

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121234.10 + phase: 0
(526 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
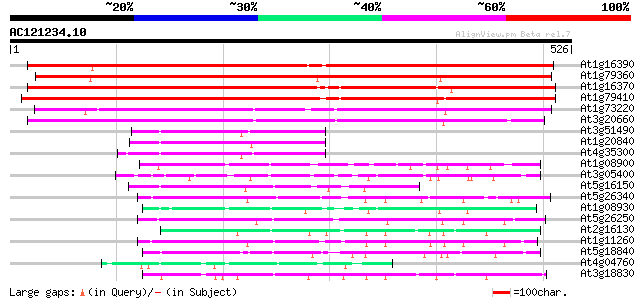
Score E
Sequences producing significant alignments: (bits) Value
At1g16390 putative transport protein 498 e-141
At1g79360 hypothetical protein 491 e-139
At1g16370 putative transport protein 428 e-120
At1g79410 unknown protein 421 e-118
At1g73220 putative transporter 334 6e-92
At3g20660 unknown protein 250 2e-66
At3g51490 sugar transporter-like protein 78 1e-14
At1g20840 putative sugar transporter protein 72 8e-13
At4g35300 sugar transporter like protein 70 4e-12
At1g08900 Beta integral like membrane protein 66 4e-11
At3g05400 sugar transporter like protein 63 4e-10
At5g16150 sugar transporter like protein 62 8e-10
At5g26340 hexose transporter - like protein 60 3e-09
At1g08930 ERD6 protein 60 3e-09
At5g26250 hexose transporter - like protein 60 4e-09
At2g16130 putative sugar transporter 59 7e-09
At1g11260 glucose transporter 59 7e-09
At5g18840 sugar transporter - like protein 59 9e-09
At4g04760 putative sugar transporter 58 1e-08
At3g18830 sugar transporter like protein 58 1e-08
>At1g16390 putative transport protein
Length = 518
Score = 498 bits (1283), Expect = e-141
Identities = 238/501 (47%), Positives = 350/501 (69%), Gaps = 11/501 (2%)
Query: 17 EQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCTNS 76
E P S + IE+ + +FGW FLQA LV+ A FFDAQQ+FI+++TD+ P WHC NS
Sbjct: 16 ESNLPPPRSLEETIERCIGDFGWAQFLQAALVSFAWFFDAQQTFITVFTDSQPMWHCDNS 75
Query: 77 -----IC-TSSSDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIGCLLG 130
+C TSSS++C LP +WSWD +P +IIS W L+CA +F+ G P SSFF+GCL+G
Sbjct: 76 DRVDSVCNTSSSNLCTLPNQTWSWDLNPHVSIISEWGLQCAGSFLKGFPASSFFLGCLIG 135
Query: 131 SFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVL 190
L+ LADSS+GRKNML+ SC+ MS++SML FST++W+Y+ L+FL G R++IGTC L
Sbjct: 136 GLALSTLADSSLGRKNMLLLSCLIMSLSSMLTAFSTSIWVYAFLRFLNGCGRATIGTCAL 195
Query: 191 VLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIA 250
VL TE V +WR +VG + +F FT+G++SLP YIN +SW++LY+W+S+P + Y +
Sbjct: 196 VLSTELVGKKWRGQVGAMGFFCFTLGFLSLPMLGYINEGNSWRNLYVWTSIPTLIYCCLV 255
Query: 251 YLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYS 310
FV ESPRWL+++GR++E + +L+ ++S + NL + + S +Y
Sbjct: 256 RSFVRESPRWLIVKGRKEEAVSILQSIAS-NAITMSFTNLCFE---VENDQSKSNPDVYD 311
Query: 311 SIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVA 370
++ L K W+ R++A M++G G+GMVY+GMPLA+ NL FN+YL VVF+A E P+ +
Sbjct: 312 ALKILVRKSWSFRRLLAAMVVGFGIGMVYYGMPLALTNLNFNLYLGVVFNALSEFPAFLI 371
Query: 371 T-YFLENLRRKPSILVFSILGGICCVTCAVLENRVPAAKVVLAMVAFFGACTAYNVFLIY 429
T +F++ + R+ +++ F+ L GI AVL ++ + ++VL +V+FF ACTA+N+ LIY
Sbjct: 372 TFFFIDKINRRDALIGFTALSGISSALIAVLGQQLGSLQIVLELVSFFSACTAFNMTLIY 431
Query: 430 IIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFL 489
IE+FPTCVRN+ S+VRQA+VFG +F P +++AGR+N +SYG+FG++I L +F L
Sbjct: 432 TIEMFPTCVRNSAISMVRQALVFGGVFSPVMVAAGRENQFWSYGLFGLIIGLCGLFVFGL 491
Query: 490 PETIGIVLCDTMDQQEKKEIA 510
PET G VLCDTMD++E K +A
Sbjct: 492 PETRGSVLCDTMDEEEYKTLA 512
>At1g79360 hypothetical protein
Length = 527
Score = 491 bits (1263), Expect = e-139
Identities = 243/497 (48%), Positives = 349/497 (69%), Gaps = 13/497 (2%)
Query: 25 SWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCT--NSICTSS- 81
S D IE + +FGW FLQA LV+ + FDAQQ+FIS++TD+ P WHCT NSIC S
Sbjct: 19 SLDDTIESYIGSFGWAQFLQAALVSFSGVFDAQQTFISVFTDSEPTWHCTDSNSICHESI 78
Query: 82 SDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSS 141
S+IC LP+++WSWD P ++IS W L+CA +F+ GLP+SSFF+GCL+G L+ LADSS
Sbjct: 79 SNICILPKTAWSWDYSPHVSVISEWGLQCAGSFVKGLPESSFFVGCLIGGLVLSTLADSS 138
Query: 142 IGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEW 201
+GRKNML SC+ M+I++ML +FS N+W+Y+ L+F+ GF R++IGTC LVL TE V +W
Sbjct: 139 LGRKNMLFLSCLVMAISTMLTVFSPNIWVYAVLRFVNGFGRATIGTCALVLSTELVGKKW 198
Query: 202 RFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWL 261
R RVGI+ +F F +G++SLP AY+NR SSW+ LY W+S+P I Y V+ FV ESPRWL
Sbjct: 199 RGRVGIMSFFGFMLGFLSLPLMAYMNRGSSWRILYAWTSIPTIIYCVLVRFFVCESPRWL 258
Query: 262 VMQGREKEILKMLKRVSSEESADDDS---VNLA-SNLPILPPKEKVSF-FQLYSSIGELF 316
++GR +E + +LKRV+S S D S ++++ S+LP +EK S +++++ L
Sbjct: 259 FVRGRREEAISILKRVASIPSTDVSSGGAISMSFSSLPFEEDEEKPSTNVNIFTTMKVLV 318
Query: 317 HKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVATYFL-E 375
KRWA+ R+ AVM + G+G+VY+GMPLA+ NL FNIYL+ F+A M+LP+ + T FL +
Sbjct: 319 EKRWALKRLSAVMAIAFGIGLVYYGMPLALSNLDFNIYLSAAFNALMDLPANLITLFLVD 378
Query: 376 NLRRKPSILVFSILGGICCVTCAVLEN----RVPAAKVVLAMVAFFGACTAYNVFLIYII 431
L R+ +++ F+ LGG+ V L N A ++ L ++++F AC+A+N+ +IY I
Sbjct: 379 KLSRRNALIGFTALGGVSSVLIFALHNMRIGNHGALQLALELISYFSACSAFNMEMIYTI 438
Query: 432 ELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPE 491
ELFPTCVRN+ ++ RQA+V G +F P +++AGRKN +S+G+FG+ I L LPE
Sbjct: 439 ELFPTCVRNSAIAMARQALVLGGVFSPIMVAAGRKNAFWSFGLFGLAIGLLGLFAVGLPE 498
Query: 492 TIGIVLCDTMDQQEKKE 508
T G LCDTMD++E K+
Sbjct: 499 TRGSDLCDTMDEEECKD 515
>At1g16370 putative transport protein
Length = 521
Score = 428 bits (1101), Expect = e-120
Identities = 222/501 (44%), Positives = 335/501 (66%), Gaps = 12/501 (2%)
Query: 17 EQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCTN- 75
E+ + L++D I+E+SLS+FG+ F Q LV +A+ FDAQQ FI++YTD YP WHC N
Sbjct: 21 EKTRLEALTFDKIVEQSLSDFGFWQFFQISLVGLALLFDAQQIFITVYTDAYPTWHCLNH 80
Query: 76 SICT-SSSDICKLPRSSWSWDT-HPSNTIISHWNLECASTFITGLPQSSFFIGCLLGSFF 133
+IC S+SDICKLPRS+W WD ++IS + LEC+S+ + G+P S+F+IG ++G FF
Sbjct: 81 TICDPSASDICKLPRSAWEWDGGSQGKSVISEFGLECSSSLLRGMPSSAFYIGAIVGGFF 140
Query: 134 LAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLL 193
LA + D S+GRK +++FS +MSITS+ +IFSTNVWIY+ LKF+IGF RS + LVL+
Sbjct: 141 LALIPDDSLGRKKLVLFSTFAMSITSISVIFSTNVWIYTFLKFIIGFSRSQTWSYALVLI 200
Query: 194 TEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLF 253
+E+VS WR R ++ + F +G+MSL G A++ ++SSW+ LY+++SVPA+ Y + YLF
Sbjct: 201 SERVSTRWRPRATMIPFTLFVLGFMSLSGIAFLAQDSSWRYLYLYTSVPAVFYCIFLYLF 260
Query: 254 VTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYSSIG 313
ESPRWL MQG++KE + +L ++S +E A +SV S LP+ +E Y SI
Sbjct: 261 ALESPRWLHMQGKDKEAIDVLTKMSPKEKAYLESV--VSKLPL--KQENFEQAPTY-SIK 315
Query: 314 ELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSCVAT-Y 372
+ F ++WA R++ VMI+ GLG+ Y+G+PLA ++ NIYL+ +A +ELP+ V T
Sbjct: 316 DFFFRKWAFRRILVVMIIMFGLGISYYGVPLAARDIDVNIYLSETLNALVELPTFVITPI 375
Query: 373 FLENLRRKPSILVFSILGGICCVTCAVLENRVPAAKVVLA--MVAFFGACTAYNVFLIYI 430
LE R+ S+LV ++LGG V C VL + + ++ A + FF A +N+ +++
Sbjct: 376 LLERFNRRSSVLVNTLLGGASGVLCFVL-SILGKTEIAFAFELGTFFCARIGFNLMAVFM 434
Query: 431 IELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLP 490
+E+FPTCVR++ T + RQA+V G CP + S GR S+ +FG+ + + LP
Sbjct: 435 VEMFPTCVRSSATMMFRQALVVGGACCPLIASIGRYIPSVSFAIFGIAMSGLGMFVLILP 494
Query: 491 ETIGIVLCDTMDQQEKKEIAL 511
ET G+ LCD+M++QEK++ A+
Sbjct: 495 ETKGLSLCDSMEEQEKRDQAV 515
>At1g79410 unknown protein
Length = 515
Score = 421 bits (1083), Expect = e-118
Identities = 223/506 (44%), Positives = 329/506 (64%), Gaps = 13/506 (2%)
Query: 12 SHSNIEQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKW 71
+H +++ L++D I+EKSLS+FG+ FLQ VLV +A+ FD+QQ FI+++TD YP W
Sbjct: 11 THIEEDEDTSSPLTFDKILEKSLSDFGFSQFLQIVLVGLALTFDSQQIFITVFTDAYPTW 70
Query: 72 HCTN-SICT-SSSDICKLPRSSWSWDT-HPSNTIISHWNLECASTFITGLPQSSFFIGCL 128
HC + +IC +++DICK+PRS+W WD ++IS ++LEC+S+F+ LP S+F++G +
Sbjct: 71 HCLDHTICNPATTDICKIPRSAWDWDGGFKGKSVISEFDLECSSSFLRSLPSSTFYVGSI 130
Query: 129 LGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTC 188
+G LA + D S+GRK +L FS +MS+T + I S+N+WIYS LKF+IGF RS GT
Sbjct: 131 VGGVVLAMIPDGSLGRKQLLFFSSFAMSLTGISIFLSSNIWIYSFLKFVIGFARSQTGTY 190
Query: 189 VLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSV 248
LVL++E++S +WR R +V + F +G+MSL G AY+ R++SWK LY+ +S+PA +S+
Sbjct: 191 ALVLISERISTKWRPRATMVPFTLFVLGFMSLSGIAYLVRHASWKVLYLCTSIPAGIHSI 250
Query: 249 IAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQL 308
Y F ESPRWL ++G+ KE +++LKR+S +SV+ L PKE +
Sbjct: 251 FIYFFALESPRWLHLEGKNKEAIEVLKRISPANRGYLESVSSR-----LRPKETLEQTSS 305
Query: 309 YSSIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSC 368
Y SI +LF +WA R+ VMI+ GLGM Y+G+PLAV ++ NIY++ +A +ELP+
Sbjct: 306 Y-SIKDLFIIKWAFRRVTLVMIIMFGLGMSYYGVPLAVRDIKVNIYMSEALNAMVELPTF 364
Query: 369 VAT-YFLENLRRKPSILVFSILGGICCVTCAV--LENRVPAAKVVLAMVAFFGACTAYNV 425
V T LE R+ S+LV ++GG V C V L R A L + +FF A +N+
Sbjct: 365 VVTPILLEQFSRRSSVLVNCLIGGASGVLCFVMSLYGRTKIA-FALELGSFFCARIGFNL 423
Query: 426 FLIYIIELFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFT 485
IY++ELFPTCVRN+ T ++RQA+V G CP + S GR S+ VFG +
Sbjct: 424 MAIYLVELFPTCVRNSATMMLRQALVVGGACCPLIASLGRNVPSLSFAVFGFAMSGLGLF 483
Query: 486 LFFLPETIGIVLCDTMDQQEKKEIAL 511
LPET G+ LCDTM++QE+++ AL
Sbjct: 484 ALLLPETKGLSLCDTMEEQEQRDQAL 509
>At1g73220 putative transporter
Length = 539
Score = 334 bits (857), Expect = 6e-92
Identities = 196/496 (39%), Positives = 286/496 (57%), Gaps = 20/496 (4%)
Query: 24 LSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYP--KWHCTNSICTSS 81
L+ D +IE+ + G+ L A+LV++A FDAQ + ISI++D P + T +I +
Sbjct: 43 LTVDEVIEQHIGALGFAQILHALLVSIAWIFDAQTTLISIFSDAQPAARLLATGAIVEGA 102
Query: 82 SDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSS 141
S +C L W W S+T++S WNL C F+ +P + FFIG L GS LADS
Sbjct: 103 S-LCGLASGEWEWIGPKSDTVVSEWNLICQHKFLVAVPSTLFFIGSLFGSGVYGYLADSW 161
Query: 142 IGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEW 201
GRK L+ SCV +T+ I FS NVW+Y+ L+F GF+RS IG+C +VL TE V +W
Sbjct: 162 FGRKKTLLLSCVLTFVTAFAISFSPNVWVYAFLRFANGFFRSGIGSCCIVLATEIVGKKW 221
Query: 202 RFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWL 261
R +VG +F FT+G++SLP AY+ R SW++LY S + Y+V F ESPRWL
Sbjct: 222 RGQVGQYGFFFFTLGFLSLPLMAYLER-KSWRNLYRIISFLPLGYAVCLLPFAYESPRWL 280
Query: 262 VMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYSSIGELFHKRWA 321
+++GR KE + +LK++ A + L ++L ++ P + SS + + +WA
Sbjct: 281 LVKGRNKEAMVVLKKL-----ARLNGKQLPADLSLVDPIPERD--DQTSSSEKFWKTKWA 333
Query: 322 VIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPSC-VATYFLENLRRK 380
V R+I VM+ G G G VY+G+ L NL FN+YL V +A ME P+ + ++ L + R+
Sbjct: 334 VKRIIMVMMAGFGSGFVYYGIQLNAENLNFNLYLTVAVNALMEFPAVFIGSFLLGVMNRR 393
Query: 381 PSILVFSILGGICCVTCAVLE-NRVPAA-------KVVLAMVAFFGACTAYNVFLIYIIE 432
P S L G C+ CAVL +RV A ++ + V F + TAY+V +Y +E
Sbjct: 394 PLFSNSSYLAGFACLLCAVLSIHRVIRAISVAKWLQLAVEAVGFMASSTAYDVLYVYCVE 453
Query: 433 LFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPET 492
LFPT VRNT SL+RQA + G P L++ GR++ + S+ VFGV +LS +L ET
Sbjct: 454 LFPTNVRNTAVSLLRQAFMLGASAAPLLVALGRESAMMSFIVFGVASVLSGIVSLWLRET 513
Query: 493 IGIVLCDTMDQQEKKE 508
L +T+ QQ K E
Sbjct: 514 RNAPLYETLAQQGKAE 529
>At3g20660 unknown protein
Length = 526
Score = 250 bits (638), Expect = 2e-66
Identities = 148/489 (30%), Positives = 241/489 (49%), Gaps = 10/489 (2%)
Query: 17 EQEKKPILSWDVIIEKSLSNFGWMDFLQAVLVAVAMFFDAQQSFISIYTDNYPKWHCTNS 76
E E + L D ++++ FG VL +A +A + + I+ D P+W C S
Sbjct: 25 EAEGEERLCIDEMLQRYCGEFGRWQLKHFVLTCIAWALEAFHTMVMIFADQEPEWRCVGS 84
Query: 77 ICTSSSDICKLPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAA 136
C S C+L SSW W ++ +S W L C + GL Q+ FF GC++G+
Sbjct: 85 DCRVGSLNCELDPSSWEWTAGKGSSTVSEWGLICGDKYKVGLVQALFFAGCMIGAGVFGH 144
Query: 137 LADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEK 196
L+DS +GRK L C+ +I + FS N W Y L+FL GF +G VL TE
Sbjct: 145 LSDSKLGRKGSLTVVCIINAIFGIATAFSPNYWTYVVLRFLTGFSTGGVGLTAFVLATEP 204
Query: 197 VSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTE 256
+ R G+ ++ F+ G L G AY+ R SW+ L+I SS+P++ + +I F++E
Sbjct: 205 IGPSKRGVAGMSTFYFFSAGIAVLSGIAYVFR--SWRELFIVSSLPSLLFLLIVIPFISE 262
Query: 257 SPRWLVMQGREKEILKMLKRVSSEESADDDS-VNLASNLPILPPKEKVSFFQLYSSIGEL 315
SPRW +++G+ E +K++ ++ + V LA + + + + + S+ ++
Sbjct: 263 SPRWYLVRGKVDEAMKLMHSIAKTNGRHIPAGVTLALDDDVENNNGERNT-AVEGSLKDV 321
Query: 316 FHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVFSASMELPS-CVATYFL 374
+R++ + + + +VY+G+ L VGNL N+YL V +A E+P+ + L
Sbjct: 322 ILSPLMRMRLVISVAISFTVSIVYYGLSLNVGNLKTNLYLNVFVNAVSEMPAFAITAVLL 381
Query: 375 ENLRRKPSILVFSILGGICCVTCAVLENRVP--AAKVVLAMVAFFGACTAYNVFLIYIIE 432
+ RKP + + C+ + P + ++V ++ FG YN+ IYI E
Sbjct: 382 DKYGRKPLSIGTQWFSCVFCLVGFSVWGAGPWKSVRMVSGVLGIFGMAGTYNLLFIYIAE 441
Query: 433 LFPTCVRNTTTSLVRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPET 492
LFPT VRN QA G I PF++ G + +GVF V ++ F+LPET
Sbjct: 442 LFPTVVRNAALGCATQAAQMGAILAPFVVVLGEE---LPFGVFAVCGLVGGGLAFYLPET 498
Query: 493 IGIVLCDTM 501
+ L DTM
Sbjct: 499 LNKPLYDTM 507
>At3g51490 sugar transporter-like protein
Length = 729
Score = 78.2 bits (191), Expect = 1e-14
Identities = 55/186 (29%), Positives = 94/186 (49%), Gaps = 6/186 (3%)
Query: 115 ITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSAL 174
I GL + IG L + F ++D +GR++MLI S V ++S+++ +S NV++
Sbjct: 44 IEGLIVAMSLIGATLITTFSGPVSDK-VGRRSMLILSSVLYFLSSIVMFWSPNVYVLLFA 102
Query: 175 KFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMG----YMSLPGFAYINRNS 230
+ L GF T V + ++E +E R + F + G Y + G + + +
Sbjct: 103 RLLDGFGIGLAVTLVPIYISETAPSEIRGLLNTFPQFCGSGGMFLSYCLVFGMS-LQESP 161
Query: 231 SWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNL 290
SW+ + S+P+I Y V+A F+ ESPRWLV +GR E ++L+R+ E + L
Sbjct: 162 SWRLMLGVLSIPSIAYFVLAAFFLPESPRWLVSKGRMDEARQVLQRLRGREDVSGELALL 221
Query: 291 ASNLPI 296
L +
Sbjct: 222 VEGLGV 227
>At1g20840 putative sugar transporter protein
Length = 734
Score = 72.0 bits (175), Expect = 8e-13
Identities = 48/187 (25%), Positives = 91/187 (47%), Gaps = 4/187 (2%)
Query: 113 TFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYS 172
T + GL + IG + + ++D +GR+ MLI S V + +++++S NV++
Sbjct: 40 TSVQGLVVAMSLIGATVITTCSGPISDW-LGRRPMLILSSVMYFVCGLIMLWSPNVYVLC 98
Query: 173 ALKFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAY---INRN 229
+ L GF T V V ++E E R ++ + F + G + ++ +
Sbjct: 99 FARLLNGFGAGLAVTLVPVYISETAPPEIRGQLNTLPQFLGSGGMFLSYCMVFTMSLSDS 158
Query: 230 SSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVN 289
SW+++ S+P++ Y + ++ ESPRWLV +GR E ++L+++ E D+
Sbjct: 159 PSWRAMLGVLSIPSLLYLFLTVFYLPESPRWLVSKGRMDEAKRVLQQLCGREDVTDEMAL 218
Query: 290 LASNLPI 296
L L I
Sbjct: 219 LVEGLDI 225
>At4g35300 sugar transporter like protein
Length = 739
Score = 69.7 bits (169), Expect = 4e-12
Identities = 54/199 (27%), Positives = 98/199 (49%), Gaps = 7/199 (3%)
Query: 102 IISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSML 161
I +NLE ++ + GL + IG L + +AD +GR+ MLI S + + S++
Sbjct: 32 IKKEFNLE-SNPSVEGLIVAMSLIGATLITTCSGGVADW-LGRRPMLILSSILYFVGSLV 89
Query: 162 IIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMG----Y 217
+++S NV++ + L GF + T V + ++E E R + + FT + G Y
Sbjct: 90 MLWSPNVYVLLLGRLLDGFGVGLVVTLVPIYISETAPPEIRGLLNTLPQFTGSGGMFLSY 149
Query: 218 MSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRV 277
+ G + + + SW+ + +P++ + + F+ ESPRWLV +GR E ++L+R+
Sbjct: 150 CMVFGMSLMP-SPSWRLMLGVLFIPSLVFFFLTVFFLPESPRWLVSKGRMLEAKRVLQRL 208
Query: 278 SSEESADDDSVNLASNLPI 296
E + L L I
Sbjct: 209 RGREDVSGEMALLVEGLGI 227
>At1g08900 Beta integral like membrane protein
Length = 462
Score = 66.2 bits (160), Expect = 4e-11
Identities = 92/415 (22%), Positives = 174/415 (41%), Gaps = 67/415 (16%)
Query: 122 SFFIGCL-LGSFFLAALA---DSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFL 177
SFF + LG A + + +GR+ + S V + + F+ ++ + + +
Sbjct: 65 SFFTSVMTLGGMITAVFSGKISALVGRRQTMWISDVCCIFGWLAVAFAHDIIMLNTGRLF 124
Query: 178 IGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYI 237
+GF I V V + E +R G Y + + + + W++L +
Sbjct: 125 LGFGVGLISYVVPVYIAEITPKTFR---GGFSYSNQLLQCLGISLMFFTGNFFHWRTLAL 181
Query: 238 WSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPIL 297
S++P+ + VI F+ ESPRWL M G+++E+ LK++ E S ++ +
Sbjct: 182 LSAIPS-AFQVICLFFIPESPRWLAMYGQDQELEVSLKKLRGENS------DILKEAAEI 234
Query: 298 PPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAV 357
++S + S I +LFH I +I+G+GL + + G+ + Y A
Sbjct: 235 RETVEISRKESQSGIRDLFH-----IGNAHSLIIGLGLML----LQQFCGSAAISAYAAR 285
Query: 358 VFSASMELPSCVATYFL---------------ENLRRKPSILVFSILGGICCVTCAV--- 399
+F + PS + T L + R+P +++ SI G+C + +
Sbjct: 286 IFDKA-GFPSDIGTTILAVILIPQSIVVMLTVDRWGRRPLLMISSI--GMCICSFFIGLS 342
Query: 400 --LENRVPAAKV--VLAMVAFFGACTAYNVFL-----IYIIELFPTCVRNTTTSLVRQA- 449
L+ K+ V+ +V G +++ + L + + E+FP V+ T SLV +
Sbjct: 343 YYLQKNGEFQKLCSVMLIVGLVGYVSSFGIGLGGLPWVIMSEIFPVNVKITAGSLVTMSN 402
Query: 450 ------IVFGCIFCPFLISAGRKNNIYSYGVF-GVVIMLSNFTLFFLPETIGIVL 497
I++ F ++G +Y +F GV ++ F +PET G L
Sbjct: 403 WFFNWIIIYSFNFMIQWSASG------TYFIFSGVSLVTIVFIWTLVPETKGRTL 451
>At3g05400 sugar transporter like protein
Length = 462
Score = 63.2 bits (152), Expect = 4e-10
Identities = 113/432 (26%), Positives = 183/432 (42%), Gaps = 65/432 (15%)
Query: 100 NTIISHWNLECASTFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITS 159
++I+S +L A + G S G ++G+ F +A A S+ G K M ++ IT
Sbjct: 52 SSIMSDLDLSLAQFSLFG---SLSTFGGMIGAIF-SAKAASAFGHK-MTLWVADLFCITG 106
Query: 160 MLIIFSTN--VWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGY 217
L I +W+ +FL+G I V V + E R FTF+
Sbjct: 107 WLAISLAKDIIWLDMG-RFLVGIGVGLISYVVPVYIAEITPKHVRGA------FTFSNQL 159
Query: 218 MSLPGFA---YINRNSSWKSLYIWSSVPAICY-SVIAYLFVTESPRWLVMQGREKEILKM 273
+ G A Y SW++L I S+P C+ VI F+ ESPRWL +GR+KE ++
Sbjct: 160 LQNCGVAVVYYFGNFLSWRTLAIIGSIP--CWIQVIGLFFIPESPRWLAKKGRDKECEEV 217
Query: 274 LKRVSSEESADDDSVNLASNLPILPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGI 333
L+++ + D V A + I K + +I LF KR+A + +GI
Sbjct: 218 LQKLRGRKY---DIVPEACEIKISVEASKKN---SNINIRSLFEKRYA-----HQLTIGI 266
Query: 334 GL----------GMVYFGMPLAVGNLGFNIYLAVVFSASMELP-SCVATYFLENLRRKPS 382
GL G+ +G L GF + ++ + + +P S + ++ R+P
Sbjct: 267 GLMLLQQLCGTAGISSYGSTL-FKLAGFPARIGMMVLSLIVVPKSLMGLILVDRWGRRPL 325
Query: 383 ILVFSILGGIC--CVTCAVL--ENRVPAAKVVLAMVAFFGACTAYNVFLI------YII- 431
++ ++ G+C C+T AV VP + + F G + +F I +II
Sbjct: 326 LMTSAL--GLCLSCITLAVAFGVKDVPGIGKITPIFCFIGILSFTMMFAIGMGALPWIIM 383
Query: 432 -ELFPTCVRNTTTSLVRQAIVF-----GCIFCPFLISAGRKNNIYSYGVFGVVIMLSNFT 485
E+FP ++ SLV A F F L+ + I S + G I+ FT
Sbjct: 384 SEIFPMDIKVLAGSLVTIANWFTGWIANYAFNFMLVWSPSGTFIISAIICGATIV---FT 440
Query: 486 LFFLPETIGIVL 497
+PET + L
Sbjct: 441 WCLVPETRRLTL 452
>At5g16150 sugar transporter like protein
Length = 546
Score = 62.0 bits (149), Expect = 8e-10
Identities = 65/283 (22%), Positives = 117/283 (40%), Gaps = 22/283 (7%)
Query: 112 STFITGLPQSSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIY 171
+T + G SS G +GSF ALAD GR + ++I + L + +V
Sbjct: 142 NTVLQGWIVSSLLAGATVGSFTGGALADK-FGRTRTFQLDAIPLAIGAFLCATAQSVQTM 200
Query: 172 SALKFLIGFWRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMS--LPGFAYINRN 229
+ L G V + ++E E R +G V +G ++ + G
Sbjct: 201 IVGRLLAGIGIGISSAIVPLYISEISPTEIRGALGSVNQLFICIGILAALIAGLPLAANP 260
Query: 230 SSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVN 289
W++++ + +P++ + I F ESPRWLV QG+ E K +K + +E V
Sbjct: 261 LWWRTMFGVAVIPSVLLA-IGMAFSPESPRWLVQQGKVSEAEKAIKTLYGKERV----VE 315
Query: 290 LASNLPIL---PPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMIL-----GIGLGMVYFG 341
L +L + + +F L+SS + W V+ + A + L GI + Y
Sbjct: 316 LVRDLSASGQGSSEPEAGWFDLFSS------RYWKVVSVGAALFLFQQLAGINAVVYYST 369
Query: 342 MPLAVGNLGFNIYLAVVFSASMELPSCVATYFLENLRRKPSIL 384
+ ++ + + AS + VA+ ++ + RK +L
Sbjct: 370 SVFRSAGIQSDVAASALVGASNVFGTAVASSLMDKMGRKSLLL 412
>At5g26340 hexose transporter - like protein
Length = 526
Score = 60.1 bits (144), Expect = 3e-09
Identities = 96/417 (23%), Positives = 180/417 (43%), Gaps = 43/417 (10%)
Query: 121 SSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGF 180
SS ++ L +FF A+ ++GR+ ++ + V I L + ++ + A + L+G
Sbjct: 88 SSLYLAGLTATFF-ASYTTRTLGRRLTMLIAGVFFIIGVALNAGAQDLAMLIAGRILLGC 146
Query: 181 WRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLP----GFAYINRNSSWKSLY 236
V + L+E R + I+ T+G + G A I W+
Sbjct: 147 GVGFANQAVPLFLSEIAPTRIRGGLNILFQLNVTIGILFANLVNYGTAKIKGGWGWRLSL 206
Query: 237 IWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPI 296
+ +PA+ +V A L VTE+P LV +GR E +L+R+ ++ + + +L
Sbjct: 207 GLAGIPALLLTVGA-LLVTETPNSLVERGRLDEGKAVLRRIRGTDNVEPEFADLL-EASR 264
Query: 297 LPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILG--IGLGMVYFGMPLAVGNLGF--- 351
L + K F L R ++ +A+ I G+ + F P+ LGF
Sbjct: 265 LAKEVKHPFRNLLQR-----RNRPQLVIAVALQIFQQCTGINAIMFYAPVLFSTLGFGSD 319
Query: 352 -NIYLAVVFSASMELPSCVATYFLENLRRKPSIL-----------VFSILGGICCVTCAV 399
++Y AVV A L + V+ Y ++ + R+ +L V +I+ G+ VT
Sbjct: 320 ASLYSAVVTGAVNVLSTLVSIYSVDKVGRRVLLLEAGVQMFFSQVVIAIILGV-KVTDTS 378
Query: 400 LENRVPAAKVVLAMVAFFGACTAYN---VFLIYIIELFPTCVRNTTTSLVRQAIVFGCIF 456
A +V+ M+ + A A++ + + E FP R+ S+ + +F
Sbjct: 379 TNLSKGFAILVVVMICTYVAAFAWSWGPLGWLIPSETFPLETRSAGQSV---TVCVNLLF 435
Query: 457 CPFLISAGRKNNI--YSYGVF----GVVIMLSNFTLFFLPETIGIVLCDTMDQQEKK 507
F+I+ + + + +G+F V+++S F +F LPET I + + ++ KK
Sbjct: 436 -TFIIAQAFLSMLCHFKFGIFIFFSAWVLIMSVFVMFLLPETKNIPIEEMTERVWKK 491
>At1g08930 ERD6 protein
Length = 496
Score = 60.1 bits (144), Expect = 3e-09
Identities = 91/401 (22%), Positives = 161/401 (39%), Gaps = 56/401 (13%)
Query: 125 IGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLII-FSTNVWIYSALKFLIGFWRS 183
+G L+G+ F +AD +GRK ++F C IT L + + N + L+G
Sbjct: 106 LGGLIGAVFSGKVADV-LGRKRTMLF-CEFFCITGWLCVALAQNAMWLDCGRLLLGIGVG 163
Query: 184 SIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPA 243
+ V + E R G + M + F I W+ L + VP
Sbjct: 164 IFSYVIPVYIAEIAPKHVR---GSFVFANQLMQNCGISLFFIIGNFIPWRLLTVVGLVPC 220
Query: 244 ICYSVIAYLFVTESPRWLVMQGREKEILKMLKR-------VSSEESADDDSVNLASNLPI 296
+ + V F+ ESPRWL GR+KE L+R +S E + D++++ N
Sbjct: 221 V-FHVFCLFFIPESPRWLAKLGRDKECRSSLQRLRGSDVDISREANTIRDTIDMTEN--- 276
Query: 297 LPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGIGL----------GMVYFGMPLAV 346
+ K+S ELF +R+A +I+G+GL G+ Y+ L
Sbjct: 277 -GGETKMS---------ELFQRRYAY-----PLIIGVGLMFLQQLCGSSGVTYYASSL-F 320
Query: 347 GNLGFNIYLAVVFSASMELP-SCVATYFLENLRRKPSILVFSILGGICCVTCAVLE---- 401
GF + A++ +P + +AT ++ + R+ ++ G+ + +V
Sbjct: 321 NKGGFPSAIGTSVIATIMVPKAMLATVLVDKMGRRTLLMASCSAMGLSALLLSVSYGFQS 380
Query: 402 -NRVPAAKVVLAMVAFFGACTAYNVFL-----IYIIELFPTCVRNTTTSLVRQA-IVFGC 454
+P + + G ++ + + I + E+FP V+ + +LV +FG
Sbjct: 381 FGILPELTPIFTCIGVLGHIVSFAMGMGGLPWIIMAEIFPMNVKVSAGTLVTVTNWLFGW 440
Query: 455 IFCPFLISAGRKNNIYSYGVFGVVIMLSNFTLFFL-PETIG 494
I N + +F +V S ++FL PET G
Sbjct: 441 IITYTFNFMLEWNASGMFLIFSMVSASSIVFIYFLVPETKG 481
>At5g26250 hexose transporter - like protein
Length = 507
Score = 59.7 bits (143), Expect = 4e-09
Identities = 86/405 (21%), Positives = 166/405 (40%), Gaps = 31/405 (7%)
Query: 121 SSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGF 180
SS ++ L+ SFF +A S +GR+ + + + I L + N+++ + L+GF
Sbjct: 85 SSLYLAALVASFFASATC-SKLGRRPTMQLASIFFLIGVGLAAGAVNIYMLIIGRILLGF 143
Query: 181 WRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNS---SWKSLYI 237
V + L+E A R + IV T+G + Y + W+
Sbjct: 144 GVGFGNQAVPLFLSEIAPARLRGGLNIVFQLMVTIGILIANIVNYFTSSIHPYGWRIALG 203
Query: 238 WSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPIL 297
+ +PA+ + L + E+P L+ + + KE + LK++ E D++ ++ I
Sbjct: 204 GAGIPALIL-LFGSLLICETPTSLIERNKTKEGKETLKKIRGVEDVDEEYESIVHACDIA 262
Query: 298 PPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGI-GLGMVYFGMPLAVGNLGFN---- 352
+ Y+ + + + VI M+ G+ + F P+ +GF
Sbjct: 263 RQVKDP-----YTKLMKPASRPPFVIGMLLQFFQQFTGINAIMFYAPVLFQTVGFGNDAA 317
Query: 353 IYLAVVFSASMELPSCVATYFLENLRRKPSILVFSILGGICCVTCAVLENRV-------- 404
+ AVV L + V + ++ R+ +L S+ IC + ++ +
Sbjct: 318 LLSAVVTGTINVLSTFVGIFLVDKTGRRFLLLQSSVHMLICQLVIGIILAKDLDVTGTLA 377
Query: 405 -PAAKVVLAMVAFFGACTAYN---VFLIYIIELFPTCVRNTTTSL-VRQAIVFGCIFCPF 459
P A VV+ V + A++ + + E FP R +L V + F +
Sbjct: 378 RPQALVVVIFVCVYVMGFAWSWGPLGWLIPSETFPLETRTEGFALAVSCNMFFTFVIAQA 437
Query: 460 LIS--AGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVLCDTMD 502
+S K+ I+ + G ++++ F LFF+PET G+ + D D
Sbjct: 438 FLSMLCAMKSGIFFF-FSGWIVVMGLFALFFVPETKGVSIDDMRD 481
>At2g16130 putative sugar transporter
Length = 511
Score = 58.9 bits (141), Expect = 7e-09
Identities = 98/400 (24%), Positives = 158/400 (39%), Gaps = 51/400 (12%)
Query: 142 IGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVLVLLTEKVSAEW 201
IGR+ ++ + ++L+ F+TN +F+ G V TE A
Sbjct: 90 IGRRYTIVLAGFFFFCGALLMGFATNYPFIMVGRFVAGIGVGYAMMIAPVYTTEVAPASS 149
Query: 202 R-FRVGIVEYFT---FTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYSVIAYLFVTES 257
R F E F +GY+S FA + + W+ + +VP++ + I L + ES
Sbjct: 150 RGFLSSFPEIFINIGILLGYVSNYFFAKLPEHIGWRFMLGIGAVPSV-FLAIGVLAMPES 208
Query: 258 PRWLVMQGREKEILKMLKRVSS---EESADDDSVNLASNLP-------ILPPKEKVSFFQ 307
PRWLVMQGR + K+L + S+ E + + + A +P I+ P +K +
Sbjct: 209 PRWLVMQGRLGDAFKVLDKTSNTKEEAISRLNDIKRAVGIPDDMTDDVIVVPNKKSAGKG 268
Query: 308 LYSSIGELFHKRWAVIRMIAVMILGI-------GLGMVYFGMPLAVGNLGF-----NIYL 355
++ +L + +R I + LGI G+ V P G +
Sbjct: 269 VWK---DLLVRPTPSVRHILIACLGIHFSQQASGIDAVVLYSPTIFSRAGLKSKNDQLLA 325
Query: 356 AVVFSASMELPSCVATYFLENLRRKPSILVFSILGGI-----CCVTCAVLENRVP----- 405
V L V T ++ R+ L+ + +GG+ T + +R P
Sbjct: 326 TVAVGVVKTLFIVVGTCLVDRFGRR--ALLLTSMGGMFFSLTALGTSLTVIDRNPGQTLK 383
Query: 406 -----AAKVVLAMVAFFGACTAYNVFLIYIIELFPTCVRNTTTSL--VRQAIVFGCIFCP 458
A V+ VA F + A V +Y E+FP +R SL + ++ G I
Sbjct: 384 WAIGLAVTTVMTFVATF-SLGAGPVTWVYASEIFPVRLRAQGASLGVMLNRLMSGIIGMT 442
Query: 459 FL-ISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVL 497
FL +S G GV + F FLPET G+ L
Sbjct: 443 FLSLSKGLTIGGAFLLFAGVAVAAWVFFFTFLPETRGVPL 482
>At1g11260 glucose transporter
Length = 522
Score = 58.9 bits (141), Expect = 7e-09
Identities = 87/405 (21%), Positives = 169/405 (41%), Gaps = 41/405 (10%)
Query: 121 SSFFIGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGF 180
SS ++ L+ S +A+ GR+ ++F + +++ F+ +VW+ + L+GF
Sbjct: 87 SSLYLAALISSL-VASTVTRKFGRRLSMLFGGILFCAGALINGFAKHVWMLIVGRILLGF 145
Query: 181 WRSSIGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLP----GFAYINRNSSWKSLY 236
V + L+E ++R + I + T+G + FA I W+
Sbjct: 146 GIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVAEVLNYFFAKIKGGWGWRLSL 205
Query: 237 IWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPI 296
+ VPA+ + I L + ++P ++ +G+ +E L+R+ + + +L +
Sbjct: 206 GGAVVPALIIT-IGSLVLPDTPNSMIERGQHEEAKTKLRRIRGVDDVSQEFDDL-----V 259
Query: 297 LPPKEKVSFFQLYSSIGELFHKRWAVIRMIAVMILGI----GLGMVYFGMPLAVGNLGF- 351
KE S + + L +++ +AVMI G+ ++ F P+ +GF
Sbjct: 260 AASKESQSIEHPWRN---LLRRKYRPHLTMAVMIPFFQQLTGINVIMFYAPVLFNTIGFT 316
Query: 352 ---NIYLAVVFSASMELPSCVATYFLENLRRKPSILVFSILGGIC------CVTCAVLEN 402
++ AVV + + V+ Y ++ R+ L IC C+ +
Sbjct: 317 TDASLMSAVVTGSVNVAATLVSIYGVDRWGRRFLFLEGGTQMLICQAVVAACIGAKFGVD 376
Query: 403 RVPA------AKVVLAMVAFFGACTAYN---VFLIYIIELFPTCVRNTTTSL-VRQAIVF 452
P A VV+ + + A A++ + + E+FP +R+ S+ V ++F
Sbjct: 377 GTPGELPKWYAIVVVTFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSITVSVNMIF 436
Query: 453 GCIFCPFLIS--AGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGI 495
I ++ K ++ F VV+M S F FLPET GI
Sbjct: 437 TFIIAQIFLTMLCHLKFGLFLVFAFFVVVM-SIFVYIFLPETKGI 480
>At5g18840 sugar transporter - like protein
Length = 482
Score = 58.5 bits (140), Expect = 9e-09
Identities = 97/399 (24%), Positives = 161/399 (40%), Gaps = 45/399 (11%)
Query: 125 IGCLLGSFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSS 184
IG +LG+ ++D S GRK + S + + F+ + +F G+
Sbjct: 92 IGAMLGAVMSGKISDFS-GRKGAMRTSACFCITGWLAVFFTKGALLLDVGRFFTGYGIGV 150
Query: 185 IGTCVLVLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAI 244
V V + E R + + +G S F I SWK+L + P I
Sbjct: 151 FSYVVPVYIAEISPKNLRGGLTTLNQLMIVIG--SSVSFL-IGSLISWKTLALTGLAPCI 207
Query: 245 CYSVIAYLFVTESPRWLVMQGREKEILKMLKRVSSEES-----ADDDSVNLASNLPILPP 299
+ F+ ESPRWL G EKE L+++ +++ AD V++ + L ILP
Sbjct: 208 VL-LFGLCFIPESPRWLAKAGHEKEFRVALQKLRGKDADITNEADGIQVSIQA-LEILPK 265
Query: 300 KEKVSFFQLYSSIGELFHKRW--AVIRMIAVMILG--IGLGMVYFGMPLAVGNLGFNI-Y 354
+ I +L K++ +VI +++M+ +G+ + F GF
Sbjct: 266 ----------ARIQDLVSKKYGRSVIIGVSLMVFQQFVGINGIGFYASETFVKAGFTSGK 315
Query: 355 LAVVFSASMELP-SCVATYFLENLRRKPSILVFS---ILGGICCVTCAVLENR------V 404
L + A +++P + + T ++ R+P I++ + LG I T +L+ + V
Sbjct: 316 LGTIAIACVQVPITVLGTILIDKSGRRPLIMISAGGIFLGCILTGTSFLLKGQSLLLEWV 375
Query: 405 PAAKV--VLAMVAFFGACTAYNVFLIYIIELFPTCVRNTTTSLVRQAIVFGC----IFCP 458
P+ V VL VA F V + + E+FP V+ SLV G
Sbjct: 376 PSLAVGGVLIYVAAFSIGMG-PVPWVIMSEIFPINVKGIAGSLVVLVNWSGAWAVSYTFN 434
Query: 459 FLISAGRKNNIYSYGVFGVVIMLSNFTLFFLPETIGIVL 497
FL+S Y Y F ++ F +PET G L
Sbjct: 435 FLMSWSSPGTFYLYSAFAAATII--FVAKMVPETKGKTL 471
>At4g04760 putative sugar transporter
Length = 457
Score = 58.2 bits (139), Expect = 1e-08
Identities = 72/294 (24%), Positives = 115/294 (38%), Gaps = 47/294 (15%)
Query: 87 LPRSSWSWDTHPSNTIISHWNLECASTFITGLPQSS---FFIGCL-------------LG 130
LP SS T S++++S + C F+ S F GC+ LG
Sbjct: 8 LPASS----TSSSSSLLSEISNACTRPFVLAFIVGSCGAFAFGCIYSLFGSILTVGLILG 63
Query: 131 SFFLAALADSSIGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKFLIGFWRSSIGTCVL 190
+ L D +GR + + + I I F+ VW+ + L G SIG V
Sbjct: 64 ALICGKLTDL-VGRVKTIWITNILFVIGWFAIAFAKGVWLLDLGRLLQGI---SIGISVY 119
Query: 191 ---VLLTEKVSAEWRFRVGIVEYFTFTMGYMSLPGFAYINRNSSWKSLYIWSSVPAICYS 247
V +TE R G F + + F + +W++L I +P++
Sbjct: 120 LGPVYITEIAPRNLR---GAASSFAQLFAGVGISVFYALGTIVAWRNLAILGCIPSLMVL 176
Query: 248 VIAYLFVTESPRWLVMQGREKEILKMLKRVSSEESADDDSVNLASNLPILPPKEKVSFFQ 307
+ + F+ ESPRWL GRE E+ +L + E+S D IL E V Q
Sbjct: 177 PLLF-FIPESPRWLAKVGREMEVEAVLLSLRGEKSDVSDEA-----AEILEYTEHVKQQQ 230
Query: 308 LYSSIG--ELFHKRWAVIRMIAVMILGIGLGMVYFGMPLAVGNLGFNIYLAVVF 359
G +LF +++A + +G+V +P G G++ Y +F
Sbjct: 231 DIDDRGFFKLFQRKYA---------FSLTIGVVLIALPQLGGLNGYSFYTDSIF 275
>At3g18830 sugar transporter like protein
Length = 539
Score = 58.2 bits (139), Expect = 1e-08
Identities = 101/430 (23%), Positives = 177/430 (40%), Gaps = 59/430 (13%)
Query: 125 IGCLLGSFFLAALADSS--------IGRKNMLIFSCVSMSITSMLIIFSTNVWIYSALKF 176
IG L GS + +L S IGR+ ++ + ++L+ S N +F
Sbjct: 75 IGILAGSLNIYSLIGSCAAGRTSDWIGRRYTIVLAGAIFFAGAILMGLSPNYAFLMFGRF 134
Query: 177 LIGFWRSSIGTCVLV--LLTEKVS--AEWRFRVGIVEYFT---FTMGYMSLPGFAYINRN 229
+ G +G +++ + T +VS + F E F +GY+S F+ +
Sbjct: 135 IAGI---GVGYALMIAPVYTAEVSPASSRGFLNSFPEVFINAGIMLGYVSNLAFSNLPLK 191
Query: 230 SSWKSLYIWSSVPAICYSVIAYLFVTESPRWLVMQGREKEILKMLKRVS---SEESADDD 286
W+ + +VP++ + I L + ESPRWLVMQGR + ++L + S +E + +
Sbjct: 192 VGWRLMLGIGAVPSVILA-IGVLAMPESPRWLVMQGRLGDAKRVLDKTSDSPTEATLRLE 250
Query: 287 SVNLASNLPILPPKEKVSFFQLYSS----IGELFHKRWAVIRMIAVMILGI-------GL 335
+ A+ +P + V + S EL + +R + + +GI G+
Sbjct: 251 DIKHAAGIPADCHDDVVQVSRRNSHGEGVWRELLIRPTPAVRRVMIAAIGIHFFQQASGI 310
Query: 336 GMVYFGMPLAVGNLGF-----NIYLAVVFSASMELPSCVATYFLENLRRKPSILVFSILG 390
V P G + V VAT+ L+ + R+P L+ + +G
Sbjct: 311 DAVVLFSPRIFKTAGLKTDHQQLLATVAVGVVKTSFILVATFLLDRIGRRP--LLLTSVG 368
Query: 391 GICCVTCA--------------VLENRVPAAKVVLAMVAFFGACTAYNVFLIYIIELFPT 436
G+ A V+ V A V+ VA F + A + +Y E+FP
Sbjct: 369 GMVLSLAALGTSLTIIDQSEKKVMWAVVVAIATVMTYVATF-SIGAGPITWVYSSEIFPL 427
Query: 437 CVRNTTTSL--VRQAIVFGCIFCPFLISAGRKNNIYSYGVFGVVIMLS-NFTLFFLPETI 493
+R+ +S+ V + G I FL + ++ +FG + ++ F FLPET
Sbjct: 428 RLRSQGSSMGVVVNRVTSGVISISFLPMSKAMTTGGAFYLFGGIATVAWVFFYTFLPETQ 487
Query: 494 GIVLCDTMDQ 503
G +L D MD+
Sbjct: 488 GRMLED-MDE 496
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.138 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,501,175
Number of Sequences: 26719
Number of extensions: 469945
Number of successful extensions: 1730
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 39
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 1603
Number of HSP's gapped (non-prelim): 97
length of query: 526
length of database: 11,318,596
effective HSP length: 104
effective length of query: 422
effective length of database: 8,539,820
effective search space: 3603804040
effective search space used: 3603804040
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC121234.10


