

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121239.4 - phase: 0
(442 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
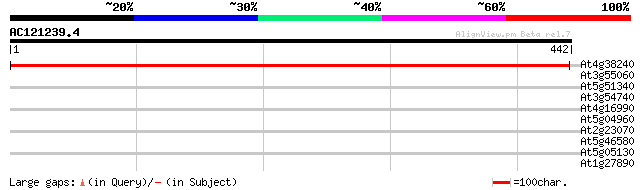
Score E
Sequences producing significant alignments: (bits) Value
At4g38240 glycosyltransferase like protein 679 0.0
At3g55060 centromere protein - like 33 0.41
At5g51340 unknown protein 32 0.92
At3g54740 unknown protein 31 1.2
At4g16990 disease resistance RPP5 like protein 30 2.0
At5g04960 pectinesterase 30 3.5
At2g23070 putative casein kinase II catalytic (alpha) subunit 30 3.5
At5g46580 putative protein 29 5.9
At5g05130 helicase-like transcription factor-like protein 28 7.7
At1g27890 hypothetical protein 28 7.7
>At4g38240 glycosyltransferase like protein
Length = 444
Score = 679 bits (1752), Expect = 0.0
Identities = 315/441 (71%), Positives = 379/441 (85%)
Query: 1 MAKFWCDFRFLLFVAALVFIYIQMRLFASQSQYADRLAAAIEAENHCTAQIRSLIDQISL 60
MA+ CD RFLL AA +FIYIQMRLF +QSQYADRL++AIE+ENHCT+Q+R LID++S+
Sbjct: 1 MARISCDLRFLLIPAAFMFIYIQMRLFQTQSQYADRLSSAIESENHCTSQMRGLIDEVSI 60
Query: 61 QQGRILDLQQERNRREQECSQIKSLVQDLERKDVRRLIDKVQVPVAAVVIMACNRADYLE 120
+Q RI+ L+ +NR+++E Q+K L+Q E+K + +L Q+PVAAVV+MAC+RADYLE
Sbjct: 61 KQSRIVALEDMKNRQDEELVQLKDLIQTFEKKGIAKLTQGGQMPVAAVVVMACSRADYLE 120
Query: 121 RTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFEPVQTERPGELI 180
RT+ SVL YQ P++S++PLF+SQDGS+ VK K+LSY++L++MQHLDFEPV TERPGEL
Sbjct: 121 RTVKSVLTYQTPVASKYPLFISQDGSDQAVKSKSLSYNQLTYMQHLDFEPVVTERPGELT 180
Query: 181 AYYKIARHYKWALGQLFDKHNFNRVIILEDDMEIAPDFFDYFEAMATLLDKDKSIMAVSS 240
AYYKIARHYKWAL QLF KH F+RVIILEDDMEIAPDFFDYFEA A+L+D+DK+IMA SS
Sbjct: 181 AYYKIARHYKWALDQLFYKHKFSRVIILEDDMEIAPDFFDYFEAAASLMDRDKTIMAASS 240
Query: 241 WNDNGQKQFVHDPYELYRSDFFPGLGWMLARSTWDELSPKWPKAYWDDWLRLKENHKGRQ 300
WNDNGQKQFVHDPY LYRSDFFPGLGWML RSTWDELSPKWPK YWDDWLRLKENHKGRQ
Sbjct: 241 WNDNGQKQFVHDPYALYRSDFFPGLGWMLKRSTWDELSPKWPKTYWDDWLRLKENHKGRQ 300
Query: 301 FIRPEVCRTYNFGEHGSSLGQFFKQYLEPIKLNEVQVDWKSMDLSYLLEDKYVIHFANII 360
FIRPEVCRTYNFGEHGSSLGQFF QYLEPIKLN+V VDWK+ DL YL E Y +F+ ++
Sbjct: 301 FIRPEVCRTYNFGEHGSSLGQFFSQYLEPIKLNDVTVDWKAKDLGYLTEGNYTKYFSGLV 360
Query: 361 KKAKPVSGADFAQKARHIDGDVRIKYKDQWDFENIAQQFGIFQEWKDGVPRTAYKGVVVF 420
++A+P+ G+D KA++I DVRI+YKDQ +FE IA +FGIF+EWKDGVPRTAYKGVVVF
Sbjct: 361 RQARPIQGSDLVLKAQNIKDDVRIRYKDQVEFERIAGEFGIFEEWKDGVPRTAYKGVVVF 420
Query: 421 RYQTTKRIFLVGPESLKLLQI 441
R QTT+R+FLVGP+S+ L I
Sbjct: 421 RIQTTRRVFLVGPDSVMQLGI 441
>At3g55060 centromere protein - like
Length = 896
Score = 32.7 bits (73), Expect = 0.41
Identities = 21/100 (21%), Positives = 50/100 (50%), Gaps = 5/100 (5%)
Query: 7 DFRFLLFVAALVFIYIQMRLFASQS---QYADRLAAAIEAENHCTAQIRSLIDQISLQQG 63
+ R L L+ ++ +L++ + Q LAAA+ +++S +D +S+
Sbjct: 729 NLRAELSAETLITSLVREKLYSKEKEIEQLQAELAAAVRGNEILRCEVQSSLDNLSVTTH 788
Query: 64 RILDLQQERNRREQECSQIKSLVQDLERKDVR--RLIDKV 101
+ DL+ + ++E+ +++S +Q+ ++ R L+ KV
Sbjct: 789 ELKDLKHQMLKKEESIRRLESNLQEAAKEMARLNALLSKV 828
>At5g51340 unknown protein
Length = 726
Score = 31.6 bits (70), Expect = 0.92
Identities = 22/84 (26%), Positives = 42/84 (49%), Gaps = 5/84 (5%)
Query: 11 LLFVAALVFIYIQMRL--FASQSQYADRLAAAIEAENHCTAQIRSLIDQIS---LQQGRI 65
L F ++ ++ ++RL + + + DRL A+ A +H +I+ L+D++S L R
Sbjct: 217 LFFYNEMLHVFYRLRLCDYKNAQHHVDRLDQAMNAHSHKMQEIQQLLDELSSLNLSLSRY 276
Query: 66 LDLQQERNRREQECSQIKSLVQDL 89
+ER+ SQ++ V L
Sbjct: 277 DLPSRERSALSARQSQLQDRVNAL 300
>At3g54740 unknown protein
Length = 390
Score = 31.2 bits (69), Expect = 1.2
Identities = 16/54 (29%), Positives = 28/54 (51%)
Query: 39 AAIEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQECSQIKSLVQDLERK 92
A I EN C A + +L Q + L+L++ERN ++ S++ L+R+
Sbjct: 15 AKILVENECAALLEALSSQRETVKDLHLELEEERNAAASAANETMSMILRLQRE 68
>At4g16990 disease resistance RPP5 like protein
Length = 582
Score = 30.4 bits (67), Expect = 2.0
Identities = 25/100 (25%), Positives = 44/100 (44%), Gaps = 15/100 (15%)
Query: 151 KRKALSYDELSHMQHLDFEPVQTERPGELIAYYKIARHYKWALGQLFDKHNFNRVIILED 210
KR+ L ++H+ + F P E +A+ R+ A G+ H FN + ++
Sbjct: 318 KRELLKAHNIAHVYEVGF-------PSEELAHQMFCRY---AFGKNSPPHGFNE--LADE 365
Query: 211 DMEIA---PDFFDYFEAMATLLDKDKSIMAVSSWNDNGQK 247
+IA P Y + LDK++ + +S + NG K
Sbjct: 366 AAKIAGNRPKALKYVGSSFRRLDKEQWVKMLSEFRSNGNK 405
>At5g04960 pectinesterase
Length = 564
Score = 29.6 bits (65), Expect = 3.5
Identities = 16/50 (32%), Positives = 26/50 (52%)
Query: 63 GRILDLQQERNRREQECSQIKSLVQDLERKDVRRLIDKVQVPVAAVVIMA 112
G+I D + R RR E K +V DL + RRL++ + A +++A
Sbjct: 212 GKIADTVKFRRRRLLETGNAKVVVADLPMMEGRRLLESGDLKKKATIVVA 261
>At2g23070 putative casein kinase II catalytic (alpha) subunit
Length = 432
Score = 29.6 bits (65), Expect = 3.5
Identities = 16/47 (34%), Positives = 28/47 (59%), Gaps = 2/47 (4%)
Query: 361 KKAKPVSGADFAQKARHIDGDVRI-KYKDQWDFENIAQQFGIFQEWK 406
K K + A KAR + DV + + KD WD+E++A Q+G+ +++
Sbjct: 88 KIGKSIRRAGAPSKAR-VYADVNVVRPKDYWDYESLAVQWGVQDDYE 133
>At5g46580 putative protein
Length = 711
Score = 28.9 bits (63), Expect = 5.9
Identities = 17/61 (27%), Positives = 32/61 (51%), Gaps = 3/61 (4%)
Query: 121 RTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFEPVQTERPGELI 180
RT+ +++K Q + LF++Q G+ + + L+ H+Q L Q++RPG +
Sbjct: 633 RTLANIIKRQEELPE---LFLAQTGTGTHRFSQGLANSFALHLQQLSAPFRQSDRPGIFV 689
Query: 181 A 181
A
Sbjct: 690 A 690
>At5g05130 helicase-like transcription factor-like protein
Length = 862
Score = 28.5 bits (62), Expect = 7.7
Identities = 14/33 (42%), Positives = 18/33 (54%), Gaps = 4/33 (12%)
Query: 253 PYELYRSDFFP----GLGWMLARSTWDELSPKW 281
P E+ +S+ F GLGW+L R EL P W
Sbjct: 204 PREVIKSELFAHQKEGLGWLLHREKSGELPPFW 236
>At1g27890 hypothetical protein
Length = 302
Score = 28.5 bits (62), Expect = 7.7
Identities = 12/53 (22%), Positives = 25/53 (46%)
Query: 294 ENHKGRQFIRPEVCRTYNFGEHGSSLGQFFKQYLEPIKLNEVQVDWKSMDLSY 346
+N K F++ GE G + +FF+ + + +K + ++ W + SY
Sbjct: 92 KNDKSIAFLKSNGLNLDKIGEEGIGIEEFFRDFSQILKEKDGKITWVNFQGSY 144
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.139 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,839,715
Number of Sequences: 26719
Number of extensions: 427527
Number of successful extensions: 1061
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 1056
Number of HSP's gapped (non-prelim): 10
length of query: 442
length of database: 11,318,596
effective HSP length: 102
effective length of query: 340
effective length of database: 8,593,258
effective search space: 2921707720
effective search space used: 2921707720
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC121239.4


