

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121233.1 - phase: 0 /pseudo
(1172 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
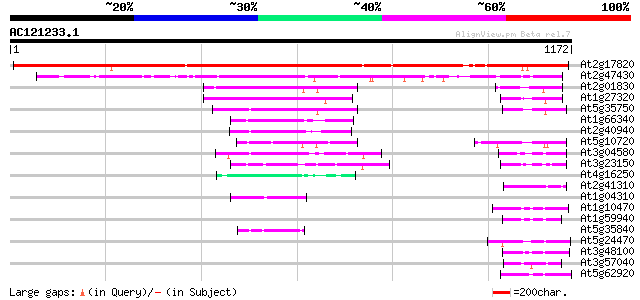
Score E
Sequences producing significant alignments: (bits) Value
At2g17820 histidine kinase 1 1432 0.0
At2g47430 putative histidine kinase 280 3e-75
At2g01830 cytokinin receptor CRE1b (CRE1) 161 2e-39
At1g27320 histidine kinase (AHK3) 154 4e-37
At5g35750 histidine kinase (AHK2) 147 3e-35
At1g66340 ethylene-response protein, ETR1 126 6e-29
At2g40940 ethylene response sensor (ERS) 119 8e-27
At5g10720 histidine kinase - like protein 112 2e-24
At3g04580 putative ethylene receptor 77 6e-14
At3g23150 thylene receptor like protein 72 2e-12
At4g16250 phytochrome D 54 5e-07
At2g41310 putative two-component response regulator 3 protein 54 7e-07
At1g04310 putative ethylene receptor ERS2 54 7e-07
At1g10470 putative response regulator 3 53 9e-07
At1g59940 putative protein 52 3e-06
At5g35840 phytochrome C (sp|P14714) 51 3e-06
At5g24470 pseudo-response regulator 5 (APRR5) 49 1e-05
At3g48100 response reactor 2 (ATRR2) 49 2e-05
At3g57040 responce reactor 4 48 3e-05
At5g62920 response regulator 6 (ARR6) 47 6e-05
>At2g17820 histidine kinase 1
Length = 1207
Score = 1432 bits (3706), Expect = 0.0
Identities = 760/1192 (63%), Positives = 917/1192 (76%), Gaps = 56/1192 (4%)
Query: 8 VFDRLLLWFATCWKKNSTLTGRRKF---HKDVEKEDFQYSTTQCLSSYYSVFVVRLAIMV 64
VFD++ W T W+ N L R+ DVE+++FQY+++ CLSSYYSVFVVRLAIMV
Sbjct: 30 VFDKMTEW-VTPWRSN--LESPREMMILRGDVEQDEFQYASSHCLSSYYSVFVVRLAIMV 86
Query: 65 MLAILIGLLTILTWHFTKIYTTKSLNSLAYDLRYELLQRPILRMWNILNSTAEITTAQVK 124
MLAILIGLLT+LTWHFT+IYT +SL +LAY LRYELLQRP+LRMW++LN+T+E+TTAQVK
Sbjct: 87 MLAILIGLLTVLTWHFTRIYTKQSLQTLAYGLRYELLQRPVLRMWSVLNTTSELTTAQVK 146
Query: 125 LSEYVIRSHGNLATQAEQVEMYESMRAVTWALFASRKALNSITVKYRNGFVQAFHRDLKD 184
LSEYVI+ + TQ E VEMY++M+ VTWALFAS KALN+IT+ YRNGFVQAFHRD
Sbjct: 147 LSEYVIKKYDKPTTQEELVEMYQAMKDVTWALFASAKALNAITINYRNGFVQAFHRDPAS 206
Query: 185 NNIFYIYTDLSYNETNSFAAHEDTNS--------NKSAIWYREQLDPVNGEKIGKAMKIA 236
++ FYI++DL N + S ED + N SAIWY++QLDPV GE +GK +KI
Sbjct: 207 SSTFYIFSDLK-NYSISGTGPEDVSGWNNKSIHGNMSAIWYQQQLDPVTGENLGKPLKIP 265
Query: 237 PEDSISIAGLSQVPDGVASWHVSVGKFTDSPLLSAALPVWDSSNKSIVAVVGVTTALYSV 296
P+D I+IAG+SQVPDG ASWHV+V K+ DSPLLSAALPV+D+SNKSIVAVVGVTTALYSV
Sbjct: 266 PDDLINIAGISQVPDGEASWHVTVSKYMDSPLLSAALPVFDASNKSIVAVVGVTTALYSV 325
Query: 297 GQLMKELVDKHSGHMYLTSQEGYLLATSTNDPLLTNSTKKPKLKMAVDCDNEVIREGAMW 356
GQLM++LV+ H GH+YLTSQEGYLLATST+ PLL N++ P+L A D + VI+ GA W
Sbjct: 326 GQLMRDLVEVHGGHIYLTSQEGYLLATSTDGPLLKNTSNGPQLMKATDSEEWVIKSGAQW 385
Query: 357 LKKTYENNFPPSHEVHEENARLGHQQYYIDSFFLILKKLPLVGVIIIPRKHIMGQADERA 416
L+KTY + P H VH EN +LG Q+YYIDSF+L LK+LP+VGV+IIPRK IMG+ DERA
Sbjct: 386 LEKTYGSKRP--HVVHAENVKLGDQRYYIDSFYLNLKRLPIVGVVIIPRKFIMGKVDERA 443
Query: 417 FKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFL 476
FKTL+ILISAS+CI IGCVCILILTNGVSKEM LRAELI L+ARR+AEASSNYKSQFL
Sbjct: 444 FKTLIILISASVCIFFIGCVCILILTNGVSKEMKLRAELIRQLDARRRAEASSNYKSQFL 503
Query: 477 ANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVES 536
ANMSHELRTPMAAVIGLLDIL+SDD L+NEQ ATVTQIRKCSTALL LLNNILD+SKVES
Sbjct: 504 ANMSHELRTPMAAVIGLLDILISDDCLSNEQYATVTQIRKCSTALLRLLNNILDLSKVES 563
Query: 537 GKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANL 596
GKLVLEEAEFD GRELEGLVDMFSVQCINHNVE +LDLSDDMP LVRGDSAR+VQIFANL
Sbjct: 564 GKLVLEEAEFDLGRELEGLVDMFSVQCINHNVETVLDLSDDMPALVRGDSARLVQIFANL 623
Query: 597 INNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNID 656
I+NSIKFT +GHIILRGWCEN NS D + +++++ KTK QH NH +K+
Sbjct: 624 ISNSIKFTTTGHIILRGWCENINSLHDEMSVSVDRRKPWAPMKTKQVQHRNHLQKSCKNA 683
Query: 657 NKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEI 716
NKM+LWFEVDDTGCGIDPSKWDSVFESFEQAD STTR HGGTGLGLCIVR+LVNKMGGEI
Sbjct: 684 NKMVLWFEVDDTGCGIDPSKWDSVFESFEQADPSTTRTHGGTGLGLCIVRNLVNKMGGEI 743
Query: 717 KIVKKEGPGTLMRLYMQLTAPVDATEQHCQVDFANNNLMVLLALHGNMSRLITSKWLQKN 776
K+V+K G GTLMRLY+ L+ P D +Q+ Q DF+ L+V+L+++G+ +R+ITSKWL+K+
Sbjct: 744 KVVQKNGLGTLMRLYLILSTP-DTVDQNIQPDFSKYGLVVMLSMYGSTARMITSKWLRKH 802
Query: 777 GVLTMEASEWNGLTQILKELFEAKTSTHNNDFDTHFPAPEGLNSKFISIQELPNPTFVIV 836
G+ T+EAS+WN LTQI+++L E T + +N FD+ + L ++ +I E+ NP FVIV
Sbjct: 803 GIATVEASDWNELTQIIRDLLE--TGSRDNSFDSQHNISDPLRAELSNIVEIKNPVFVIV 860
Query: 837 VDIDLLDLSTDIWKEQLNFLHKYYARAKFVWLQNHDSSNTVKTELRKKGHILTVNKPLYK 896
VDI +LDL+T+IWKEQLN+L ++ +AKF WL HD+SNTVKTELR+KGH++ VNKPLYK
Sbjct: 861 VDIGVLDLTTNIWKEQLNYLDRFSNKAKFAWLLKHDTSNTVKTELRRKGHVMMVNKPLYK 920
Query: 897 AKMIHILEDVIKERN----VEVQKKNMKEDDLHESLEIEYTHHCDVASSDGSDISEIGSS 952
AKMI ILE VIK R +++ + D+ H+ LEI+ T +S D S+ S
Sbjct: 921 AKMIQILEAVIKNRKRGLCNDLRNRGNGSDESHDCLEIDPTQFDTCSSDDSSETS----- 975
Query: 953 NLVTANGEKQREEVVRINPSSLYQK--SNCLLGLSNGYMEHKEAFISTRAKGEDSKGGET 1010
GEKQ ++ V+ PS+L+ N L+ + + A S K + + +
Sbjct: 976 ------GEKQVDKSVK--PSTLHSPVLKNYLIDATTSNDDSTSA--SMTQKNPEEEDWKD 1025
Query: 1011 SRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALN- 1069
SG + +KSLEG+RILLAEDT V+QRVATIMLEKMGATV AV DGQQAVD+LN
Sbjct: 1026 RLYSGIALDGKNQKSLEGIRILLAEDTPVLQRVATIMLEKMGATVTAVWDGQQAVDSLNY 1085
Query: 1070 -----GMPGVERNTI---------TSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKS 1115
P E + T +T + + YDLILMDCQMPKMDGYEATK IR++
Sbjct: 1086 KSINAQAPTEEHKSFEEETANKVTTRETSLRNSSPYDLILMDCQMPKMDGYEATKAIRRA 1145
Query: 1116 EIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTK 1167
EIGT HIPIVALTAHAMS DEAKCLEVGMDAYLTKPID KLM STILSLTK
Sbjct: 1146 EIGTELHIPIVALTAHAMSSDEAKCLEVGMDAYLTKPIDRKLMVSTILSLTK 1197
>At2g47430 putative histidine kinase
Length = 1122
Score = 280 bits (717), Expect = 3e-75
Identities = 297/1192 (24%), Positives = 498/1192 (40%), Gaps = 191/1192 (16%)
Query: 57 VVRLAIMVMLAILIGLLTILTWHFTKIYTTKSLNSLAYDLRYELLQRPILRMWNILNSTA 116
+V ++ L ++ + I W T K + S DLR L+ + +
Sbjct: 14 IVVFCVLAFLVVVFECIWISNWRTTTENLVKEVASFTEDLRTSLVSE--------IENIG 65
Query: 117 EITTAQVKLSEYVIRSHGNLATQAEQVEMYESMRAVTWALFASRKALNSITVKYRNGFVQ 176
+ T A+ LS + + E + LF + + ++ V
Sbjct: 66 KFTYAKTNLSTIGLARVIDSYITNNDTGFTEIQTQIAPLLFVAYSTILQVSQ------VS 119
Query: 177 AFHRDLKDNNIFYIYTDLSYNETNSFAAHEDTNSNKSAIWYREQLDPVNGEKIGKAMKIA 236
RD + + Y S FA +S WY + +D + G G + K
Sbjct: 120 YISRD----GLMFSYIAESNTSVAVFANSSSNSSRGDYTWYTQTVDQLTGRLNGNSTKSQ 175
Query: 237 PEDSISIAGLSQVPDG---VASWHVSVGKFTDSPLLSAALPVWDSSNKSIVAVVGVTTAL 293
D A S+G + L+ + + ++ S K +V++ L
Sbjct: 176 SLDVTHTDWFQAAQSNNYTTAFVGTSLGGEDNETLIQSVVSLY--SKKGLVSLGFPVKTL 233
Query: 294 YSVGQLMKELVDKHSGHMYLTSQEGYLLAT--STNDPLLTNSTKKPKLKMAVDCDNEVIR 351
V ++ H +Y+ +++G +L S ND ++ C
Sbjct: 234 TEV----LNSLNLHGEELYMWTKDGTVLVREGSLNDSFFISNGSI--------CFGR--E 279
Query: 352 EGAMWLKKTYENNFPPSHEVHEENARLGHQQYYIDSFFLILKKLPLVGVIIIPRK----H 407
++W + EN +EV E RL +Q + + + +PL ++ P K
Sbjct: 280 SNSLWSQCIPENCSSSGYEV--EIKRLRYQAF---CSVIEVSGVPLRYTLMFPNKGGATR 334
Query: 408 IMGQADERAFKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEA 467
I QA++ ++ +V++I V + + +EM++RA LI+ +EA ++AE
Sbjct: 335 IKHQAEKAKYQLIVVMIFLGFGWPVW---FVWFMMQATRREMHMRATLINQMEATQQAER 391
Query: 468 SSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNN 527
S KSQ AN SH++R +A + GL+DI + ++ T+ Q+ C+ L+ LLN+
Sbjct: 392 KSMNKSQAFANASHDIRGALAGMKGLIDICRDGVKPGSDVDTTLNQVNVCAKDLVALLNS 451
Query: 528 ILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMP---KLVRG 584
+LD+SK+ESGK+ L E +F+ + LE ++D + + V+++LD D VRG
Sbjct: 452 VLDMSKIESGKMQLVEEDFNLSKLLEDVIDFYHPVAMKKGVDVVLDPHDGSVFKFSNVRG 511
Query: 585 DSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFG--------- 635
DS R+ QI NL++N++KFT+ GHI +R W + P G + + L P G
Sbjct: 512 DSGRLKQILNNLVSNAVKFTVDGHIAVRAWAQRP---GSNSSVVLASYPKGVSKFVKSMF 568
Query: 636 CSRKTKMKQHENH-SKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRL 694
C K + +E S N N M FEVDDTG GI SVFE++ Q T +
Sbjct: 569 CKNKEESSTYETEISNSIRNNANTMEFVFEVDDTGKGIPMEMRKSVFENYVQV-RETAQG 627
Query: 695 HGGTGLGLCIVRSLVNKMGGEIKIVKKE--GPGTLMRLYMQLTA----PVDATEQHCQVD 748
H GTGLGL IV+SLV MGGEI+I K GT + + LT PV + +++
Sbjct: 628 HQGTGLGLGIVQSLVRLMGGEIRITDKAMGEKGTCFQFNVLLTTLESPPVSDMKVRQEIE 687
Query: 749 FAN-----------------------------NNLM----------VLLALHGNMSRLIT 769
NN + V+L L R +T
Sbjct: 688 AGGDYVSTPNLGLTINTSLGGSMNIRNLSPRFNNCLSSSPKQEGSRVVLLLKNEERRRVT 747
Query: 770 SKWLQKNGVLTMEASEWNGLTQILKELFEAKTSTHNNDFDTHFPAPEGLNSKFI------ 823
K+++ G+ +W L+ L+ LF + + P FI
Sbjct: 748 EKYIKNLGIKVTVVEKWEHLSYALERLFGFSPQSSMGRAECSLSCPSSRELPFIGMDGID 807
Query: 824 SIQELPNPTFVIVVDIDLL--DLSTDIWKEQLNFLHKYY------ARAKFVWLQNHDSSN 875
S +LP + + LL D T + E + + ++ K VWL N
Sbjct: 808 SRSQLPKRRSISFSAVVLLVIDAKTGPFFELCDIVKQFRRGLPHGISCKVVWL------N 861
Query: 876 TVKTELRKKGHILTVNKPLYKAKMIHILE-------DVIKERNVEVQKKNMKEDDLHESL 928
T + ++G I + ++PL+ ++++ +L+ V+KE E+Q++++
Sbjct: 862 ESSTRVSERGDI-SCSRPLHGSRLMEVLKMLPEFGGTVLKEPPTELQRESL--------- 911
Query: 929 EIEYTHHCDVASSDGSDISEIGSSNLVTANGEKQREEVVRINPSSLYQK--SNCLLGLSN 986
H VA + +V PSS++ K ++ ++
Sbjct: 912 ----LRHSFVAE-------------------RSPKHKVQEEGPSSMFNKKLGKRIMASTD 948
Query: 987 GYMEHKEAFISTRAK--GEDSKGGETSRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVA 1044
E + + T K G ETS+ S + L G R+L+ +D + ++VA
Sbjct: 949 SESETRVKSVRTGRKPIGNPEDEQETSKPSDD-------EFLRGKRVLVVDDNFISRKVA 1001
Query: 1045 TIMLEKMGATVVAVGD-GQQAVDALNGMPGVERNTITSQTEILSFPSYDLILMDCQMPKM 1103
T L+KMG + V D G++A+ + G+ + + L F D I MDCQMP+M
Sbjct: 1002 TGKLKKMGVSEVEQCDSGKEALRLVT--EGLTQREEQGSVDKLPF---DYIFMDCQMPEM 1056
Query: 1104 DGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAK-CLEVGMDAYLTKPID 1154
DGYEAT+EIRK E PI+A++ H +EA+ ++ GMDA+L K ++
Sbjct: 1057 DGYEATREIRKVEKSYGVRTPIIAVSGHDPGSEEARETIQAGMDAFLDKSLN 1108
>At2g01830 cytokinin receptor CRE1b (CRE1)
Length = 844
Score = 161 bits (407), Expect = 2e-39
Identities = 114/347 (32%), Positives = 177/347 (50%), Gaps = 27/347 (7%)
Query: 405 RKHIM-GQADERAFKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARR 463
RKH M + ++A L +L + L IG + IL + + + E +
Sbjct: 170 RKHKMICRYHQKAPIPLNVLTTVPL-FFAIGFLVGYILYGAAMHIVKVEDDFHEMQELKV 228
Query: 464 KAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLH 523
+AEA+ KSQFLA +SHE+RTPM ++G+L +LL D L++ Q + C AL+
Sbjct: 229 RAEAADVAKSQFLATVSHEIRTPMNGILGMLAMLL-DTELSSTQRDYAQTAQVCGKALIA 287
Query: 524 LLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVR 583
L+N +LD +K+E+GKL LE FD L+ ++ +FS + N ++E+ + +SD +P++V+
Sbjct: 288 LINEVLDRAKIEAGKLELESVPFDIRSILDDVLSLFSEESRNKSIELAVFVSDKVPEIVK 347
Query: 584 GDSARVVQIFANLINNSIKFTLSGHIILR----------GWCENPNSYGDSENFTLEQKP 633
GDS R QI NL+ NS+KFT GHI ++ +N + G SE + K
Sbjct: 348 GDSGRFRQIIINLVGNSVKFTEKGHIFVKVHLAEQSKDESEPKNALNGGVSEEMIVVSKQ 407
Query: 634 FGCSRKT--------------KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDS 679
+ + K E S +I + + L ++DTG GI
Sbjct: 408 SSYNTLSGYEAADGRNSWDSFKHLVSEEQSLSEFDISSNVRLMVSIEDTGIGIPLVAQGR 467
Query: 680 VFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGT 726
VF F QAD ST+R +GGTG+GL I + LV M G+I + + G+
Sbjct: 468 VFMPFMQADSSTSRNYGGTGIGLSISKCLVELMRGQINFISRPHIGS 514
Score = 83.6 bits (205), Expect = 6e-16
Identities = 54/147 (36%), Positives = 78/147 (52%), Gaps = 26/147 (17%)
Query: 1015 GSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGV 1074
GSS A K L G +IL+ +D V +RVA L+K GA VV GQ A+
Sbjct: 696 GSSPATL-KSLLTGKKILVVDDNIVNRRVAAGALKKFGAEVVCAESGQVALG-------- 746
Query: 1075 ERNTITSQTEILSFP-SYDLILMDCQMPKMDGYEATKEIR------KSEIGTSFHIPIVA 1127
+L P ++D MD QMP+MDG+EAT++IR K + +H+PI+A
Sbjct: 747 ----------LLQIPHTFDACFMDIQMPQMDGFEATRQIRMMEKETKEKTNLEWHLPILA 796
Query: 1128 LTAHAMSCDEAKCLEVGMDAYLTKPID 1154
+TA + +CL+ GMD Y++KP +
Sbjct: 797 MTADVIHATYEECLKSGMDGYVSKPFE 823
>At1g27320 histidine kinase (AHK3)
Length = 1036
Score = 154 bits (388), Expect = 4e-37
Identities = 112/321 (34%), Positives = 161/321 (49%), Gaps = 10/321 (3%)
Query: 405 RKHIMGQADERAFKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRK 464
RKH M ++ V+ + S I+VI + I+ VS+ + + + ++K
Sbjct: 384 RKHEMRCRFKQKPPWPVLSMVTSFGILVIALLVAHIIHATVSRIHKVEEDCDKMKQLKKK 443
Query: 465 AEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHL 524
AEA+ KSQFLA +SHE+RTPM V+G+L +L+ D L Q V + AL+ L
Sbjct: 444 AEAADVAKSQFLATVSHEIRTPMNGVLGMLHMLM-DTELDVTQQDYVRTAQASGKALVSL 502
Query: 525 LNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRG 584
+N +LD +K+ESGKL LEE FD L+ ++ +FS + VE+ + +SD +P ++ G
Sbjct: 503 INEVLDQAKIESGKLELEEVRFDLRGILDDVLSLFSSKSQQKGVELAVYISDRVPDMLIG 562
Query: 585 DSARVVQIFANLINNSIKFTLSGHIILRGWC--ENPNSYGDSENFTLEQKPFGCSRKTKM 642
D R QI NL+ NSIKFT GHI + E S + E G +
Sbjct: 563 DPGRFRQILTNLMGNSIKFTEKGHIFVTVHLVDELFESIDGETASSPESTLSGLPVADRQ 622
Query: 643 KQHENHSKKASNIDNK-------MILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLH 695
+ EN +SN + L V+DTG GI +F F Q S +R H
Sbjct: 623 RSWENFKAFSSNGHRSFEPSPPDINLIVSVEDTGVGIPVEAQSRIFTPFMQVGPSISRTH 682
Query: 696 GGTGLGLCIVRSLVNKMGGEI 716
GGTG+GL I + LV M GEI
Sbjct: 683 GGTGIGLSISKCLVGLMKGEI 703
Score = 75.1 bits (183), Expect = 2e-13
Identities = 50/147 (34%), Positives = 71/147 (48%), Gaps = 35/147 (23%)
Query: 1026 LEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEI 1085
L G +IL+ +D V RVA L+K GA VV G +A+ L P E
Sbjct: 887 LLGRKILIVDDNNVNLRVAAGALKKYGADVVCAESGIKAISLLK--PPHE---------- 934
Query: 1086 LSFPSYDLILMDCQMPKMDGYEATKEIRKSEIG------------------TSFHIPIVA 1127
+D MD QMP+MDG+EAT+ IR E TS+H+P++A
Sbjct: 935 -----FDACFMDIQMPEMDGFEATRRIRDMEEEMNKRIKNGEALIVENGNKTSWHLPVLA 989
Query: 1128 LTAHAMSCDEAKCLEVGMDAYLTKPID 1154
+TA + +CL+ GMD Y++KP +
Sbjct: 990 MTADVIQATHEECLKCGMDGYVSKPFE 1016
>At5g35750 histidine kinase (AHK2)
Length = 1176
Score = 147 bits (372), Expect = 3e-35
Identities = 104/319 (32%), Positives = 160/319 (49%), Gaps = 17/319 (5%)
Query: 424 ISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHEL 483
I+ S+ ++VI + IL +++ + + E + +AEA+ KSQFLA +SHE+
Sbjct: 540 ITPSILVLVITFLVGYILYEAINRIATVEEDCQKMRELKARAEAADIAKSQFLATVSHEI 599
Query: 484 RTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEE 543
RTPM V+G+L +L+ D L +Q L L+N +LD +K+ESG+L LE
Sbjct: 600 RTPMNGVLGMLKMLMDTD-LDAKQMDYAQTAHGSGKDLTSLINEVLDQAKIESGRLELEN 658
Query: 544 AEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKF 603
FD L+ + + S + +E+ + +S +P +V GD +R QI NL+ NSIKF
Sbjct: 659 VPFDMRFILDNVSSLLSGKANEKGIELAVYVSSQVPDVVVGDPSRFRQIITNLVGNSIKF 718
Query: 604 TLS-GHIILR-GWCENPNSYGDSENFTLEQK-PFGCSRKTK-------------MKQHEN 647
T GHI + + E+ L+Q+ GCS + K +
Sbjct: 719 TQERGHIFISVHLADEVKEPLTIEDAVLKQRLALGCSESGETVSGFPAVNAWGSWKNFKT 778
Query: 648 HSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRS 707
S +++ L V+DTG GI +F F QAD ST+R +GGTG+GL I +
Sbjct: 779 CYSTESQNSDQIKLLVTVEDTGVGIPVDAQGRIFTPFMQADSSTSRTYGGTGIGLSISKR 838
Query: 708 LVNKMGGEIKIVKKEGPGT 726
LV M GE+ V + G G+
Sbjct: 839 LVELMQGEMGFVSEPGIGS 857
Score = 72.0 bits (175), Expect = 2e-12
Identities = 44/151 (29%), Positives = 75/151 (49%), Gaps = 35/151 (23%)
Query: 1030 RILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSFP 1089
+IL+ +D V +RVA L+K GA V V G+ A+ L
Sbjct: 1036 QILVVDDNLVNRRVAEGALKKYGAIVTCVESGKAALAMLKPPH----------------- 1078
Query: 1090 SYDLILMDCQMPKMDGYEATKEIRKSEIG------------------TSFHIPIVALTAH 1131
++D MD QMP+MDG+EAT+ +R+ E +S+H+PI+A+TA
Sbjct: 1079 NFDACFMDLQMPEMDGFEATRRVRELEREINKKIASGEVSAEMFCKFSSWHVPILAMTAD 1138
Query: 1132 AMSCDEAKCLEVGMDAYLTKPIDFKLMESTI 1162
+ +C++ GMD Y++KP + +++ + +
Sbjct: 1139 VIQATHEECMKCGMDGYVSKPFEEEVLYTAV 1169
>At1g66340 ethylene-response protein, ETR1
Length = 738
Score = 126 bits (317), Expect = 6e-29
Identities = 85/258 (32%), Positives = 127/258 (48%), Gaps = 26/258 (10%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
ARR+AE + ++ FLA M+HE+RTPM A+I L LL + LT EQ V I K S
Sbjct: 333 ARREAETAIRARNDFLAVMNHEMRTPMHAIIALSS-LLQETELTPEQRLMVETILKSSNL 391
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
L L+N++LD+S++E G L LE F+ ++++ + + I L+L+ D+P+
Sbjct: 392 LATLMNDVLDLSRLEDGSLQLELGTFNLHTLFREVLNLIKPIAVVKKLPITLNLAPDLPE 451
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
V GD R++QI N++ N++KF+ G I + + D+ P G
Sbjct: 452 FVVGDEKRLMQIILNIVGNAVKFSKQGSISVTALV----TKSDTRAADFFVVPTG----- 502
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
+ L +V D+G GI+P +F F Q TR GG+GL
Sbjct: 503 ----------------SHFYLRVKVKDSGAGINPQDIPKIFTKFAQTQSLATRSSGGSGL 546
Query: 701 GLCIVRSLVNKMGGEIKI 718
GL I + VN M G I I
Sbjct: 547 GLAISKRFVNLMEGNIWI 564
Score = 43.1 bits (100), Expect = 0.001
Identities = 38/172 (22%), Positives = 70/172 (40%), Gaps = 26/172 (15%)
Query: 1005 SKGGETSRVSGSSK--AMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQ 1062
S+ S+ SG K A+ + GL++L+ ++ V + V +L +G V
Sbjct: 584 SERSNESKQSGIPKVPAIPRHSNFTGLKVLVMDENGVSRMVTKGLLVHLGCEVT------ 637
Query: 1063 QAVDALNGMPGVERNTITSQTEILSFPSYD--LILMDCQMPKMDGYEATKEIRKSEIGTS 1120
T++S E L S++ ++ MD MP ++ Y+ I +
Sbjct: 638 ---------------TVSSNEECLRVVSHEHKVVFMDVCMPGVENYQIALRIHEKFTKQR 682
Query: 1121 FHIPI-VALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTKRETL 1171
P+ VAL+ + + KC+ G+D L KP+ + + L + L
Sbjct: 683 HQRPLLVALSGNTDKSTKEKCMSFGLDGVLLKPVSLDNIRDVLSDLLEPRVL 734
>At2g40940 ethylene response sensor (ERS)
Length = 613
Score = 119 bits (299), Expect = 8e-27
Identities = 81/257 (31%), Positives = 126/257 (48%), Gaps = 26/257 (10%)
Query: 460 EARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCST 519
+AR++AE + + ++ FLA M+HE+RTPM A+I L +LL + L+ EQ + I K S
Sbjct: 332 KARQEAEMAVHARNDFLAVMNHEMRTPMHAIISLSSLLLETE-LSPEQRVMIETILKSSN 390
Query: 520 ALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMP 579
+ L++++LD+S++E G L+LE F E ++ + + L LS D+P
Sbjct: 391 LVATLISDVLDLSRLEDGSLLLENEPFSLQAIFEEVISLIKPIASVKKLSTNLILSADLP 450
Query: 580 KLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRK 639
GD R++Q N++ N++KFT G+I + P S L++ P
Sbjct: 451 TYAIGDEKRLMQTILNIMGNAVKFTKEGYISIIASIMKPES--------LQELP------ 496
Query: 640 TKMKQHENHSKKASNI--DNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGG 697
S + + D+ L +V DTGCGI +F F Q T R H G
Sbjct: 497 ---------SPEFFPVLSDSHFYLCVQVKDTGCGIHTQDIPLLFTKFVQPRTGTQRNHSG 547
Query: 698 TGLGLCIVRSLVNKMGG 714
GLGL + + V MGG
Sbjct: 548 GGLGLALCKRFVGLMGG 564
>At5g10720 histidine kinase - like protein
Length = 950
Score = 112 bits (279), Expect = 2e-24
Identities = 84/273 (30%), Positives = 133/273 (47%), Gaps = 25/273 (9%)
Query: 474 QFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISK 533
Q LA MSHE+R+P++ V+G+ +IL S +L EQ + + +L L+N+ILD+SK
Sbjct: 365 QMLATMSHEIRSPLSGVVGMAEIL-STTKLDKEQRQLLNVMISSGDLVLQLINDILDLSK 423
Query: 534 VESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIF 593
VESG + LE +F RE+ V + + ++ + +++DD+P V GD R+ QI
Sbjct: 424 VESGVMRLEATKFR-PREVVKHVLQTAAASLKKSLTLEGNIADDVPIEVVGDVLRIRQIL 482
Query: 594 ANLINNSIKFTLSGHI-----------ILRGWCENPNSYGDSENFTLEQKPFGCSR---- 638
NLI+N+IKFT G++ +R N ++ +N E + C
Sbjct: 483 TNLISNAIKFTHEGNVGIKLQVISEPSFVRDNALNADTEEHEQNGLTETSVWICCDVWDT 542
Query: 639 ----KTKMKQHENHSKKASNIDNKMIL-WFEVDDTGCGIDPSKWDSVFESFEQADLSTTR 693
K N +K + IL WF C + +F+ + QA R
Sbjct: 543 GIGIPGKQAILTNEHEKTKKLQLTYILTWFMF---WCSCAENALPCLFKKYMQASADHAR 599
Query: 694 LHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGT 726
+GGTGLGL I + LV MGG++ + + G+
Sbjct: 600 KYGGTGLGLAICKQLVELMGGQLTVTSRVSEGS 632
Score = 69.7 bits (169), Expect = 9e-12
Identities = 59/221 (26%), Positives = 98/221 (43%), Gaps = 50/221 (22%)
Query: 971 PSSLYQKSNCLLGLSNGYMEHKEAFISTRAKGEDSKGGETSRVSGSS-----KAMNGKKS 1025
P S +Q N G + + E+ S++A E S ++ SS KA K
Sbjct: 746 PESAHQYEN---GNGRCFSKESESCSSSQASSEGGTLEMESELTVSSHREEEKAETEVKE 802
Query: 1026 LEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEI 1085
+ILL ED + VA M++++G T+ +G +A+ A+N
Sbjct: 803 TSKPKILLVEDNKINIMVAKSMMKQLGHTMDIANNGVEAITAINSS-------------- 848
Query: 1086 LSFPSYDLILMDCQMPKMDGYEATKEIRKSE--------------IGTSFH--------- 1122
SYDL+LMD MP +DG +AT+ IR E I TS +
Sbjct: 849 ----SYDLVLMDVCMPVLDGLKATRLIRSYEETGNWNAAIEAGVDISTSENEQVCMRPTN 904
Query: 1123 -IPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTI 1162
+PI+A+TA+ ++ +C GMD++++KP+ + + +
Sbjct: 905 RLPIIAMTANTLAESSEECYANGMDSFISKPVTLQKLRECL 945
>At3g04580 putative ethylene receptor
Length = 766
Score = 77.0 bits (188), Expect = 6e-14
Identities = 51/143 (35%), Positives = 73/143 (50%), Gaps = 20/143 (13%)
Query: 1021 NGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTIT 1080
N L GLRI LA+D V + V +LEK+G V AV G + LN + VE
Sbjct: 634 NSNSILRGLRITLADDDDVNRTVTKRLLEKLGCEVTAVSSG---FECLNALSNVEM---- 686
Query: 1081 SQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIP-IVALTAHAMSCDEAK 1139
SY ++++D QMP+MDG+E +IRK H P I+ALTA +
Sbjct: 687 ---------SYRVVILDLQMPEMDGFEVAMKIRKF---CGHHWPLIIALTASTEDHVRER 734
Query: 1140 CLEVGMDAYLTKPIDFKLMESTI 1162
CL++GM+ + KP+ +M S +
Sbjct: 735 CLQMGMNGMIQKPVLLHVMASEL 757
Score = 70.1 bits (170), Expect = 7e-12
Identities = 89/366 (24%), Positives = 149/366 (40%), Gaps = 41/366 (11%)
Query: 431 IVIGCVCILILTNGVSKEMNLRAELI-----SHLEARRKAEASSNYKSQFLANMSHELRT 485
+V V + I V +E L E + + L A++ A +S ++ MSH +R
Sbjct: 322 VVADQVAVAISHASVLEESQLMREKLGIQNRALLRAKQNAMMASQARNTCQKVMSHGMRR 381
Query: 486 PMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAE 545
PM ++GLL + S+ ++ +Q V + K ST L L+N+++DIS ++GK LE
Sbjct: 382 PMHTILGLLSMFQSES-MSLDQKIIVDALMKTSTVLSALINDVIDISPKDNGKSALEVKR 440
Query: 546 FDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTL 605
F + + + +D+ +P LV GD R Q+ ++ + T
Sbjct: 441 FQLHSLIREAACVAKCLSVYKGYGFEMDVQTRLPNLVVGDEKRTFQLVMYMLGYILDMTD 500
Query: 606 SGH-IILRGWCENPNSYGDSENFTLEQKPFGCSRKTKM-KQHENHSKKASNIDNKMILWF 663
G + R CE + D R+T M K H + D+ + + F
Sbjct: 501 GGKTVTFRVICEGTGTSQDKS-----------KRETGMWKSHMS--------DDSLGVKF 541
Query: 664 EVDDTGCGIDPSKWDSV-FESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKI-VKK 721
EV+ P ++ + + G LG+C R L M G I I K
Sbjct: 542 EVEINEIQNPPLDGSAMAMRHIPNRRYHSNGIKEGLSLGMC--RKLAQMMQGNIWISPKS 599
Query: 722 EGPGTLMRLYMQL-TAPV--------DATE-QHCQVDFANNNLMVLLALHGNMSRLITSK 771
G M+L ++ T P +A E QH + L + LA +++R +T +
Sbjct: 600 HGQTQSMQLVLRFQTRPSIRRSILAGNAPELQHPNSNSILRGLRITLADDDDVNRTVTKR 659
Query: 772 WLQKNG 777
L+K G
Sbjct: 660 LLEKLG 665
>At3g23150 thylene receptor like protein
Length = 773
Score = 72.0 bits (175), Expect = 2e-12
Identities = 79/350 (22%), Positives = 143/350 (40%), Gaps = 39/350 (11%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
A+R A +S ++ F MS +R PM +++GLL ++ D++L++EQ V + K
Sbjct: 357 AKRDALRASQARNAFQKTMSEGMRRPMHSILGLLS-MIQDEKLSDEQKMIVDTMVKTGNV 415
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
+ +L+ + +D V G+ E F R + M C+ + + ++D +P
Sbjct: 416 MSNLVGDSMD---VPDGRFGTEMKPFSLHRTIHEAACMARCLCLCNGIRFLVDAEKSLPD 472
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
V GD RV Q+ +++ + +K G S F + ++ R
Sbjct: 473 NVVGDERRVFQVILHIVGSLVK-------------PRKRQEGSSLMFKVLKERGSLDRSD 519
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
+ AS+ D + + FE++ + SV S ++ R GG GL
Sbjct: 520 --HRWAAWRSPASSADGDVYIRFEMNVENDDSSSQSFASV--SSRDQEVGDVRFSGGYGL 575
Query: 701 G----LCIVRSLVNKMGGEIKIVK-KEGPGTLMRLYMQL------------TAPVDATEQ 743
G + + +V + G I +V +G M L ++ +P
Sbjct: 576 GQDLSFGVCKKVVQLIHGNISVVPGSDGSPETMSLLLRFRRRPSISVHGSSESPAPDHHA 635
Query: 744 HCQVDFANNNLMVLLALHGNMSRLITSKWLQKNGV-LTMEASEWNGLTQI 792
H + L VLL + +R +T K L+K G +T +S ++ LT I
Sbjct: 636 HPHSNSLLRGLQVLLVDTNDSNRAVTRKLLEKLGCDVTAVSSGFDCLTAI 685
Score = 58.2 bits (139), Expect = 3e-08
Identities = 42/140 (30%), Positives = 68/140 (48%), Gaps = 23/140 (16%)
Query: 1026 LEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEI 1085
L GL++LL + + V +LEK+G V AV G + A+ PG +
Sbjct: 643 LRGLQVLLVDTNDSNRAVTRKLLEKLGCDVTAVSSGFDCLTAI--APGSSSPST------ 694
Query: 1086 LSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEA---KCLE 1142
S+ ++++D QM +MDGYE IR S P++ T +S DE KC +
Sbjct: 695 ----SFQVVVLDLQMAEMDGYEVAMRIR------SRSWPLIVAT--TVSLDEEMWDKCAQ 742
Query: 1143 VGMDAYLTKPIDFKLMESTI 1162
+G++ + KP+ + MES +
Sbjct: 743 IGINGVVRKPVVLRAMESEL 762
>At4g16250 phytochrome D
Length = 1164
Score = 53.9 bits (128), Expect = 5e-07
Identities = 58/292 (19%), Positives = 115/292 (38%), Gaps = 53/292 (18%)
Query: 432 VIGCVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVI 491
+IG C L + + EL LE +R+ E+ + + LA + ++ P++ +
Sbjct: 901 IIGAFCFLQIPS---------PELQQALEVQRRQESEYFSRRKELAYIFQVIKNPLSG-L 950
Query: 492 GLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRE 551
+ LL D L +Q + C + ++ + +D+ ++ G +LE EF G
Sbjct: 951 RFTNSLLEDMDLNEDQKQLLETSVSCEKQISKIVGD-MDVKSIDDGSFLLERTEFFIGNV 1009
Query: 552 LEGLVDMFSVQCINHNVEIILDLSDDMPKL-VRGDSARVVQIFANLINNSIKFT-LSGHI 609
+V + N+++I ++ ++ + V GD R+ Q+ A + + +++ + G +
Sbjct: 1010 TNAVVSQVMLVVRERNLQLIRNIPTEVKSMAVYGDQIRLQQVLAEFLLSIVRYAPMEGSV 1069
Query: 610 ILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTG 669
L C N D F R L F + G
Sbjct: 1070 ELH-LCPTLNQMADG---------FSAVR----------------------LEFRMACAG 1097
Query: 670 CGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKK 721
G+ P K +F S +R GLGL + R ++ M G ++ +++
Sbjct: 1098 EGVPPEKVQDMFHS--------SRWTSPEGLGLSVCRKILKLMNGGVQYIRE 1141
>At2g41310 putative two-component response regulator 3 protein
Length = 225
Score = 53.5 bits (127), Expect = 7e-07
Identities = 35/134 (26%), Positives = 70/134 (52%), Gaps = 5/134 (3%)
Query: 1031 ILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALN-GMPGVERNTITSQTEILSFP 1089
+L +D+ +++ +L+K V V G +A++ L + + N +++ +I
Sbjct: 11 VLAVDDSLFDRKMIERLLQKSSCQVTTVDSGSKALEFLGLRVDDNDPNALSTSPQIHQEV 70
Query: 1090 SYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAYL 1149
+LI+ D MP M GY+ K++++S S IP+V +++ + ++CLE G + +
Sbjct: 71 EINLIITDYCMPGMTGYDLLKKVKESAAFRS--IPVVIMSSENVPARISRCLEEGAEEFF 128
Query: 1150 TKPIDFKLMESTIL 1163
KP+ KL + T L
Sbjct: 129 LKPV--KLADLTKL 140
>At1g04310 putative ethylene receptor ERS2
Length = 645
Score = 53.5 bits (127), Expect = 7e-07
Identities = 39/161 (24%), Positives = 75/161 (46%), Gaps = 4/161 (2%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
AR A ++ K+ F MS +R P+ +++GLL ++L D +L Q V +R+ S
Sbjct: 372 ARENALRANQAKAAFEQMMSDAMRCPVRSILGLLPLILQDGKLPENQTVIVDAMRRTSEL 431
Query: 521 LLHLLNNILDISKVESGKLVLEEAE-FDFGRELEGLVDMFSVQCINHNVEIILDLSDDMP 579
L+ L+NN DI+ +G + E F ++ + C+ + ++ +P
Sbjct: 432 LVQLVNNAGDIN---NGTIRAAETHYFSLHSVVKESACVARCLCMANGFGFSAEVYRALP 488
Query: 580 KLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNS 620
V GD +V Q +++ + + G++ + E+ NS
Sbjct: 489 DYVVGDDRKVFQAILHMLGVLMNRKIKGNVTFWVFPESGNS 529
>At1g10470 putative response regulator 3
Length = 259
Score = 53.1 bits (126), Expect = 9e-07
Identities = 42/159 (26%), Positives = 74/159 (46%), Gaps = 9/159 (5%)
Query: 1009 ETSRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDAL 1068
E V G + L+ + +L +D+ V + V +L V AV G +A++ L
Sbjct: 14 EMMSVGGIGGIESAPLDLDEVHVLAVDDSLVDRIVIERLLRITSCKVTAVDSGWRALEFL 73
Query: 1069 NGMPGVERNTITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVAL 1128
G++ +++ + L DLI+ D MP M GYE K+I++S +P+V +
Sbjct: 74 ----GLDNEKASAEFDRLKV---DLIITDYCMPGMTGYELLKKIKES--SNFREVPVVIM 124
Query: 1129 TAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTK 1167
++ + +CLE G +L KP+ ++ LTK
Sbjct: 125 SSENVLTRIDRCLEEGAQDFLLKPVKLADVKRLRSHLTK 163
>At1g59940 putative protein
Length = 231
Score = 51.6 bits (122), Expect = 3e-06
Identities = 38/126 (30%), Positives = 66/126 (52%), Gaps = 11/126 (8%)
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSF 1088
+ +L +D+ V + V +L V AV G +A++ L G++ + + + L
Sbjct: 33 VHVLAVDDSLVDRIVIERLLRITSCKVTAVDSGWRALEFL----GLDDDKAAVEFDRLKV 88
Query: 1089 PSYDLILMDCQMPKMDGYEATKEIRKSEIGTSF-HIPIVALTAHAMSCDEAKCLEVGMDA 1147
DLI+ D MP M GYE K+I++S TSF +P+V +++ + +CLE G +
Sbjct: 89 ---DLIITDYCMPGMTGYELLKKIKES---TSFKEVPVVIMSSENVMTRIDRCLEEGAED 142
Query: 1148 YLTKPI 1153
+L KP+
Sbjct: 143 FLLKPV 148
>At5g35840 phytochrome C (sp|P14714)
Length = 1111
Score = 51.2 bits (121), Expect = 3e-06
Identities = 41/141 (29%), Positives = 70/141 (49%), Gaps = 7/141 (4%)
Query: 476 LANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVE 535
LA + HE++ P A+ L D+L S L+ +Q + C L ++++ DI +E
Sbjct: 887 LAYLRHEVKDPEKAISFLQDLLHSSG-LSEDQKRLLRTSVLCREQLAKVISDS-DIEGIE 944
Query: 536 SGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKL-VRGDSARVVQIFA 594
G + L+ +EF LE +V I V+I D ++ + + GD+ R+ QI +
Sbjct: 945 EGYVELDCSEFGLQESLEAVVKQVMELSIERKVQISCDYPQEVSSMRLYGDNLRLQQILS 1004
Query: 595 NLINNSIKFTLSGHIILRGWC 615
+ +SI+FT + LRG C
Sbjct: 1005 ETLLSSIRFTPA----LRGLC 1021
>At5g24470 pseudo-response regulator 5 (APRR5)
Length = 667
Score = 49.3 bits (116), Expect = 1e-05
Identities = 45/181 (24%), Positives = 75/181 (40%), Gaps = 26/181 (14%)
Query: 998 TRAKGEDSKGGETSR--VSGSSKAMNGKKSLE------GLRILLAEDTAVIQRVATIMLE 1049
T K ++ GG+ SR + ++G E LR+LL E +++ +L
Sbjct: 120 TVVKAPEAGGGKLSRRKIRKKDAGVDGLVKWERFLPKIALRVLLVEADDSTRQIIAALLR 179
Query: 1050 KMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSFPSYDLILMDCQMPKMDGYEAT 1109
K V AV DG +A + L G P S DLIL + +P + GY
Sbjct: 180 KCSYRVAAVPDGLKAWEMLKGKP----------------ESVDLILTEVDLPSISGYALL 223
Query: 1110 KEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTKRE 1169
I + +I +IP++ ++ KC+ G YL KP+ + + + +R+
Sbjct: 224 TLIMEHDI--CKNIPVIMMSTQDSVNTVYKCMLKGAADYLVKPLRRNELRNLWQHVWRRQ 281
Query: 1170 T 1170
T
Sbjct: 282 T 282
>At3g48100 response reactor 2 (ATRR2)
Length = 184
Score = 48.9 bits (115), Expect = 2e-05
Identities = 38/143 (26%), Positives = 69/143 (47%), Gaps = 11/143 (7%)
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSF 1088
L +L +D+ V ++ +L V V +A+ L G+ G E N+ ++
Sbjct: 25 LHVLAVDDSMVDRKFIERLLRVSSCKVTVVDSATRALQYL-GLDG-ENNSSVGFEDL--- 79
Query: 1089 PSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAY 1148
+LI+ D MP M GYE K+I++S IP+V +++ + +CLE G + +
Sbjct: 80 -KINLIMTDYSMPGMTGYELLKKIKES--SAFREIPVVIMSSENILPRIDRCLEEGAEDF 136
Query: 1149 LTKPI---DFKLMESTILSLTKR 1168
L KP+ D K + +++ +R
Sbjct: 137 LLKPVKLADVKRLRDSLMKAEER 159
>At3g57040 responce reactor 4
Length = 234
Score = 48.1 bits (113), Expect = 3e-05
Identities = 31/130 (23%), Positives = 64/130 (48%), Gaps = 13/130 (10%)
Query: 1031 ILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILS--- 1087
+L +D+ +++ +L+K V V G +A++ L G+ ++T ++ S
Sbjct: 11 VLAVDDSLFDRKLIERLLQKSSCQVTTVDSGSKALEFL----GLRQSTDSNDPNAFSKAP 66
Query: 1088 ----FPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEV 1143
+LI+ D MP M GY+ K++++S IP+V +++ + ++CLE
Sbjct: 67 VNHQVVEVNLIITDYCMPGMTGYDLLKKVKESSAFRD--IPVVIMSSENVPARISRCLEE 124
Query: 1144 GMDAYLTKPI 1153
G + + KP+
Sbjct: 125 GAEEFFLKPV 134
>At5g62920 response regulator 6 (ARR6)
Length = 186
Score = 47.0 bits (110), Expect = 6e-05
Identities = 38/148 (25%), Positives = 70/148 (46%), Gaps = 9/148 (6%)
Query: 1025 SLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTE 1084
S + L +L +D+ V ++ +L V V +A+ L G+ E++ +
Sbjct: 21 SPDPLHVLAVDDSHVDRKFIERLLRVSSCKVTVVDSATRALQYL-GLDVEEKSVGFEDLK 79
Query: 1085 ILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVG 1144
+ +LI+ D MP M GYE K+I++S +P+V +++ + +CLE G
Sbjct: 80 V------NLIMTDYSMPGMTGYELLKKIKES--SAFREVPVVIMSSENILPRIDRCLEEG 131
Query: 1145 MDAYLTKPIDFKLMESTILSLTKRETLT 1172
+ +L KP+ ++ SL K E L+
Sbjct: 132 AEDFLLKPVKLSDVKRLRDSLMKVEDLS 159
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.133 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,268,508
Number of Sequences: 26719
Number of extensions: 1080545
Number of successful extensions: 3119
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 34
Number of HSP's that attempted gapping in prelim test: 3035
Number of HSP's gapped (non-prelim): 73
length of query: 1172
length of database: 11,318,596
effective HSP length: 110
effective length of query: 1062
effective length of database: 8,379,506
effective search space: 8899035372
effective search space used: 8899035372
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC121233.1


