRI 201
+E R+
Sbjct: 191 TYEDRV 196
>At3g21010 unknown protein
Length = 438
Score = 72.0 bits (175), Expect = 4e-13
Identities = 54/221 (24%), Positives = 100/221 (44%), Gaps = 24/221 (10%)
Query: 8 VHSIIPKFDGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMN 67
+H + D +Y+ WA + L K WD++EN + + V + A + + K +
Sbjct: 6 IHDAVVSDDLNYEIWAPIAKTTLTEKGLWDVVENGVPPDPSKVPELAATIQAEEFSKWRD 65
Query: 68 LKTKNY-----LFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFE 122
L K+ L +++ L+ + K++WDS++ G+N +A+L+ L + FE
Sbjct: 66 LAEKDMKALQILQSSLTDSAFRKTLSASSAKDLWDSLEK---GNNE--QAKLRRLEKQFE 120
Query: 123 ILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNN 182
L M EGE IN Y R + I +++ S + EK+L ++ D V + + +
Sbjct: 121 ELSMNEGEPINPYIDRVIEIVEQLRRFKIAKSDYQVIEKVLATLSGSYDDVAPVLGDLVD 180
Query: 183 LDTMTI----------DELQSSLVH----EQRIKSNGEEEN 209
L M++ D + +H R++S EEE+
Sbjct: 181 LKNMSLKSFVELFYVYDSITEESIHLMLKANRLRSMAEEES 221
>At1g36110 hypothetical protein
Length = 745
Score = 70.5 bits (171), Expect = 1e-12
Identities = 47/194 (24%), Positives = 94/194 (48%), Gaps = 20/194 (10%)
Query: 18 HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQA 77
+Y WA+ M+ + W+ IE I + N + +FQ+
Sbjct: 26 NYTVWAIRMQIARSVNKMWETIEPGI------------------DDGYKNTMARGLIFQS 67
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFS 137
I + + T K +WDS+KT + G++RV A+LQ L +FE + MKE E I+ +
Sbjct: 68 IPESLTLQVGTLATAKLVWDSIKTRYVGADRVKEARLQTLMAEFEKMKMKESEKIDVFAG 127
Query: 138 RTLTIANKMKAHSEIMSQTIITEKILRSM-VSKIDYVVCSIEESNNLDTMTIDELQSSL- 195
R +A + A + + + +K L ++ + K +++ S+E+ +L+ + +++ +
Sbjct: 128 RLAELATRSDALGSNIETSKLVKKFLNALPLRKYIHIIASLEQVLDLNNTSFEDIVGRIK 187
Query: 196 VHEQRIKSNGEEEN 209
V+E+R+ E+E+
Sbjct: 188 VYEERVWDGEEQED 201
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 65.5 bits (158), Expect = 4e-11
Identities = 39/141 (27%), Positives = 80/141 (56%), Gaps = 4/141 (2%)
Query: 74 LFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETIN 133
LFQ++ + + T K +W+++KT G+ RV A+LQ L +F+ L+MK+ ETI+
Sbjct: 31 LFQSVPESTILQVGKHKTSKAMWEAIKTRNLGAERVKEAKLQTLMAEFDRLNMKDNETID 90
Query: 134 TYFSRTLTIANKMKAHSEIMSQTIITEKILRSMV-SKIDYVVCSIEESNNLDTMTIDELQ 192
+ R I+ K ++ E + ++ I +K L+S+ K +++ ++E+ +L+T +++
Sbjct: 91 EFVGRISEISTKSESLGEEIEESKIVKKFLKSLPRKKYIHIIAALEQILDLNTTGFEDIV 150
Query: 193 SSL-VHEQRI--KSNGEEENG 210
+ +E R+ + + EE G
Sbjct: 151 GRMKTYEDRVCDEDDSPEEQG 171
>At3g31380 hypothetical protein
Length = 262
Score = 65.1 bits (157), Expect = 5e-11
Identities = 42/145 (28%), Positives = 73/145 (49%), Gaps = 2/145 (1%)
Query: 71 KNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGE 130
+ L Q+I + N T K +WDS+KT G+ RV A++Q L DFE + MKE E
Sbjct: 3 RGLLVQSIPEAFTLQVGNLQTAKEVWDSIKTRHVGAERVKEARVQTLMADFEKMKMKEAE 62
Query: 131 TINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMV-SKIDYVVCSIEESNNLDTMTID 189
I+ + R ++ K + + +K L S+ K +++ S+E+ +L+ T +
Sbjct: 63 KIDDFAGRLSELSTKSAPLGVNIEVPKLVKKFLNSLPRKKYIHIIASLEQVLDLNNTTFE 122
Query: 190 ELQSSL-VHEQRIKSNGEEENGGDN 213
++ + V+E+R+ EE N N
Sbjct: 123 DIVGCMKVNEERVYDPEEETNEDQN 147
>At3g20990 unknown protein
Length = 379
Score = 60.8 bits (146), Expect = 1e-09
Identities = 44/182 (24%), Positives = 85/182 (46%), Gaps = 19/182 (10%)
Query: 16 DGHYDHWAMFMENFLRSKEYWDLIENEI---------LVVEAGVEQTEAQRKL-IKEQKL 65
D Y+ W+ + L KE WD++EN + L + E+ R L +K+ K
Sbjct: 14 DFDYEIWSSVTKATLTEKELWDVVENGVPPDPSKIPELATKIQPEELSRWRDLAVKDMKA 73
Query: 66 MNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILH 125
+ + + +++ L+ + K++WDS++ G+N +A+L+ L + FE +
Sbjct: 74 LQILQSS----SLTDSAFRKTLSASSAKDLWDSLEK---GNNE--QAKLRRLEKQFEEIK 124
Query: 126 MKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDT 185
+E E++++Y R I K++ S+ I +K+ SM + +EE N+
Sbjct: 125 REENESLDSYIDRITEIVGKLRVLKVEKSEYEICKKVFASMSDSYGDIHSMMEELMNVHG 184
Query: 186 MT 187
MT
Sbjct: 185 MT 186
>At3g21020 unknown protein
Length = 310
Score = 58.9 bits (141), Expect = 4e-09
Identities = 44/173 (25%), Positives = 82/173 (46%), Gaps = 10/173 (5%)
Query: 16 DGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNY-L 74
D +Y+ WA + L K WD++EN + + V + A+ + + K +L K+
Sbjct: 14 DLNYEIWARIAKTTLTEKGLWDVVENGVPPDPSKVPELAAKIQPEELSKWRDLAVKDMKA 73
Query: 75 FQAISHDVLETILNKDTY----KNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGE 130
Q + + ++ L K Y K++WD ++ G+N +A+L+ L + FE L M E E
Sbjct: 74 LQNLQSSLTDSALRKTLYASSAKDVWDLLEK---GNNE--QAKLRRLEKQFEELSMDEAE 128
Query: 131 TINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNL 183
N+Y + IA +++ S + K+L S+ + + +EE +L
Sbjct: 129 QTNSYLDKVRKIAEQLRRLKNGKSDYEVVTKVLGSLSESHEDLATLLEEQVDL 181
>At3g21050 hypothetical protein
Length = 420
Score = 57.4 bits (137), Expect = 1e-08
Identities = 42/198 (21%), Positives = 86/198 (43%), Gaps = 21/198 (10%)
Query: 4 SSNFVHSIIPKFDGHYDHWAMFMENFLRSKEYWDLIEN----------EILVVEAGVEQT 53
++N + FD Y+HW+ + L WD+++N ++ V+
Sbjct: 2 AANLQDGVSDDFD--YEHWSPIAKKRLVENGVWDVVQNGVSPNPTKIPQLAATIQAVDLA 59
Query: 54 EAQRKLIKEQKLMNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQ 113
+ + ++IK+ K + + + ++ V ++ + K +WD +K N A+
Sbjct: 60 QWRNRVIKDNKALKI-----MQSSLPDSVFRKTISIASAKELWDLLKK----GNDTKEAK 110
Query: 114 LQALRRDFEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYV 173
L+ L + FE L M EGE ++ Y R I + + +S + K+L S+ D
Sbjct: 111 LRRLEKQFEKLMMYEGEPMDLYLKRVEEITERFEVLGNPISDDKVITKLLTSLSWPYDDS 170
Query: 174 VCSIEESNNLDTMTIDEL 191
+ ++E L +T+ +L
Sbjct: 171 IPVLKEFMTLPDLTLRDL 188
>At2g05930 copia-like retroelement pol polyprotein
Length = 916
Score = 56.6 bits (135), Expect = 2e-08
Identities = 46/182 (25%), Positives = 87/182 (47%), Gaps = 11/182 (6%)
Query: 17 GHYDHWAMFMENFLRS--KEYWDLI----ENEILVVEAG--VEQTEAQRKLIKEQKLM-N 67
G+Y HW + M +R KE W + ++ E G V +TE Q +E K N
Sbjct: 31 GNYGHWKVKMRALIRGLGKEAWIATSIGWKAPVIKGEDGEDVLKTEDQWNDAEEAKATAN 90
Query: 68 LKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMK 127
+ + +F +++ + + I N ++ K WD + + G++ V R+++ L FE L M+
Sbjct: 91 SRALSLIFNSVNQNQFKQIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENLTME 150
Query: 128 EGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMT 187
E E I + + IA++ + + +K+LR + S+ + ++ +LDT +
Sbjct: 151 ETENIEEFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAM--GTSLDTNS 208
Query: 188 ID 189
ID
Sbjct: 209 ID 210
>At4g04420 putative transposon protein
Length = 1008
Score = 55.1 bits (131), Expect = 5e-08
Identities = 44/188 (23%), Positives = 88/188 (46%), Gaps = 9/188 (4%)
Query: 17 GHYDHWAMFMENFLRS--KEYWDLI----ENEILVVEAG--VEQTEAQRKLIKEQKLM-N 67
G+Y HW + M +R KE W + ++ E G V +T+ Q +E K N
Sbjct: 19 GNYGHWKVKMRALIRGLGKEAWIATSIGWKAPVIKGEDGEDVLKTKDQWNDAEEAKAKAN 78
Query: 68 LKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMK 127
+ + +F ++ + + I N ++ K WD + + G++ V R+++ L FE L M+
Sbjct: 79 SRALSLIFNFVNQNQFKRIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENLSME 138
Query: 128 EGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMT 187
E E I + + IA++ + + +K+LR + S+ + ++ S + D++
Sbjct: 139 ETENIEEFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTDSID 198
Query: 188 IDELQSSL 195
+E+ L
Sbjct: 199 FEEVVGML 206
>At3g21040 hypothetical protein
Length = 534
Score = 53.5 bits (127), Expect = 2e-07
Identities = 40/186 (21%), Positives = 79/186 (41%), Gaps = 18/186 (9%)
Query: 16 DGHYDHWAMFMENFLRSKEYWDLIEN----------EILVVEAGVEQTEAQRKLIKEQKL 65
D Y+HW+ E L WD+++N E+ + T+ + ++I++ K
Sbjct: 12 DFDYEHWSPIAETRLVENGVWDVVQNGVSPNPTTDPEVAATIKVEDLTQWRNRMIEDNKA 71
Query: 66 MNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILH 125
+ + + A+ V ++ + K +WD +K G++ +L+ L + E L
Sbjct: 72 LKI-----MQSALPDSVFRKTISVASSKELWDLLKK---GNDNEEAKKLRRLEKQLENLT 123
Query: 126 MKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDT 185
M EGE + +Y R I + +S + K+L S+ D + ++E L
Sbjct: 124 MYEGERMKSYLKRVEKIIEEFFVWGNPISDEKVIAKLLTSLPRPYDDSITVLKEFMTLPD 183
Query: 186 MTIDEL 191
+T +L
Sbjct: 184 LTHRDL 189
Score = 43.9 bits (102), Expect = 1e-04
Identities = 26/86 (30%), Positives = 44/86 (50%)
Query: 106 SNRVIRAQLQALRRDFEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRS 165
+N + A+L L + FE L M +GE++++Y R L + +A +S + K+L S
Sbjct: 230 NNPIKDAKLLRLEKQFEKLMMYDGESMDSYLERVLDTIEQFRASGNPLSDDDVIAKLLTS 289
Query: 166 MVSKIDYVVCSIEESNNLDTMTIDEL 191
+ D + +EE NL MT +L
Sbjct: 290 LSWPYDDAIPVLEELMNLPDMTPHDL 315
>At2g05960 putative retroelement pol polyprotein
Length = 1200
Score = 53.5 bits (127), Expect = 2e-07
Identities = 36/127 (28%), Positives = 66/127 (51%), Gaps = 9/127 (7%)
Query: 91 TYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFSRTLTIANKMKAHS 150
T K +WD +++ G+ RV A+L+ L +F+ L MK+ ETI+ R I+ K +
Sbjct: 42 TSKAVWDKIQSRNLGAERVKEAKLKTLMAEFDKLKMKDNETIDECAGRLSEISTKSTSLG 101
Query: 151 EIMSQTIITEKILRSM-VSKIDYVVCSIEESNNLDTMTIDELQSSL-VHEQRI------- 201
E + +T + +K L+S+ K ++V ++E+ +L T ++ + +E +I
Sbjct: 102 EDIEETKVVKKFLKSLPTKKYIHIVAALEQVLDLKNTTFKDIVGRIKTYEDKIWVLITCL 161
Query: 202 KSNGEEE 208
K EEE
Sbjct: 162 KKEAEEE 168
>At2g07080 putative gag-protease polyprotein
Length = 627
Score = 52.4 bits (124), Expect = 3e-07
Identities = 38/169 (22%), Positives = 74/169 (43%), Gaps = 4/169 (2%)
Query: 35 YWDLIENEILV-VEAGVEQTEAQRKLIKEQKLM---NLKTKNYLFQAISHDVLETILNKD 90
YW + +I+ E G + T+ + E+KL N + +F + D + I
Sbjct: 23 YWKVCMTQIIRGQEDGFKITKPKANWTAEEKLQSKFNARAMKAIFNGVDEDEFKLIQGCK 82
Query: 91 TYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFSRTLTIANKMKAHS 150
+ K WD+++ G++ V R +L + FE L M+ E I + S+ +AN+ +
Sbjct: 83 SAKQAWDTLQKSHEGTSSVKRTRLDHIATQFEYLKMEPDEKIVKFSSKISALANEAEVMG 142
Query: 151 EIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVHEQ 199
+ + +K+LR + K + + N D ++ +L L E+
Sbjct: 143 KTYKDQKLVKKLLRCLPPKFAAHKAVMRVAGNTDKISFVDLVGMLKLEE 191
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.334 0.144 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,582,835
Number of Sequences: 26719
Number of extensions: 253801
Number of successful extensions: 1171
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 1121
Number of HSP's gapped (non-prelim): 57
length of query: 330
length of database: 11,318,596
effective HSP length: 100
effective length of query: 230
effective length of database: 8,646,696
effective search space: 1988740080
effective search space used: 1988740080
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC121241.10


