

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121233.5 - phase: 0
(301 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
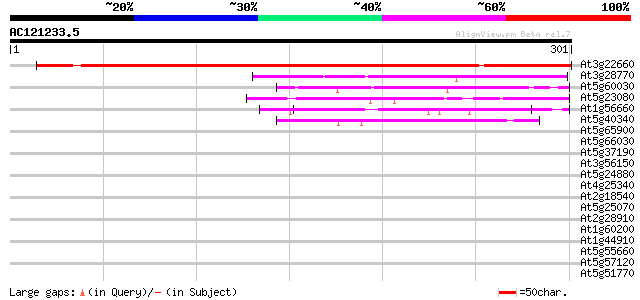
Sequences producing significant alignments: (bits) Value
At3g22660 unknown protein 362 e-100
At3g28770 hypothetical protein 45 4e-05
At5g60030 KED - like protein 44 1e-04
At5g23080 unknown protein 44 1e-04
At1g56660 hypothetical protein 43 2e-04
At5g40340 unknown protein 42 3e-04
At5g65900 ATP-dependent RNA helicase-like 40 0.001
At5g66030 Golgi-localized protein GRIP 40 0.002
At5g37190 COP1-interacting protein 4 (CIP4) 39 0.003
At3g56150 PROBABLE EUKARYOTIC TRANSLATION INITIATION FACTOR 3 SU... 39 0.003
At5g24880 glutamic acid-rich protein 39 0.003
At4g25340 unknown protein 39 0.003
At2g18540 putative vicilin storage protein (globulin-like) 39 0.003
At5g25070 unknown protein 39 0.004
At2g28910 unknown protein 39 0.004
At1g60200 38 0.006
At1g44910 splicing factor like protein 38 0.008
At5g55660 putative protein 37 0.010
At5g57120 unknown protein 37 0.017
At5g51770 putative protein 37 0.017
>At3g22660 unknown protein
Length = 293
Score = 362 bits (929), Expect = e-100
Identities = 185/288 (64%), Positives = 229/288 (79%), Gaps = 7/288 (2%)
Query: 15 ELPLVNDDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPE 74
E+ ++++D D EAE LS+ S++E+E KLAEP+KTA+ NRD LLDKL DISWPE
Sbjct: 12 EMNMIDEDDATDSEAESLSD----SDTENEITEKLAEPTKTAIYNRDGLLDKLQDISWPE 67
Query: 75 NVEWIHKLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYA 134
+V+W HKL+++IDQ VDVNDDLARE AFYTQALEGTR+AF KL +G+ FLRPA+YYA
Sbjct: 68 DVDWTHKLTVEIDQGGAVDVNDDLARETAFYTQALEGTREAFGKLNEMGVNFLRPANYYA 127
Query: 135 EMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIES 194
EMVK+D HMEKVKSRLL EK+++ E+EERRKAR+ KR++KEVQSQK+KERAK+KKD+IES
Sbjct: 128 EMVKSDVHMEKVKSRLLHEKKQIEESEERRKARDNKRMAKEVQSQKMKERAKEKKDNIES 187
Query: 195 VKKWRKQRQQSGFADDGADKALDFEDGKVFER-SKKKRPGVSPGDRSGGKAKQAFGKGKK 253
VKKWRKQRQQSGF+D + LDFE GK F+R KKRPGVSPGDRSGGK + G
Sbjct: 188 VKKWRKQRQQSGFSDKAGEPELDFESGKSFQRGGGKKRPGVSPGDRSGGKGRPTSRMGN- 246
Query: 254 PKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKKRKR 301
KK + ++SKFG GG+KGL KQNTA+TTNDF G + G G+K++KR
Sbjct: 247 -KKREFRDSKFGHGGRKGLSKQNTAETTNDFKGGFRGGKASGNKRQKR 293
>At3g28770 hypothetical protein
Length = 2081
Score = 45.4 bits (106), Expect = 4e-05
Identities = 40/173 (23%), Positives = 80/173 (46%), Gaps = 6/173 (3%)
Query: 131 DYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKD 190
+Y + KT +K K + +K++ ++EER+ +E K S++++++K +E K+KK+
Sbjct: 1014 EYEEKKSKTKEEAKKEKKKSQDKKREEKDSEERKSKKE-KEESRDLKAKKKEEETKEKKE 1072
Query: 191 DIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGD----RSGGKAKQ 246
E+ K +K+ ++ + K D ++ K E SK ++ D K+
Sbjct: 1073 S-ENHKSKKKEDKKEHEDNKSMKKEEDKKEKKKHEESKSRKKEEDKKDMEKLEDQNSNKK 1131
Query: 247 AFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKKR 299
K +K K VK K K+ + + ++T S+K V +K+
Sbjct: 1132 KEDKNEKKKSQHVKLVKKESDKKEKKENEEKSETKEIESSKSQKNEVDKKEKK 1184
Score = 38.9 bits (89), Expect = 0.003
Identities = 37/145 (25%), Positives = 65/145 (44%), Gaps = 14/145 (9%)
Query: 138 KTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKK 197
K + EK KS+ + +K + +E+ K E K +KE++S K ++ KK+ S +
Sbjct: 1131 KKEDKNEKKKSQHVKLVKKESDKKEK-KENEEKSETKEIESSKSQKNEVDKKEKKSSKDQ 1189
Query: 198 WRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKG 257
+K+ ++ + +K L K E +KK+ V K ++ K K K
Sbjct: 1190 QKKKEKE---MKESEEKKL-----KKNEEDRKKQTSVEE-----NKKQKETKKEKNKPKD 1236
Query: 258 DVKNSKFGFGGKKGLKKQNTADTTN 282
D KN+ GGKK + + + N
Sbjct: 1237 DKKNTTKQSGGKKESMESESKEAEN 1261
Score = 35.0 bits (79), Expect = 0.050
Identities = 49/273 (17%), Positives = 107/273 (38%), Gaps = 40/273 (14%)
Query: 33 SEEMSPSESESEEDVKLAEPSK----TAVNNRDALLDKLGDISWPENVEWIHKLSIDIDQ 88
SE++ E + +D K E ++ NRD ++ G+ + + E S++ +
Sbjct: 782 SEKVEKGEKKESKDAKSVETKDNKKLSSTENRDEAKERSGEDNKEDKEESKDYQSVEAKE 841
Query: 89 EQE--------------VDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYA 134
+ E D+ DD + E+ + E ++ E++Q + +A
Sbjct: 842 KNENGGVDTNVGNKEDSKDLKDDRSVEVKANKE--ESMKKKREEVQRNDKSSTKEVRDFA 899
Query: 135 EMVKTDSHM----------EKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKER 184
+ D ++ K E + + ++K ++ K+ KE ++ +K++
Sbjct: 900 NNMDIDVQKGSGESVKYKKDEKKEGNKEENKDTINTSSKQKGKDKKKKKKESKNSNMKKK 959
Query: 185 AKQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSK-KKRPGVSPGDRSGGK 243
+ KK+ + + + +KQ +D + E+ K+ E +K K S S +
Sbjct: 960 EEDKKEYVNN--ELKKQ-------EDNKKETTKSENSKLKEENKDNKEKKESEDSASKNR 1010
Query: 244 AKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQN 276
K+ + + K K + K K KK +K +
Sbjct: 1011 EKKEYEEKKSKTKEEAKKEKKKSQDKKREEKDS 1043
Score = 33.5 bits (75), Expect = 0.14
Identities = 28/126 (22%), Positives = 54/126 (42%), Gaps = 11/126 (8%)
Query: 138 KTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKK 197
K +S +K + E +K E +K + K+ + + ++ KLKE K K+ ES
Sbjct: 948 KKESKNSNMKKK--EEDKKEYVNNELKKQEDNKKETTKSENSKLKEENKDNKEKKESEDS 1005
Query: 198 WRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKG 257
K R++ + + + K E +KK++ R +++ K +K +
Sbjct: 1006 ASKNREKKEYEE---------KKSKTKEEAKKEKKKSQDKKREEKDSEERKSKKEKEESR 1056
Query: 258 DVKNSK 263
D+K K
Sbjct: 1057 DLKAKK 1062
Score = 32.0 bits (71), Expect = 0.42
Identities = 41/177 (23%), Positives = 67/177 (37%), Gaps = 44/177 (24%)
Query: 144 EKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQ---KKDDIESVKKWRK 200
EK + +++K EE +K +E K+ + + K K KQ KK+ +ES K +
Sbjct: 1202 EKKLKKNEEDRKKQTSVEENKKQKETKKEKNKPKDDK-KNTTKQSGGKKESMESESKEAE 1260
Query: 201 QRQQS--------------------GFADDGADKALDFEDGK---------------VFE 225
+Q+S AD +D D ++ K E
Sbjct: 1261 NQQKSQATTQADSDESKNEILMQADSQADSHSDSQADSDESKNEILMQADSQATTQRNNE 1320
Query: 226 RSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTN 282
+KK+ V+ K K+ + KP K D KN+ GGKK + + + N
Sbjct: 1321 EDRKKQTSVA----ENKKQKETKEEKNKP-KDDKKNTTKQSGGKKESMESESKEAEN 1372
Score = 29.6 bits (65), Expect = 2.1
Identities = 54/270 (20%), Positives = 101/270 (37%), Gaps = 23/270 (8%)
Query: 35 EMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPENVEWIHKLSIDIDQEQEVDV 94
+ + S+ S ++ V+ D LD +G +N D+ EV
Sbjct: 528 DANESDGNSTKERHQEAQVNNGVSTEDKNLDNIGADEQKKN-----------DKSVEVTT 576
Query: 95 ND-DLARELAFYTQALEGTRQAFEKLQSI-GLPFLRPADYYAEMVKTDSHMEKVKSRLLA 152
ND D +E TQ G E L++ L+ + ++ +E+ + +
Sbjct: 577 NDGDHTKEKREETQGNNGESVKNENLENKEDKKELKDDESVGAKTNNETSLEEKREQTQK 636
Query: 153 EKQKMVEAE--ERRKAREAKRLSKEVQ-SQKLKERAKQKKDDIESVKKWRKQRQQSGFAD 209
+ ++ + + KEV + + K+D +S + +K S +
Sbjct: 637 GHDNSINSKIVDNKGGNADSNKEKEVHVGDSTNDNNMESKEDTKSEVEVKKNDGSSEKGE 696
Query: 210 DGADKALDFEDGKVFERSKKKRPGVSPGDRS-GGKAKQAFGKGKKPKKGDVKNSKFGFGG 268
+G + D + K E + + S D+S K ++A G + K +K G
Sbjct: 697 EGKENNKDSMEDKKLENKESQTD--SKDDKSVDDKQEEAQIYGGESKDDKSVEAK----G 750
Query: 269 KKGLKKQNTADTTNDFGGFSKKGAVGGSKK 298
KK K+N TN+ +K+ V G+KK
Sbjct: 751 KKKESKENKKTKTNENRVRNKEENVQGNKK 780
Score = 29.6 bits (65), Expect = 2.1
Identities = 19/74 (25%), Positives = 35/74 (46%), Gaps = 9/74 (12%)
Query: 20 NDDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPENVEWI 79
N++ ++G+ E + + ES+EDVK + A N+ ++ + LG+ V
Sbjct: 272 NNEKEVEGQGESIGDSAIEKNLESKEDVK--SEVEAAKNDGSSMTENLGEAQGNNGVS-- 327
Query: 80 HKLSIDIDQEQEVD 93
ID E+EV+
Sbjct: 328 -----TIDNEKEVE 336
Score = 29.3 bits (64), Expect = 2.7
Identities = 64/303 (21%), Positives = 121/303 (39%), Gaps = 45/303 (14%)
Query: 19 VNDDTMIDGEAEYLSEEMSPSESESEEDVKL-AEPSKTAVNNRDALLDKL---GDISWPE 74
++++ ++G+ E + + ES+EDVK E +K A ++ L++ +S E
Sbjct: 329 IDNEKEVEGQGESIEDSDIEKNLESKEDVKSEVEAAKNAGSSMTGKLEEAQRNNGVSTNE 388
Query: 75 NVEWIHKLSIDIDQEQEVDV---NDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPAD 131
+ +K S + ++ V+ ++D +E T G E L++
Sbjct: 389 TMNSENKGSGESTNDKMVNATTNDEDHKKENKEETHENNGESVKGENLEN---KAGNEES 445
Query: 132 YYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKR-----LSKE--------VQS 178
E ++ E++K E + E+ + K E++R ++KE +Q
Sbjct: 446 MKGENLENKVGNEELKGNASVEAKTNNESSKEEKREESQRSNEVYMNKETTKGENVNIQG 505
Query: 179 QKL-----------KERAKQKKDDIESVKKWRKQRQQSGFADDGA---DKALD-FEDGKV 223
+ + KE K K D ES K+R Q ++G DK LD +
Sbjct: 506 ESIGDSTKDNSLENKEDVKPKVDANESDGNSTKERHQEAQVNNGVSTEDKNLDNIGADEQ 565
Query: 224 FERSKKKRPGVSPGDRSGGKAKQAFG-KGKKPKKGDVKNSKFGFGGKKGLKKQNT--ADT 280
+ K + GD + K ++ G G+ K +++N + KK LK + A T
Sbjct: 566 KKNDKSVEVTTNDGDHTKEKREETQGNNGESVKNENLENKE----DKKELKDDESVGAKT 621
Query: 281 TND 283
N+
Sbjct: 622 NNE 624
Score = 29.3 bits (64), Expect = 2.7
Identities = 26/141 (18%), Positives = 61/141 (42%), Gaps = 7/141 (4%)
Query: 140 DSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWR 199
D+ E KS K++ + +E++++ ++ SK + ++ +E+ + K++ + KK
Sbjct: 976 DNKKETTKSENSKLKEENKDNKEKKESEDSA--SKNREKKEYEEKKSKTKEEAKKEKKKS 1033
Query: 200 KQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGK-----GKKP 254
+ +++ + + E+ + + KK+ + K+K+ K K
Sbjct: 1034 QDKKREEKDSEERKSKKEKEESRDLKAKKKEEETKEKKESENHKSKKKEDKKEHEDNKSM 1093
Query: 255 KKGDVKNSKFGFGGKKGLKKQ 275
KK + K K K KK+
Sbjct: 1094 KKEEDKKEKKKHEESKSRKKE 1114
Score = 28.5 bits (62), Expect = 4.6
Identities = 38/168 (22%), Positives = 62/168 (36%), Gaps = 44/168 (26%)
Query: 153 EKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQ---KKDDIESVKKWRKQRQQS---- 205
+++K E +K +E K + + K K KQ KK+ +ES K + +Q+S
Sbjct: 1322 DRKKQTSVAENKKQKETKEEKNKPKDDK-KNTTKQSGGKKESMESESKEAENQQKSQATT 1380
Query: 206 ----------------GFADDGADKALDFEDGK---------------VFERSKKKRPGV 234
AD +D D ++ K E +KK+ V
Sbjct: 1381 QADSDESKNEILMQADSQADSHSDSQADSDESKNEILMQADSQATTQRNNEEDRKKQTSV 1440
Query: 235 SPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTN 282
+ K K+ + KP K D KN+ GGKK + + + N
Sbjct: 1441 A----ENKKQKETKEEKNKP-KDDKKNTTEQSGGKKESMESESKEAEN 1483
>At5g60030 KED - like protein
Length = 292
Score = 43.5 bits (101), Expect = 1e-04
Identities = 42/164 (25%), Positives = 80/164 (48%), Gaps = 15/164 (9%)
Query: 144 EKVKSRLLAEKQKMVEAEERRKAREAKRLSK--EVQSQKLKERAKQKKDDIESVKKWRKQ 201
EKV +L AE Q+ E ER+K ++ K+ +K +V +K+KE+ + ++ + K+ +K+
Sbjct: 131 EKVNEKLEAE-QRSEERRERKKEKKKKKNNKDEDVVDEKVKEKLEDEQKSADR-KERKKK 188
Query: 202 RQQSGFADDGADKALDFEDGKVFERSKKKRPG-----VSPGDRSGGKAKQAFGKGKKPKK 256
+ + +D D+ ED + K+K+ V ++ + +Q G+ KK KK
Sbjct: 189 KSKKNNDEDVVDEKEKLEDEQKSAEIKEKKKNKDEDVVDEKEKEKLEDEQRSGERKKEKK 248
Query: 257 GDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKKRK 300
K+ + ++ KK+ +D + G +K KKRK
Sbjct: 249 KKRKSDEEIVSEERKSKKKRKSD--EEMGSEERK----SKKKRK 286
Score = 40.4 bits (93), Expect = 0.001
Identities = 38/130 (29%), Positives = 63/130 (48%), Gaps = 20/130 (15%)
Query: 144 EKVKSRLLAEKQKMVEAEERRKAR-----------EAKRLSKEVQSQKLKERAKQKKDDI 192
EKVK +L ++QK + +ER+K + E ++L E +S ++KE+ K K +D+
Sbjct: 167 EKVKEKL-EDEQKSADRKERKKKKSKKNNDEDVVDEKEKLEDEQKSAEIKEKKKNKDEDV 225
Query: 193 --ESVKKWRKQRQQSGFADDGADKALDFEDGKVFE--RSKKKRPGVSPGDRSGGKAKQAF 248
E K+ + Q+SG K ++ V E +SKKKR D G ++
Sbjct: 226 VDEKEKEKLEDEQRSGERKKEKKKKRKSDEEIVSEERKSKKKR----KSDEEMGSEERKS 281
Query: 249 GKGKKPKKGD 258
K +K K+ D
Sbjct: 282 KKKRKLKEID 291
>At5g23080 unknown protein
Length = 930
Score = 43.5 bits (101), Expect = 1e-04
Identities = 43/183 (23%), Positives = 76/183 (41%), Gaps = 21/183 (11%)
Query: 128 RPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQ 187
RP D Y + DS ++ + K+ E +E++ A L++ + L+ K+
Sbjct: 715 RPVDLYKAIFSDDSEDDEDQPM----NGKIQEGQEKKNEAAATTLNRLIAGDFLESLGKE 770
Query: 188 KKDDI---ESVKKWRKQRQQS-------GFADDGADKALDFEDGKVFERSKKKRPGVSPG 237
++ E +K K S G + +K G E+S+KKR SPG
Sbjct: 771 LGFEVPMEEEIKSRSKPEDSSDKRLDRPGLKEKVEEKTSSLTLGSEEEKSRKKREK-SPG 829
Query: 238 DRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSK 297
RSGG GD + K + K + + +D+++D+ K+G+ SK
Sbjct: 830 KRSGGN-----DLSSSESSGDERRRK-RYNKKDRHRNDSESDSSSDYHSRDKQGSRSRSK 883
Query: 298 KRK 300
+R+
Sbjct: 884 RRE 886
>At1g56660 hypothetical protein
Length = 522
Score = 42.7 bits (99), Expect = 2e-04
Identities = 39/155 (25%), Positives = 71/155 (45%), Gaps = 15/155 (9%)
Query: 135 EMVKTDSHMEKVKSR---LLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDD 191
EM + DS K K + EK+K + E++ K ++ K+++ +K K +K+D+
Sbjct: 241 EMKEKDSKKNKKKEKDESCAEEKKKKPDKEKKEKDESTEKEDKKLKGKKGKGEKPEKEDE 300
Query: 192 IESVKKWRKQRQQSGFADDGADKALDFEDGKV---FERSKKKRPGVSPGDRSGGKAK--- 245
+K ++ + D+A D ++GK +++KKK + K K
Sbjct: 301 ------GKKTKEHDATEQEMDDEAADHKEGKKKKNKDKAKKKETVIDEVCEKETKDKDDD 354
Query: 246 QAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADT 280
+ K KK KK + K+ K K+ KK+N +T
Sbjct: 355 EGETKQKKNKKKEKKSEKGEKDVKEDKKKENPLET 389
Score = 42.4 bits (98), Expect = 3e-04
Identities = 40/153 (26%), Positives = 68/153 (44%), Gaps = 18/153 (11%)
Query: 153 EKQKMVEAEERRKAREAKRLSKEVQSQKLK-ERAKQKKDDIESVKKWRKQRQQSGFADDG 211
EK+K + E+++K K+V+ +K K E+ +K+D E K+ + Q+ D
Sbjct: 189 EKKKKPKKEKKQKEESKSNEDKKVKGKKEKGEKGDLEKEDEEKKKEHDETDQEMKEKDSK 248
Query: 212 ADKALDFEDGKVFERSKK----KRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFG 267
+K + ++ E+ KK K+ ++ K K GKG+KP+K D
Sbjct: 249 KNKKKEKDESCAEEKKKKPDKEKKEKDESTEKEDKKLKGKKGKGEKPEKED--------E 300
Query: 268 GKKGLKKQNTADTTNDFGGFSKKGAVGGSKKRK 300
GKK + T +D K+G KK+K
Sbjct: 301 GKKTKEHDATEQEMDDEAADHKEG-----KKKK 328
Score = 40.0 bits (92), Expect = 0.002
Identities = 33/142 (23%), Positives = 64/142 (44%), Gaps = 3/142 (2%)
Query: 161 EERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKA-LDFE 219
+E +K +E K E + + K++ K++KD+ +K +K ++ D +K L+ E
Sbjct: 113 KEHKKGKEKKHEELEEEKEGKKKKNKKEKDESGPEEKNKKADKEKKHEDVSQEKEELEEE 172
Query: 220 DGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTAD 279
DGK ++ +K G + K K+ + K + VK K G+KG ++ +
Sbjct: 173 DGKKNKKKEKDESGTEEKKKKPKKEKKQKEESKSNEDKKVKGKK--EKGEKGDLEKEDEE 230
Query: 280 TTNDFGGFSKKGAVGGSKKRKR 301
+ ++ SKK K+
Sbjct: 231 KKKEHDETDQEMKEKDSKKNKK 252
Score = 39.3 bits (90), Expect = 0.003
Identities = 31/116 (26%), Positives = 53/116 (44%), Gaps = 9/116 (7%)
Query: 135 EMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIES 194
E++ D +E+ + AEK++ + EE++K++ S+E + +K K++ K KK D +
Sbjct: 390 EVMSRDIKLEEPE----AEKKEEDDTEEKKKSKVEGGESEEGKKKKKKDKKKNKKKDTKE 445
Query: 195 VKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGK 250
K + ++ DD D + E K E K K G GK K K
Sbjct: 446 PKMTEDEEEKK---DDSKD--VKIEGSKAKEEKKDKDVKKKKGGNDIGKLKTKLAK 496
Score = 38.5 bits (88), Expect = 0.004
Identities = 45/229 (19%), Positives = 98/229 (42%), Gaps = 23/229 (10%)
Query: 33 SEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPENVEWIHKLSIDIDQEQEV 92
++E+ P E +V++ +K+ + A D+ + S + K ++D E +
Sbjct: 19 TQELDPKEKGENVEVEMEVKAKS-IEKVKAKKDE--ESSGKSKKDKEKKKGKNVDSEVKE 75
Query: 93 DVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRLLA 152
D +DD ++ ++ E E +S VK + H ++ K
Sbjct: 76 DKDDDKKKDGKMVSKKHEEGHGDLEVKESD--------------VKVEEHEKEHKKGKEK 121
Query: 153 EKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKK-DDIESVKKWRKQRQQSGFADDG 211
+ +++ E +E +K + K + +K K+ K+KK +D+ K+ + ++ G +
Sbjct: 122 KHEELEEEKEGKKKKNKKEKDESGPEEKNKKADKEKKHEDVSQEKE--ELEEEDGKKNKK 179
Query: 212 ADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVK 260
+K E G ++ K K+ + + K+ GK +K +KGD++
Sbjct: 180 KEKD---ESGTEEKKKKPKKEKKQKEESKSNEDKKVKGKKEKGEKGDLE 225
Score = 32.3 bits (72), Expect = 0.32
Identities = 28/128 (21%), Positives = 56/128 (42%), Gaps = 8/128 (6%)
Query: 138 KTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKK 197
K + EK + + +K+K E +R+ K E + ++ + ++KK +E +
Sbjct: 365 KKEKKSEKGEKDVKEDKKKENPLETEVMSRDIKLEEPEAEKKEEDDTEEKKKSKVEGGES 424
Query: 198 WRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKG 257
++++ D +K D ++ K+ E ++K+ G KAK+ + K
Sbjct: 425 EEGKKKKK--KDKKKNKKKDTKEPKMTEDEEEKKDDSKDVKIEGSKAKE------EKKDK 476
Query: 258 DVKNSKFG 265
DVK K G
Sbjct: 477 DVKKKKGG 484
Score = 31.2 bits (69), Expect = 0.72
Identities = 25/128 (19%), Positives = 55/128 (42%), Gaps = 3/128 (2%)
Query: 159 EAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDF 218
+ + ++K +++++ K+V+ K KE + + +K + ++ DD +K
Sbjct: 360 QKKNKKKEKKSEKGEKDVKEDKKKENPLETEVMSRDIKLEEPEAEKKE-EDDTEEKKKSK 418
Query: 219 EDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTA 278
+G E KKK+ ++ + + ++ KK D K+ K G K +++
Sbjct: 419 VEGGESEEGKKKKKKDKKKNKKKDTKEPKMTEDEEEKKDDSKDVK--IEGSKAKEEKKDK 476
Query: 279 DTTNDFGG 286
D GG
Sbjct: 477 DVKKKKGG 484
>At5g40340 unknown protein
Length = 1008
Score = 42.4 bits (98), Expect = 3e-04
Identities = 38/150 (25%), Positives = 68/150 (45%), Gaps = 12/150 (8%)
Query: 144 EKVKSRLLAEKQKMVEAEERRKAREAKRLSKE-----VQSQKLKERAKQ---KKDDIESV 195
+K + R +E +K + EE + ++ KE +S+K E ++ +K+ +ES
Sbjct: 758 KKERKRKKSESKKQSDGEEETQKEPSESTKKERKRKNPESKKKAEAVEEEETRKESVEST 817
Query: 196 KKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKK-P 254
KK RK+++ ++ ++ E K +R K + + + SG + FG G P
Sbjct: 818 KKERKRKKPKHDEEEVPNETEKPEKKKKKKREGKSKKKETETEFSGAELYVTFGPGSSLP 877
Query: 255 KKGDVKNSKFGFGGKKGLKKQNTADTTNDF 284
KK D+ FG L K+ T N+F
Sbjct: 878 KKEDLIEIYEKFG---ALDKERTDTVDNNF 904
Score = 35.0 bits (79), Expect = 0.050
Identities = 26/100 (26%), Positives = 44/100 (44%), Gaps = 10/100 (10%)
Query: 156 KMVEAEERRKAR-EAKRLSKEVQSQKLKERAKQKKDD--------IESVKKWRKQRQQSG 206
+++E E K R E + +V+S R K K+ D + + WR+++ +
Sbjct: 386 EIIEKESAAKVRFETEPADGDVKSNVKSGRKKTKRHDEVNGDLENVTTTALWRRRKSEVA 445
Query: 207 FADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQ 246
+DG +K + E K KKK+ V GD G K+
Sbjct: 446 TIEDGGNKQV-VESSKGKTSRKKKKMDVDDGDDDGSGDKE 484
Score = 31.2 bits (69), Expect = 0.72
Identities = 49/244 (20%), Positives = 97/244 (39%), Gaps = 16/244 (6%)
Query: 32 LSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPENV---EWIHKLSIDID- 87
+ E S S+S+ D ++ + S + + + LDK + S + E +LS +D
Sbjct: 559 IESESSKVSSQSQVDERVTDASDSLMEVEEDTLDKPCEPSSDNGLGQEELSRELSNAVDF 618
Query: 88 -----QEQEVDVNDDLARELAFYTQALE--GTRQAFEKLQSIGLPFLRPADYYAEMVKTD 140
+E+ DL R A TQ + +R + +I F + + +
Sbjct: 619 LRLGATPKEMQ---DLIRVAALGTQYPKDSSSRDMVREFMTIYRSFTYHDGANHKFLGSY 675
Query: 141 SHMEKVKSRLLAEKQKMVEAEERR-KAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWR 199
+K K L + + + +E++ K +AK+ ++E++ +E K ++ +K R
Sbjct: 676 DSSDKEKEELSEMGKPVTKGKEKKDKKGKAKQKAEEIEVTGKEENETDKHGKMKKERK-R 734
Query: 200 KQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDV 259
K+ + +G + + + ER +KK D K+ KK +K
Sbjct: 735 KKSESKKEGGEGEETQKEANESTKKERKRKKSESKKQSDGEEETQKEPSESTKKERKRKN 794
Query: 260 KNSK 263
SK
Sbjct: 795 PESK 798
Score = 31.2 bits (69), Expect = 0.72
Identities = 27/125 (21%), Positives = 48/125 (37%), Gaps = 14/125 (11%)
Query: 159 EAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDF 218
+ E+ + K ++K + + K +AKQK ++IE K + + G
Sbjct: 679 DKEKEELSEMGKPVTKGKEKKDKKGKAKQKAEEIEVTGKEENETDKHGKMKK-------- 730
Query: 219 EDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTA 278
ER +KK G K+A KK +K SK G++ +K+ +
Sbjct: 731 ------ERKRKKSESKKEGGEGEETQKEANESTKKERKRKKSESKKQSDGEEETQKEPSE 784
Query: 279 DTTND 283
T +
Sbjct: 785 STKKE 789
>At5g65900 ATP-dependent RNA helicase-like
Length = 633
Score = 40.4 bits (93), Expect = 0.001
Identities = 29/118 (24%), Positives = 58/118 (48%), Gaps = 9/118 (7%)
Query: 152 AEKQKMVEAEERRKAR-EAKRLSKEVQSQKLKERAKQKKD-----DIESVKKWRKQRQQS 205
+E +++ + + +++AR EAK+L + ++ K+ D E KK +K+ ++
Sbjct: 11 SENEEIKKKKHKKRARDEAKKLKQPAMEEEPDHEDGDAKENNALIDEEPKKKKKKKNKKR 70
Query: 206 GFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSK 263
G DDG D+A+ E+ K + KKK + + + + + ++PKK K K
Sbjct: 71 GDTDDGEDEAVAEEEPK---KKKKKNKKLQQRGDTNDEEDEVIAEEEEPKKKKKKQRK 125
>At5g66030 Golgi-localized protein GRIP
Length = 788
Score = 40.0 bits (92), Expect = 0.002
Identities = 55/233 (23%), Positives = 101/233 (42%), Gaps = 32/233 (13%)
Query: 14 MELPLVNDDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWP 73
ME + + AE L E++ +SE+E++ + E S DAL KL
Sbjct: 331 MEAAATGEAARLRAAAETLKGELAHLKSENEKEKETWEASC------DALKSKL------ 378
Query: 74 ENVEWIHKLSIDIDQEQEVDVNDDLARELAFYTQAL-------EGTRQAFEKLQSIGLPF 126
+ + L +I+ + + L E++ TQ L +G R+ +LQS +
Sbjct: 379 -EIAESNYLQAEIEVAK---MRSQLGSEMSMQTQILSTKDAELKGAREEINRLQSEFSSY 434
Query: 127 -------LRPADYYAEMVKTDSHMEKVKSRLL-AEKQKMVEAEERRKAREAKRLSKEVQS 178
L+ D K ++ ++ L AEK+ + + ER +A++ + +
Sbjct: 435 KIRAHALLQKKDMELAAAKDSEQIKSLEEALKEAEKEVYLVSAERDRAQQDLQSALASLE 494
Query: 179 QKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKR 231
++L+ERA KD E +K + + S A + A+K ED +V E + ++R
Sbjct: 495 KELEERAGALKDASEQIKS-LEVKLDSTVARNQAEKQAWEEDLRVLEETWRRR 546
Score = 30.0 bits (66), Expect = 1.6
Identities = 43/178 (24%), Positives = 79/178 (44%), Gaps = 21/178 (11%)
Query: 32 LSEEMSPSESESEEDVKLAEPSKT---AVNN--RDALLDKLGDISWPEN-VEWIHKLSID 85
+ +E+ + ++ E +K + + + NN RD + + G + EN +E + + +D
Sbjct: 200 MQQELERTRQQANEALKAMDAERQQLRSANNKLRDTIEELRGSLQPKENKIETLQQSLLD 259
Query: 86 IDQEQEVDVNDDLARELAFYTQALEGTRQ-AFEKLQSIGLPFLR--PADYYAEMVKTDSH 142
DQ + +DL ++L QA+E +Q A +L + L A + + D
Sbjct: 260 KDQ-----ILEDLKKQL----QAVEERKQIAVTELSAKHQKNLEGLEAQVVDALSERDKA 310
Query: 143 MEKVKSR--LLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKW 198
E + S LLAEK+ + E EA RL ++ K E A K ++ + + W
Sbjct: 311 AETISSLQVLLAEKESKIAEMEAAATGEAARLRAAAETLK-GELAHLKSENEKEKETW 367
>At5g37190 COP1-interacting protein 4 (CIP4)
Length = 876
Score = 39.3 bits (90), Expect = 0.003
Identities = 64/293 (21%), Positives = 119/293 (39%), Gaps = 32/293 (10%)
Query: 22 DTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKT--AVNNRDALLDKLGDISWPENVEWI 79
D + GEA + + E ++ ++++A + T A LD +G ++ E+
Sbjct: 386 DKVGSGEATLIGAD----EIQAASNLQVAGMASTPHAFVQESKTLDHIGKVTDTEHKVPQ 441
Query: 80 HKLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKT 139
++ ID DQ + V + A T A E T L G + P ++V
Sbjct: 442 ERVEIDADQAKSVKSTKKKSSRKA-KTPAKEDT------LVDFGAQNVEPI----KVVDG 490
Query: 140 DSHMEKVKSRLLAEKQKM-VEAEERRKAREAKRLSKEVQSQKLKERAK------QKKDDI 192
+ H+ +++ L + +Q+ VE + +++ + SK+ S + E A+ QKK+
Sbjct: 491 EGHVNDIRNVLDSLQQRTEVEENMEKSGKKSSKRSKKKDSLNIVEEAQVVDSLQQKKEAE 550
Query: 193 ESVKKWRKQRQQSGFADDGAD-----KALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQA 247
E+++K K+ + D + + L E V + D S AK+
Sbjct: 551 ENLEKSGKKSSKKTKKKDSLNIVEEAQVLSVEVNNVAQEEASPINNPKDTDASFTPAKKT 610
Query: 248 FGKGKKPKK--GDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKK 298
P K +V ++ + K+N AD ++FG K VGG+ K
Sbjct: 611 TESNASPLKKISEVTDNTEDLNRSMQVLKEN-ADMGDNFGSSQKDDIVGGTNK 662
>At3g56150 PROBABLE EUKARYOTIC TRANSLATION INITIATION FACTOR 3
SUBUNIT 8
Length = 900
Score = 39.3 bits (90), Expect = 0.003
Identities = 52/222 (23%), Positives = 91/222 (40%), Gaps = 34/222 (15%)
Query: 29 AEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGD-------ISWPENVEWIHK 81
+ + ++ S SE ES+ +V++ E VNNR D + P + +
Sbjct: 3 SRFFTQVGSESEDESDYEVEVNEVQNDDVNNRYLQSGSEDDDDTDTKRVVKPAKDKRFEE 62
Query: 82 LSIDIDQ-EQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTD 140
++ +DQ + + +ND ++ + F + + EK+ I P Y +KT
Sbjct: 63 MTYTVDQMKNAMKINDWVSLQENF-----DKVNKQLEKVMRITEAVKPPTLY----IKTL 113
Query: 141 SHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVK---- 196
+E + LA K EA+++ +K L+ QKLK+ K +DDI +
Sbjct: 114 VMLEDFLNEALANK----EAKKKMSTSNSKALNS--MKQKLKKNNKLYEDDINKYREAPE 167
Query: 197 -KWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPG 237
+ KQ + DD D+ D +D + P V PG
Sbjct: 168 VEEEKQPEDDDDDDDDDDEVEDDDDSSI------DGPTVDPG 203
>At5g24880 glutamic acid-rich protein
Length = 443
Score = 38.9 bits (89), Expect = 0.003
Identities = 40/176 (22%), Positives = 80/176 (44%), Gaps = 9/176 (5%)
Query: 49 LAEPSKTAVNNRDALLDKLGDISWPENVEWIHKLSIDIDQEQEVDVNDDLAR---ELAFY 105
+AE + + N D + +K ++ +N + +K + D ++++D N+ + E
Sbjct: 267 VAELEEKLIKNEDDIEEKTEEMKEQDNNQ-ANKSEEEEDVKKKIDENETPEKVDTESKEV 325
Query: 106 TQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRK 165
E T++ E+++ G + + E VK D EKV+ EK+K+ EE+ K
Sbjct: 326 ESVEETTQEKEEEVKEEGKERVEEEEKEKEKVKEDDQKEKVEEE---EKEKVKGDEEKEK 382
Query: 166 AREAKRL-SKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGA-DKALDFE 219
+E + K+ + K K+ + +D+ + K R+ A GA +D+E
Sbjct: 383 VKEEESAEGKKKEVVKGKKESPSAYNDVIASKMQENPRKNKVLALAGAFQTVIDYE 438
Score = 30.4 bits (67), Expect = 1.2
Identities = 24/97 (24%), Positives = 44/97 (44%), Gaps = 2/97 (2%)
Query: 138 KTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKK 197
K D + K +++ + VE + K E K KE ++ KE+ K K+DD + +
Sbjct: 308 KIDENETPEKVDTESKEVESVEETTQEKEEEVKEEGKERVEEEEKEKEKVKEDDQKEKVE 367
Query: 198 WRKQRQQSGFADDGADKALDFEDGKVFE--RSKKKRP 232
++ + G + K + +GK E + KK+ P
Sbjct: 368 EEEKEKVKGDEEKEKVKEEESAEGKKKEVVKGKKESP 404
>At4g25340 unknown protein
Length = 477
Score = 38.9 bits (89), Expect = 0.003
Identities = 56/259 (21%), Positives = 107/259 (40%), Gaps = 43/259 (16%)
Query: 20 NDDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPENVEWI 79
+D+ DG SEE E +SEED +L E + D+ L++ + + P++ I
Sbjct: 107 HDEDDSDGIDVGESEEDDSCEYDSEEDEQLDE----FEDFLDSNLERYRNAAAPKSGVII 162
Query: 80 HKLSIDIDQEQEVDVNDDLARELAFYTQALEG--TRQAFEKLQSIGLPFLRPADYYAEMV 137
++ +++E D+ A++ +QA EG ++ ++ +P L D + +
Sbjct: 163 EEI-----EDEEKPAKDNKAKQTKKKSQASEGENAKKQIVAIEGAHVPVLESEDEDEDGL 217
Query: 138 K----TDSHMEKVKSRLL--------AEKQKMVEAEERRKAREAKRLSKEVQSQKLKERA 185
S +E + + K++ +A E+ +E+ SK+ ++QK K++
Sbjct: 218 PIPKGKSSEVENASGEKMVVDNDEQGSNKKRKAKAAEQDDGQESANKSKKKKNQKEKKKG 277
Query: 186 KQ---------------KKDDIESVKKWRKQRQQSGFADDG---ADKALDFEDGKVFERS 227
+ KK DI + + Q G A++ + K D K +
Sbjct: 278 ENVLNEEAGQVQTGNVLKKQDISQISS--NTKAQDGTANNAMSESSKTPDKSAEKKTKNK 335
Query: 228 KKKRPGVSPGDRSGGKAKQ 246
KKK+P + SG KQ
Sbjct: 336 KKKKPSDEAAEISGTVEKQ 354
>At2g18540 putative vicilin storage protein (globulin-like)
Length = 699
Score = 38.9 bits (89), Expect = 0.003
Identities = 25/92 (27%), Positives = 47/92 (50%), Gaps = 4/92 (4%)
Query: 144 EKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQ-- 201
E++ R E+QK E RK RE + +E + K++E +Q+K+ + +K R++
Sbjct: 597 EEMAKRREQERQKKEREEMERKKREEEARKREEEMAKIREEERQRKEREDVERKRREEEA 656
Query: 202 --RQQSGFADDGADKALDFEDGKVFERSKKKR 231
R++ ++ A K + E K E +K+R
Sbjct: 657 MRREEERKREEEAAKRAEEERRKKEEEEEKRR 688
Score = 37.4 bits (85), Expect = 0.010
Identities = 28/98 (28%), Positives = 49/98 (49%), Gaps = 13/98 (13%)
Query: 135 EMVKTDSHMEKVKSRLLAE-KQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIE 193
E+ K +E+ K R E +++ E EE RK EAKR +E ++ +E ++KK + E
Sbjct: 416 ELSKLMREIEERKRREEEEIERRRKEEEEARKREEAKRREEEEAKRREEEETERKKREEE 475
Query: 194 SVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKR 231
+K ++R++ + E+ K E +KKR
Sbjct: 476 EARKREEERKR------------EEEEAKRREEERKKR 501
Score = 36.6 bits (83), Expect = 0.017
Identities = 26/131 (19%), Positives = 63/131 (47%), Gaps = 4/131 (3%)
Query: 144 EKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQR- 202
E+ K R K++ EAE+ RK E + +E+ ++ +ER +++++++E ++ ++R
Sbjct: 489 EEAKRREEERKKREEEAEQARKREEEREKEEEMAKKREEERQRKEREEVERKRREEQERK 548
Query: 203 ---QQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDV 259
+++ ++ + + + ER +K+R V R + K+ K+ ++
Sbjct: 549 RREEEARKREEERKREEEMAKRREQERQRKEREEVERKIREEQERKREEEMAKRREQERQ 608
Query: 260 KNSKFGFGGKK 270
K + KK
Sbjct: 609 KKEREEMERKK 619
Score = 36.6 bits (83), Expect = 0.017
Identities = 25/106 (23%), Positives = 48/106 (44%)
Query: 149 RLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFA 208
R + E+Q+ EE K RE +R KE + + K+R ++ + E + K R++ +Q
Sbjct: 585 RKIREEQERKREEEMAKRREQERQKKEREEMERKKREEEARKREEEMAKIREEERQRKER 644
Query: 209 DDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKP 254
+D K + E + E K++ + K ++ K + P
Sbjct: 645 EDVERKRREEEAMRREEERKREEEAAKRAEEERRKKEEEEEKRRWP 690
Score = 34.7 bits (78), Expect = 0.065
Identities = 18/58 (31%), Positives = 32/58 (55%)
Query: 144 EKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQ 201
E++ R E+Q+ E RK RE + +E + K +E+ +QKK+ E +K R++
Sbjct: 565 EEMAKRREQERQRKEREEVERKIREEQERKREEEMAKRREQERQKKEREEMERKKREE 622
Score = 33.1 bits (74), Expect = 0.19
Identities = 24/93 (25%), Positives = 44/93 (46%), Gaps = 5/93 (5%)
Query: 144 EKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKE-----RAKQKKDDIESVKKW 198
E+ K R E ++ E E RK RE + K + +K +E R +++K E ++
Sbjct: 449 EEAKRREEEEAKRREEEETERKKREEEEARKREEERKREEEEAKRREEERKKREEEAEQA 508
Query: 199 RKQRQQSGFADDGADKALDFEDGKVFERSKKKR 231
RK+ ++ ++ A K + K E ++KR
Sbjct: 509 RKREEEREKEEEMAKKREEERQRKEREEVERKR 541
Score = 32.0 bits (71), Expect = 0.42
Identities = 19/73 (26%), Positives = 37/73 (50%), Gaps = 9/73 (12%)
Query: 135 EMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQ------K 188
E K + M K++ E+Q+ + RK RE + + +E + ++ +E AK+ K
Sbjct: 623 EARKREEEMAKIREE---ERQRKEREDVERKRREEEAMRREEERKREEEAAKRAEEERRK 679
Query: 189 KDDIESVKKWRKQ 201
K++ E ++W Q
Sbjct: 680 KEEEEEKRRWPPQ 692
>At5g25070 unknown protein
Length = 736
Score = 38.5 bits (88), Expect = 0.004
Identities = 39/163 (23%), Positives = 73/163 (43%), Gaps = 9/163 (5%)
Query: 28 EAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPENVEWIHKLSIDID 87
E E L + E E +E+ E + +NN +L S + + + ++D
Sbjct: 392 ELEELLALVKAKEKEIDENDSQIEAVEERINNVVTGFKEL-QTSMDKMLNDVQAGLTEVD 450
Query: 88 QEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVK 147
+E E DL+R+ + + ++ KL+ + A Y E++K +
Sbjct: 451 KETE-----DLSRKKKDVDEFMTSEKERGAKLRDLARVSADEACEYEEVIKLRKGLMSYV 505
Query: 148 SRLLAEKQKMVEAEERRKAREAKRLSKEVQSQK--LKERAKQK 188
S+ E+ K+V EE + + E ++L +EV S + LKER+ +K
Sbjct: 506 SKTREERAKLVNIEE-KLSEEVQKLQEEVSSTRELLKERSSKK 547
>At2g28910 unknown protein
Length = 332
Score = 38.5 bits (88), Expect = 0.004
Identities = 24/95 (25%), Positives = 50/95 (52%), Gaps = 4/95 (4%)
Query: 137 VKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVK 196
VK S + K K R+ E ++ +R++ +R S + +S + + +D+ E
Sbjct: 172 VKKTSSVRKKKKRVSDESDSDSDSGDRKR----RRRSMKKRSSHKRRSLSESEDEEEGRS 227
Query: 197 KWRKQRQQSGFADDGADKALDFEDGKVFERSKKKR 231
K RK+R+ +D +D++ D +D +V +S+K++
Sbjct: 228 KRRKERRGRKRDEDDSDESEDEDDRRVKRKSRKEK 262
Score = 31.6 bits (70), Expect = 0.55
Identities = 31/130 (23%), Positives = 54/130 (40%), Gaps = 8/130 (6%)
Query: 141 SHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRK 200
S +EK++ + + + V +EE ++ + + + ER +KK SVKK
Sbjct: 119 SGLEKIRRGVGKGEVEEVSSEEEEESESSDSDVDSEMERIIAERFGKKKGG-SSVKKTSS 177
Query: 201 QRQQSGFADDGADKALDFEDGKVFERSKKKRP-------GVSPGDRSGGKAKQAFGKGKK 253
R++ D +D D D K RS KKR S + G ++ +G+K
Sbjct: 178 VRKKKKRVSDESDSDSDSGDRKRRRRSMKKRSSHKRRSLSESEDEEEGRSKRRKERRGRK 237
Query: 254 PKKGDVKNSK 263
+ D S+
Sbjct: 238 RDEDDSDESE 247
Score = 29.3 bits (64), Expect = 2.7
Identities = 29/125 (23%), Positives = 58/125 (46%), Gaps = 12/125 (9%)
Query: 146 VKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIE-------SVKKW 198
+K R +++ + E+E+ + R +R KE + +K E + +D + S K+
Sbjct: 205 MKKRSSHKRRSLSESEDEEEGRSKRR--KERRGRKRDEDDSDESEDEDDRRVKRKSRKEK 262
Query: 199 RKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGD 258
R++R + +DD +D +D + R+K S + SG + G+G K+ +
Sbjct: 263 RRRRSRRNHSDD-SDSESSEDDRRQKRRNKVAASSDSEANVSGDDVSRV-GRGSS-KRSE 319
Query: 259 VKNSK 263
K+ K
Sbjct: 320 KKSRK 324
>At1g60200
Length = 781
Score = 38.1 bits (87), Expect = 0.006
Identities = 30/120 (25%), Positives = 55/120 (45%), Gaps = 20/120 (16%)
Query: 132 YYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKK-- 189
Y E + S E+ + R L + ++ + R+ R + KE Q +K KE+ K++K
Sbjct: 308 YEREAERERSRKEREQRRKLEDAERAYQTRLRQWERREREKEKERQYEKEKEKEKERKRK 367
Query: 190 -----------DDIESVKKW-------RKQRQQSGFADDGADKALDFEDGKVFERSKKKR 231
DD +S ++W R++RQ DD AD+ + E+ +RS +++
Sbjct: 368 KEIRYEEEEEEDDDDSRRRWHRAALDERRRRQLREKEDDLADRLKEEEEVAEAKRSAEEQ 427
Score = 30.0 bits (66), Expect = 1.6
Identities = 28/125 (22%), Positives = 56/125 (44%), Gaps = 11/125 (8%)
Query: 88 QEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYA-------EMVKTD 140
+ ++ D N D+AR FY ++ E P D+ E + +
Sbjct: 245 KSKDGDSNTDVARSGCFYH--VDAAANDVETSGEHNRPDTSSPDWSKRNDRRSRERGEKE 302
Query: 141 SHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSK-EVQSQKLKERAKQKKDDIESVKKWR 199
M++ + E+ + E E+RRK +A+R + ++ + +ER K+K+ E K+
Sbjct: 303 QEMDRYEREAERERSRK-EREQRRKLEDAERAYQTRLRQWERREREKEKERQYEKEKEKE 361
Query: 200 KQRQQ 204
K+R++
Sbjct: 362 KERKR 366
>At1g44910 splicing factor like protein
Length = 958
Score = 37.7 bits (86), Expect = 0.008
Identities = 36/150 (24%), Positives = 67/150 (44%), Gaps = 6/150 (4%)
Query: 135 EMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIES 194
E V E + S K+K + +E + +E +R KE + K KER +++++ +
Sbjct: 788 ESVSQGLFEEYITSLQEKAKEKERKRDEEKVRKEKERDEKEKRKDKDKERREKEREREKE 847
Query: 195 VKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKP 254
K R +R++S DG + A+D +G E+ K K R + + +
Sbjct: 848 KGKERSKREES----DG-ETAMDVSEGHKDEKRKGKDRDRKHRRRHHNNSDEDVSSDRDD 902
Query: 255 KKGDVKNS-KFGFGGKKGLKKQNTADTTND 283
+ K+S K G KK K N+ ++ ++
Sbjct: 903 RDESKKSSRKHGNDRKKSRKHANSPESESE 932
>At5g55660 putative protein
Length = 759
Score = 37.4 bits (85), Expect = 0.010
Identities = 28/109 (25%), Positives = 50/109 (45%), Gaps = 18/109 (16%)
Query: 160 AEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWR----KQRQQSGFADDGADKA 215
A+ ++K EA R +K+ + E ++K+DD E K+ ++ ++G D D+A
Sbjct: 486 AKSQKKTEEATRTNKKSVAHSDDESEEEKEDDEEEEKEQEVEEEEEENENGIPDKSEDEA 545
Query: 216 ---------LDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPK 255
++ E+ E KKKR G R+ K++ GK + K
Sbjct: 546 PQLSESEENVESEEESEEETKKKKR-----GSRTSSDKKESAGKSRSKK 589
>At5g57120 unknown protein
Length = 330
Score = 36.6 bits (83), Expect = 0.017
Identities = 41/169 (24%), Positives = 73/169 (42%), Gaps = 17/169 (10%)
Query: 138 KTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKK----DDIE 193
+TD+ +E +K+ V+ E K +E + ++ +K K+++K K DD E
Sbjct: 112 ETDAVIEDGVKEKKKKKETKVKVTEEEKVKETDAVIEDGVKEKKKKKSKSKSVEADDDKE 171
Query: 194 SVKKWRKQRQ----QSGFADDGADKALDFEDGKVFERSK--KKRPGVSPGDRSGGKAKQA 247
V K RK+ + + DD + ++ V E + ++ P + G A+++
Sbjct: 172 KVSKKRKRSEPEETKEETEDDDEESKRRKKEENVVENDEGVQETPVKETETKENGNAEKS 231
Query: 248 FGKGKKPKKG-DVKNSKFGFGGKKGLKKQNTAD---TTNDFGGFSKKGA 292
K K G + NSK KK ++ N + T N +SK GA
Sbjct: 232 ETKSTNQKSGKGLSNSK---EPKKPFQRVNVDEIVYTENSNSYYSKGGA 277
Score = 30.8 bits (68), Expect = 0.94
Identities = 23/100 (23%), Positives = 50/100 (50%), Gaps = 8/100 (8%)
Query: 134 AEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIE 193
++ V+ D EKV ++K+K E EE ++ E + +E + +K +E + + ++
Sbjct: 161 SKSVEADDDKEKV-----SKKRKRSEPEETKE--ETEDDDEESKRRKKEENVVENDEGVQ 213
Query: 194 SVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKK-KRP 232
+ +++G A+ K+ + + GK SK+ K+P
Sbjct: 214 ETPVKETETKENGNAEKSETKSTNQKSGKGLSNSKEPKKP 253
>At5g51770 putative protein
Length = 654
Score = 36.6 bits (83), Expect = 0.017
Identities = 32/110 (29%), Positives = 56/110 (50%), Gaps = 13/110 (11%)
Query: 130 ADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEV-QSQKLKERAKQK 188
+D+ AE + +EK KS E ++ E+ + ++ +R+ +E + + KE AK+
Sbjct: 367 SDWIAETAEGGKKVEKKKSSKRLEWWLSLDEEKEKGKKKKRRMVREWWKDEYRKELAKRM 426
Query: 189 KDDIESVKKWRKQRQQSGFADDGADKALDFE---DGKVFERSKKKRPGVS 235
K KK +K+ +S F D ++D DG+V+ +KKR GVS
Sbjct: 427 K------KKKKKKTLESEFYSDDVSGSVDQRRHGDGEVY---RKKRRGVS 467
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.310 0.130 0.355
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,034,349
Number of Sequences: 26719
Number of extensions: 328596
Number of successful extensions: 2387
Number of sequences better than 10.0: 283
Number of HSP's better than 10.0 without gapping: 76
Number of HSP's successfully gapped in prelim test: 208
Number of HSP's that attempted gapping in prelim test: 1881
Number of HSP's gapped (non-prelim): 564
length of query: 301
length of database: 11,318,596
effective HSP length: 99
effective length of query: 202
effective length of database: 8,673,415
effective search space: 1752029830
effective search space used: 1752029830
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC121233.5


